Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is an old version of this entry, which may differ significantly from the current revision.
Subjects:
Microbiology
Cedecea, a genus in the Enterobacteriaceae family, includes several opportunistic pathogens reported to cause an array of sporadic acute infections, most notably of the lung and bloodstream. One species, Cedecea neteri, is associated with cases of bacteremia in immunocompromised hosts and has documented resistance to different antibiotics, including β-lactams and colistin. Despite the potential to inflict serious infections, knowledge about drug resistance determinants in Cedecea is limited.
- Cedecea neteri
- opportunistic pathogen
- drug-resistance genes
- β-lactamases
- multidrug efflux pumps
- genomic islands
1. Introduction
The genus Cedecea comprises Gram-negative, facultatively anaerobic bacilli that are non-sporulating and fermentative [1]. Like other genera in the Enterobacteriaceae family, Cedecea species are widely distributed in aquatic and soil environments, as well as associated with plants, insects, the human gut microbiome, and non-human animals (reviewed in [2][3]). Three validly recognized species (Cedecea davisae, Cedecea lapagei, and Cedecea neteri) have documented clinical relevance in humans and collectively, have been reported to cause such diverse acute infections as pneumonia [4][5][6][7][8][9], bacteremia [8][9][10][11][12][13][14][15][16][17][18][19], cutaneous and oral ulcers [13][14][20][21], and dialysis-related peritonitis [22]. Infections attributed to Cedecea occur predominantly in severely immunocompromised hosts, underscoring the opportunistic nature of its pathogenicity.
While Cedecea infections have been reported sporadically in the literature, clinical cases of this emerging opportunistic human pathogen continue to occur worldwide. Clinical strains of Cedecea species exhibit natural resistance to various antimicrobial agents, including ampicillin, ampicillin-sulbactam, cefazolin, cephalothin, cefoxitin, and colistin [3]. The multidrug-resistant phenotype of one C. lapagei isolate was attributed, in part, to the acquisition of the blaNDM-1 gene, which encodes the New Delhi metallo-β-lactamase-1 (NDM-1) [18]. This clinical isolate was resistant to carbapenems, monobactams, and extended-spectrum cephalosporins. Recently, additional blaNDM-1-harboring Cedecea isolates have been reported [23][24]. The dissemination of NDM-1 is of serious public health concern as this metallo-β-lactamase severely limits the chemotherapeutic options available for treating bacterial infections. Apart from the detection of β-lactamase expression in certain Cedecea strains [18][25], knowledge on the genetic basis of drug resistance phenotypes in this genus is limited. The pathogenic potential of Cedecea species is likely under-recognized due to its low reported frequency of association with human infections and the fact that these organisms are not linked to specific disease states but, instead, cause a wide spectrum of acute infections in compromised hosts.
Opportunistic C. neteri infections typically present as bacteremia [10][12], although recently C. neteri was identified as the etiological agent of a urinary tract infection in a pregnant female with polyhydramnios [26]. One of the rare fatal patient outcomes associated with clinical isolates of Cedecea spp. involved a C. neteri infection in the bloodstream of an individual with systemic lupus erythematosus (SLE) [12]. Given the potentially serious nature of these opportunistic infections, research is needed on identifying the genes implicated in drug resistance phenotypes that impact treatment of Cedecea. In this study, we focused our attention on C. neteri because whole-genome sequence (WGS) information is available for multiple strains of this species. WGS data for three environmental strains of C. neteri and different bioinformatics tools were utilized to provide insight into the predicted drug resistance determinants of this species. Only a draft genome at the unassembled contig level is available for the type strain C. neteri ATCC 33855 [10], isolated from a human foot, so we focused our queries on the completed genome sequence of C. neteri SSMD04 [27] and compared results to the genomes of C. neteri M006 [28] and ND14a. Our findings highlight the value of genome-based explorations in contributing to our understanding of antimicrobial resistance in little studied environmental microorganisms with the capability of behaving as clinically relevant opportunistic pathogens.
2. Predicted Multidrug Efflux Pumps in C. neteri
A ubiquitous microbial mechanism of resistance is drug extrusion via integral membrane transporters, which reduces exposure of the bacterial target to active concentrations of the drug. Five different transporter families confer clinical resistance to various antibiotic classes [29]: ATP-binding cassette (ABC), multidrug and toxic compound extrusion (MATE), major facilitator superfamily (MFS), resistance-nodulation-division (RND), and small multidrug resistance (SMR). Database searches revealed the presence of at least 34, 32, and 33 annotated efflux pump or transporter genes in C. neteri strains SSMD04, M006, and ND14a, respectively. Classifications of these predicted efflux pump/transporter genes were distributed across the five different families (ABC, MATE, MFS, RND, and SMR). Efflux pumps of the RND family represent some of the most clinically significant transporter proteins in Gram-negative bacteria because of their broad substrate specificity and association with multidrug resistance (MDR) [30]. Genomes for all three C. neteri strains encoded several RND MDR transporters generally annotated as efflux pumps.
By performing in-depth homology searches against the KEGG database, we found homologs of the RND MDR efflux systems AcrAB-TolC, AcrD, OqxAB, and MdtABCD in the SSMD04 genome (Figure 3). BLAST searches indicated that one predicted RND efflux system resembled both OqxAB and AcrAB. Cnt17100 (ORF JT31_17100) has 95% identity to OqxB in Enterobacter cloacae and 84% identity to AcrB in Xanthomonas citri at the amino acid level. Both OqxB and AcrB are RND transporters that share a consistent transmembrane helical structure [31]. Immediately upstream of JT31_17100 is a gene with 88% amino acid identity to AcrA in E. cloacae and 85% identity to OqxA in Klebsiella variicola. OqxA and AcrA function as membrane fusion proteins (MFPs) in the RND MDR efflux system. Upstream of the oqxAB/acrAB gene cluster is a gene (JT31_17110) encoding an AraC family transcriptional regulator that is transcribed divergently. The deduced gene product shares 84% amino acid identity to the transcriptional activator MarA in Enterobacter ludwigii and 78% identity to RarA in Klebsiella pneumoniae. A second putative regulatory gene (JT31_17095), encoding a Rrf2-type regulator, is located immediately downstream of the oqxAB/acrAB gene cluster and may be involved in repressing transcription of the putative RND MDR efflux system based on its homology to OqxR. Substrates of the AcrAB efflux pump include cationic dyes (acriflavine), detergents, and antibiotics such as penicillins, cephalosporins, fluoroquinolones, macrolides, chloramphenicol, and tetracycline [32]. OqxAB confers resistance to multiple antimicrobial agents (quinoxalines, quinolones, tigecycline, nitrofurantoin, and chloramphenicol), as well as detergents and disinfectants [31]. The function of the E. coli AcrB transporter is dependent on TolC, a multifunctional outer membrane channel [33][34], and a homologous gene with 75% amino acid sequence identity to TolC in Salmonella enterica was identified upstream of the C. neteri acrB/oqxB gene (JT31_17100), suggesting a similar tripartite system in this species (Figure 1). Additionally, the small adaptor protein, AcrZ, which interacts with the AcrAB efflux pump, was identified some distance downstream of the acrB gene in a similar genetic organization as the E. coli cluster [35]. The AcrZ protein was shown to aid in the binding and export of chloramphenicol and tetracycline by the AcrAB efflux pump, thus enhancing the drug resistance phenotype of E. coli [36].
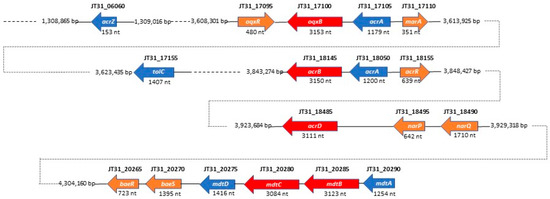
Figure 1. Schematic representation showing the position and size of predicted RND family efflux pump systems and associated genes in the C. neteri SSMD04 genome. RND efflux pumps are displayed in red, structural components in blue, and regulatory genes in orange.
In our analysis, a BLAST search suggested that C. neteri JT31_18485 is a homolog of E. coli AcrD, an aminoglycoside efflux pump from the RND family [37]. The JT31_18485 gene product showed 87% identity in amino acid sequence to AcrD from E. coli and Shigella sonnei. In addition, homologous genes encoding the MdtABCD efflux pump system were identified on the C. neteri SSMD04 chromosome (Figure 3). The C. neteri mdtABCD locus encodes a putative membrane fusion protein (mdtA), two RND-type transporters (mdtB and mdtC), and an MFS-type transporter (mdtD). The predicted MdtB and MdtC transmembrane exporter subunits in C. neteri exhibited high amino acid sequence identity (89% and 90%, respectively) to their counterparts in E. cloacae and E. coli. MdtABCD has previously been shown to comprise a multidrug efflux system that confers resistance to novobiocin and deoxycholate [38][39]. The genes baeS and baeR, which encode a two-component signal transduction system, were identified immediately downstream of the C. neteri mdtABCD locus in a similar genetic organization as described for E. coli. The BaeSR two-component system positively regulates drug resistance in E. coli via the MdtABCD multidrug efflux system [39][40], suggesting that the predicted mdtABCD locus in C. neteri may be under the transcriptional control of the BaeR response regulator. These putative RND efflux pump systems likely contribute to the intrinsic antimicrobial drug resistance reported for clinical isolates of C. neteri. However, the functional role of these homologs of RND efflux pumps and the identity of the specific drug substrates of each pump remain to be established.
A KEGG database search also revealed two pairs of linked genes annotated as emrA-like and emrB-like MDR transporters of the MFS family on the chromosomes of all three C. neteri strains. In C. neteri SSMD04, these emrA/emrB-like gene pairs are JT31_07770/JT31_07775 and JT31_16920/JT31_16915. Multiple sequence alignment showed that the JT31_16920/JT31_16915 pair had the highest sequence identity with the known EmrAB counterpart in Klebsiella pneumoniae, Salmonella enterica, and E. coli. The emrAB locus in E. coli encodes a MDR pump involved in the extrusion of chemically unrelated antimicrobial agents, including the antibiotics nalidixic acid and thiolactomycin [41]. The putative SSMD04 emrB, which encodes a MFS-type efflux pump, shared 93% amino acid sequence identity with K. pneumoniae EmrB, 92% identity with E. coli EmrB, and 90% identity with S. enterica EmrB. The putative emrA, which encodes a membrane fusion protein, shared high amino acid sequence identity (82%-86%) with EmrA in K. pneumoniae, E. coli, and S. enterica. In addition, a gene (JT31_16925) annotated as mprA is located adjacent to emrA. Previous research demonstrated that mprA (renamed emrR) is part of the emrAB operon and functions to repress transcription of emrAB [42]. A recent study showed that the EmrAB pump system contributes to colistin resistance in the nosocomial pathogen Acinetobacter baumannii [43]. Colistin resistance has been noted as one of the defining properties characterizing Cedecea species and is a trait shared by established opportunistic pathogens in the genus Serratia [1][10], but the specific mechanism conferring this resistance in Cedecea is not known. The possibility that the predicted MFS-type EmrAB efflux pump system may be responsible, at least in part, for the colistin resistance phenotype in Cedecea should be explored further, particularly since these species are gaining increased recognition as opportunistic pathogens in the clinical setting.
3. Conclusions
Cedecea species are members of the Enterobacteriaceae family that have been found in a wide range of natural environments, as well as in human clinical specimens. Reported clinical isolates have been associated with a spectrum of acute infections (e.g., pneumonia, bacteremia, oral ulcers, and dialysis-related peritonitis) in primarily immunocompromised hosts, and antibiotic susceptibility testing has indicated varying degrees of drug resistance among documented isolates. As emerging opportunistic pathogens of environmental origin, C. neteri and the closely related species C. davisae and C. lapagei have received little research attention to date. This study exploited whole-genome sequence information for three C. neteri strains (SSMD04, M006, and ND14a) to gain a deeper understanding of the genetic potential for drug resistance in this species and to identify drug-resistance candidate genes for further investigation. We focused our genomic analyses on C. neteri SSMD04, an isolate originating from retailed sashimi, since only a draft genome was available for type strain C. neteri ATCC 33855. Our work reports the presence of multiple β-lactamase-encoding genes in the C. neteri SSMD04 chromosome, including four putative MBLs, a CMY/ACT-type AmpC variant, and a novel β-lactamase gene not described previously. Homologous genes encoding RND- and MFS-type efflux pumps were also identified, along with associated regulatory genes known to be involved in the control of these efflux systems in other bacteria. Comparative analysis of predicted genomic islands suggested the acquisition of some drug resistance determinants and virulence factors by horizontal genetic transfer. The findings of this study advance our currently limited understanding of the molecular basis of antimicrobial resistance in C. neteri. Future research is needed to correlate the genetic data with resistance phenotypes impacting public health management of this opportunistic pathogen.
This entry is adapted from the peer-reviewed paper 10.3390/microorganisms9081741
References
- Grimont, P.; Grimont, F.; Farmer III, J.; Asbury, M. Cedecea davisae gen. nov., sp. nov. and Cedecea lapagei sp. nov., new Enterobacteriaceae from clinical specimens. Int. J. Syst. Bacteriol. 1981, 31, 317–326.
- Dalamaga, M.; Vrioni, G. Cedecea. In Molecular Detection of Human Bacterial Pathogens; Liu, D., Ed.; CRC Press/Taylor and Francis Group: Boca Raton, FL, USA, 2011; pp. 817–825.
- Thompson, D.K.; Sharkady, S.M. Expanding spectrum of opportunistic Cedecea infections: Current clinical status and multidrug resistance. Int. J. Infect. Dis. 2020, 100, 461–469.
- Bae, B.H.C.; Sureka, S.B.; Ajamy, J.A. Enteric group 15 (Enterobacteriaceae) associated with pneumonia. J. Clin. Microbiol. 1981, 14, 596–597.
- Ismaael, T.G.; Zamora, E.M.; Khasawneh, F.A. Cedecea davisae’s role in a polymicrobial lung infection in a cystic fibrosis patient. Case Rep. Infect. Dis. 2012, 2012, 176864.
- Lopez, L.A.; Ibarra, B.S.; de la Garza, J.A.; Rada Fde, J.; Nuñez, A.I.; López, M.G. First reported case of pneumonia caused by Cedecea lapagei in America. Braz. J. Infect. Dis. 2013, 17, 626–628.
- Hong, S.K.; Lee, J.S.; Kim, E.C. First Korean case of Cedecea lapagei pneumonia in a patient with chronic obstructive pulmonary disease. Ann. Lan. Med. 2015, 35, 266–268.
- Kury, C.M.H.; Yabrudi, A.A.; de Souza, T.B.; de Souza, E.C.; Silva Costa, L.T.E.; Soares, C.B.; Calixto, G.A.; Gramático, M.R. First reported case of ventilator-associated pneumonia and sepsis caused by Cedecea lapagei in a Brazilian neonatal intensive care unit. J. Pediatric Infect. Dis. Soc. 2017, 6, 209–210.
- Ramaswamy, V.V.; Gummadapu, S.; Suryanarayana, N. Nosocomial pneumonia and sepsis caused by a rare organism Cedecea lapagei in an infant and a review of literature. BMJ Case Rep. 2019, 12, e229854.
- Farmer III, J.J.; Sheth, N.K.; Hudzinski, J.A.; Rose, H.D.; Asbury, M.F. Bacteremia due to Cedecea neteri sp. nov. J. Clin. Microbiol. 1982, 16, 775–778.
- Perkins, S.R.; Beckett, T.A.; Bump, C.M. Cedecea davisae bacteremia. J. Clin. Microbiol. 1986, 24, 675–676.
- Aguilera, A.; Pascual, J.; Loza, E.; Lopez, J.; Garcia, G.; Liaño, F.; Quereda, C.; Ortuño, J. Bacteraemia with Cedecea neteri in a patient with systemic lupus erythematosus. Postgrad. Med. J. 1995, 71, 179–180.
- Dalamaga, M.; Karmaniolas, K.; Arsenis, G.; Pantelaki, M.; Daskalopoulou, K.; Papadavid, E.; Migdalis, I. Cedecea lapagei bacteremia following cement-related chemical burn injury. Burns 2008, 34, 1205–1207.
- Dalamaga, M.; Pantelaki, M.; Karmaniolas, K.; Matekovits, A.; Daskalopoulou, K. Leg ulcer and bacteremia due to Cedecea davisae. Eur. J. Dermatol. 2008, 18, 204–205.
- Abate, G.; Qureshi, S.; Mazumder, S.A. Cedecea davisae bacteremia in a neutropenic patient with acute myeloid leukemia. J. Infect. 2011, 63, 83–85.
- Akinosoglou, K.; Perperis, A.; Siagris, D.; Goutou, P.; Spiliopoulou, I.; Gogos, C.A.; Marangos, M. Bacteraemia due to Cedecea davisae in a patient with sigmoid colon cancer: A case report and brief review of the literature. Diagn. Microbiol. Infect. Dis. 2012, 74, 303–306.
- Peretz, A.; Simsolo, C.; Farber, E.; Roth, A.; Brodsky, D.; Nakhoul, F. A rare bacteremia caused by Cedecea davisae in patient with chronic renal disease. Am. J. Case Rep. 2013, 14, 216–218.
- Ahmad, N.; Ali, S.M.; Khan, A.U. First reported New Delhi metallo-β-lactamase-1-producing Cedecea lapagei. Int. J. Antimicrob. Agents 2017, 49, 118–119.
- Kanakadandi, V.S.; Sarao, M.S.; Cunningham, J.M. A rare case of Cedecea davisae bacteremia presenting as biliary sepsis. Cureus 2019, 11, e5298.
- Mawardi, H.; Pavlakis, M.; Mandelbrot, D.; Woo, S.B. Sirolimus oral ulcer with Cedecea davisae superinfection. Transpl. Infect. Dis. 2010, 12, 446–450.
- Biswal, I.; Hussain, N.A.; Grover, R.K. Cedecea lapagei in a patient with malignancy: Report of a rare case. J. Cancer Res. Ther. 2015, 11, 646.
- Davis, O.; Wall, B.M. “Broom straw peritonitis” secondary to Cedecea lapagei in a liver transplant recipient. Perit. Dial. Int. 2006, 26, 512–513.
- Ejaz, H.; Alzahrani, B.; Hamad, M.F.S.; Abosalif, K.O.A.; Junaid, K.; Abdalla, A.E.; Elamir, M.Y.M.; Aljaber, N.J.; Hamam, S.S.M.; Younas, S. Molecular analysis of the antibiotic resistant NDM-1 gene in clinical isolates of Enterobacteriaceae. Clin. Lab. 2020, 66, 409–417.
- Khalid, S.; Ahmad, N.; Ali, S.M.; Khan, A.U. Outbreak of efficiently transferred carbapenem-resistant blaNDM-producing gram-negative bacilli isolated from neonatal intensive care unit of an Indian hospital. Microb. Drug Resist. 2020, 26, 284–289.
- Ammenouche, N.; Dupont, H.; Mammeri, H. Characterization of a novel AmpC β-lactamase produced by a carbapenem-resistant Cedecea davisae clinical isolate. Antimicrob. Agents Chemother. 2014, 58, 6942–6945.
- Ahmad, H.; Masroor, T.; Parmar, S.A.; Panigrahi, D. Urinary tract infection by a rare pathogen Cedecea neteri in a pregnant female with polyhydramnios: Rare case report from UAE. BMC Infect. Dis. 2021, 21, 637.
- Chan, K.G.; Tan, K.H.; Yin, W.F.; Tan, J.Y. Complete genome sequence of Cedecea neteri strain SSMD04, a bacterium isolated from pickled mackerel sashimi. Genome Announc. 2014, 2, e01339-14.
- Chan, K.G.; Tan, W.S. Insights into Cedecea neteri strain M006 through complete genome sequence, a rare bacterium from aquatic environment. Stand. Genomic Sci. 2017, 12, 40.
- Li, X.Z.; Nikaido, H. Efflux-mediated drug resistance in bacteria: An update. Drugs 2009, 69, 1555–1623.
- Blair, J.M.A.; Richmond, G.E.; Piddock, L.J.V. Multidrug efflux pumps in gram-negative bacteria and their role in antibiotic resistance. Future Microbiol. 2014, 9, 1165–1177.
- Li, J.; Zhang, H.; Ning, J.; Sajid, A.; Cheng, G.; Yuan, Z.; Hao, H. The nature and epidemiology of OqxAB, a multidrug efflux pump. Antimicrob. Resist. Infect. Control 2019, 8, 44.
- Nikaido, H.; Takatsuka, Y. Mechanisms of RND multidrug efflux pumps. Biochim. Biophys. Acta. 2009, 1794, 769–781.
- Fralick, J.A. Evidence that TolC is required for functioning of the Mar/AcrAB efflux pump of Escherichia coli. J. Bacteriol. 1996, 178, 5803–5805.
- Nishino, K.; Yamada, J.; Hirakawa, H.; Hirata, T.; Yamaguchi, A. Roles of TolC-dependent multidrug transporters of Escherichia coli in resistance to β-lactams. Antimicrob. Agents Chemother. 2003, 47, 3030–3033.
- Anes, J.; McCusker, M.P.; Fanning, S.; Martins, M. The ins and outs of RND efflux pumps in Escherichia coli. Front. Microbiol. 2015, 6, 587.
- Hobbs, E.C.; Yin, X.; Paul, B.J.; Astarita, J.L.; Storz, G. Conserved small protein associates with the multidrug efflux pump AcrB and differentially affects antibiotic resistance. Proc. Natl. Acad. Sci. USA 2012, 109, 16696–16701.
- Rosenberg, E.Y.; Ma, D.; Nikaido, H. AcrD of Escherichia coli is an aminoglycoside efflux pump. J. Bacteriol. 2000, 182, 1754–1756.
- Nishino, K.; Yamaguchi, A. Analysis of a complete library of putative drug transporter genes in Escherichia coli. J. Bacteriol. 2001, 183, 5803–5812.
- Baranova, N.; Nikaido, H. The BaeSR two-component regulatory system activates transcription of the yegMNOB (mdtABCD) transporter gene cluster in Escherichia coli and increases its resistance to novobiocin and deoxycholate. J. Bacteriol. 2002, 184, 4168–4176.
- Nagakubo, S.; Nishino, K.; Hirata, T.; Yamaguchi, A. The putative response regulator BaeR stimulates multidrug resistance of Escherichia coli via a novel multidrug exporter system, MdtABC. J. Bacteriol. 2002, 184, 4161–4167.
- Lomovskaya, O.; Lewis, K. emr, an Escherichia coli locus for multidrug resistance. Proc. Natl. Acad. Sci. USA 1992, 89, 8938–8942.
- Lomovskaya, O.; Lewis, K.; Matin, A. EmrR is a negative regulator of the Escherichia coli multidrug resistance pump EmrAB. J. Bacteriol. 1995, 177, 2328–2334.
- Lin, M.F.; Lin, Y.Y.; Lan, C.Y. Contribution of EmrAB efflux pumps to colistin resistance in Acinetobacter baumannii. J. Microbiol. 2017, 55, 130–136.
This entry is offline, you can click here to edit this entry!
