Acinetobacter baumannii is an emerging pathogen, and over the last three decades it has proven to be particularly difficult to treat by healthcare services. It is now regarded as a formidable infectious agent with a genetic setup for prompt development of resistance to most of the available antimicrobial agents. we provide a comprehensive updated overview of the available data about A. baumannii, the multi-drug resistant (MDR) phenotype spread, carbapenem-resistance, and the associated genetic resistance determinants in low-income countries (LIICs) since the beginning of the 21st century.
- A. baumannii
- low-income
- surveillance
- developing countries
- carbapenem-resistance
- MDR
- Syria
- Ethiopia
- COVID-19
1. Introduction
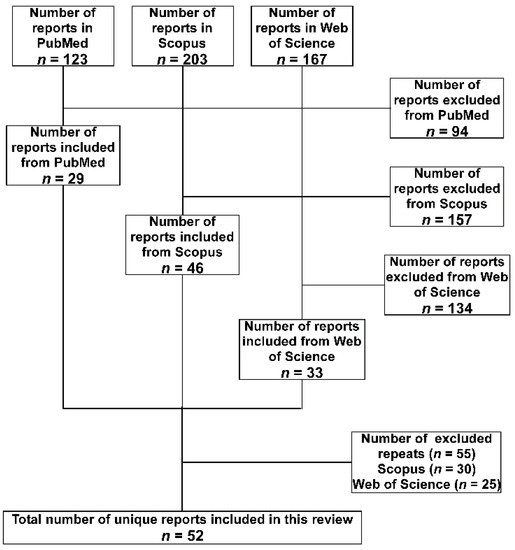
2. Methodology
3. A. baumannii in the LICs Distributed over the Different Geographical Regions
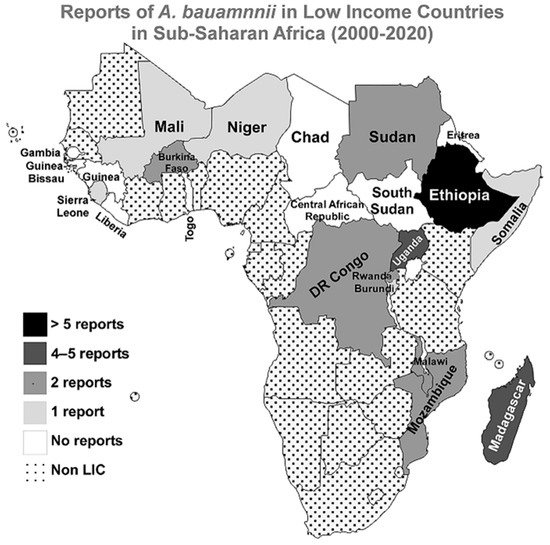
3.1. Sub-Saharan Africa
3.2. Middle East and North Africa
3.3. South Asia
3.4. Latin America and the Caribbean
3.5. Europe and Central Asia
3.6. East Asia and Pacific
| Country | Study | Isolates (n) | MDR % * | CRAB% | Isolates Characterization | References |
|---|---|---|---|---|---|---|
| Sub-Saharan Africa | ||||||
| Ethiopia | Kempf et al., 2012 | 40 | NA | NA | rpoB and recA sequencing for genotyping | [10] |
| Lema et al., 2012 | 5 | ≥20% | NA | AST with KB | [11] | |
| Pritsch et al., 2017 | 3 | 100% | 100% | AST with KB and VITEK 2, CT102 Micro-Array, real-time PCR, WGS, MLST, and detection of the blaNDM-1 | [12] | |
| Solomon et al., 2017 | 43 | 81% | 37% | AST with KB and phenotypic detection of ESBLs and MBLs | [13] | |
| Bitew et al., 2017 | 2 | 100% | NA | Identification and AST with VITEK 2 | [15] | |
| Demoz et al., 2018 | 1 | 100% | 100% | AST with KB | [18] | |
| Gashaw et al., 2018 | 2 | 50% XDR and 50% PDR | 100% | AST with KB and phenotypic detection of ESBLs and AmpC | [19] | |
| Moges et al., 2019 | 15 | ≥63% | Yes | AST with KB and phenotypic detection of ESBLs and carbapenemases | [14] | |
| Admas et al., 2020 | 6 | 100% | NA | Identification and AST with VITEK 2 | [16] | |
| Motbainor et al., 2020 | 9 | 100% | 33% | Identification with VITEK 2 and AST with KB | [17] | |
| Madagascar | Randrianirina et al., 2010 | 50 | ≥44% | 44% | AST with KB and phenotypic detection of ESBLs | [20] |
| Andriamanantena et al., 2010 | 53 | 100% | 100% | AST with KB and MIC determination, phenotypic detection of carbapenemases, ReP-PCR for genotyping and PCR for detection of; blaAmpC, blaoxa51, blaoxa23, blaoxa24, blaVIM, blaIMP, and isAba-1 | [21] | |
| Rasamiravaka et al., 2015 | 10 | ≥50% | 0% | AST with KB | [22] | |
| Tchuinte et al., 2019 | 15 | 100% | 100% | MALDI-TOF MS for identification, AST with KB and MIC determination, WGS, MLST for genotyping and WGS detecting; blaoxa51, blaoxa23, blaoxa24, blaoxa58, and isAba-1 | [23] | |
| Eremeeva et al., 2019 | 14 | NA | NA | TaqMan PCR of the rpoB for identification, and PCR for detecting: blaoxa51-like, blaoxa23, blaoxa24, blaVIM, and blaIMP | [24] | |
| Uganda | Kateete et al., 2016 | 40 | 60% | 38% | AST with Phoenix Automated Microbiology System, PCR for: blaoxa51-like, blaoxa51, blaoxa23, blaoxa24, blaoxa58, blaVIM, blaSPM, and blaIMP | [26] |
| Kateete et al., 2017 | 20 | 40% | 35% | AST with MIC determination, PAMS, Rep-PCR for genotyping and phenotypic detection of ESBLs and AmpC | [27] | |
| Moore et al., 2019 | 3 | NA | NA | qPCR TAC | [25] | |
| Aruhomukama et al., 2019 | 1077 | 3% | 3% | AST with KB, PCR for detecting: blaoxa23, blaoxa24, blaoxa58, blaVIM, blaSPM, blaKPC, and blaIMP, phenotypic detection of carbapenemases, and conjugation to show transferability of blaVIM. | [28] | |
| Burkina Faso | Kaboré et al., 2016 | 3 | 100% | NA | AST with KB and phenotypic detection of ESBLs | [29] |
| Sanou et al., 2021 # | 5 | 100% | 60% | MALDI-TOF MS for identification, AST with KB and MIC determination, phenotypic detection of ESBLs, PCR and sequencing of multiple resistance genes including; blaoxa1-like, blaoxa48-like, blaNDM, blaVIM, blaSPM, blaKPC, blaCTX-M, and blaIMP, and MLST for genotyping. | [30] | |
| DR of the Congo | Lukuke et al., 2017 | 2 | 0% | NA | API for identification and AST with KB | [32] |
| Koyo et al., 2019 | 15 | NA | NA | qPCR and phylogenetic analysis using the rpoB gene | [33] | |
| Malawi | Bedell et al., 2012 | 1 | NA | NA | Identification with standard diagnostic techniques | [34] |
| Iroh Tam et al., 2019 | 84 | ≥44% | NA | API for identification, AST with KB, and phenotypic detection of ESBLs | [35] | |
| Mozambique | Martínez et al., 2016 | 1 | NA | NA | 16S rRNA PCR and MALDI-TOF MS for identification | [37] |
| Hurtado et al., 2019 | 1 | 100% | 0% | 16S rRNA for identification and AST with KB | [36] | |
| Sudan | Mohamed et al., 2019 | 1 | NA | NA | API for identification followed by WGS | [38] |
| Dirar et al., 2020 | 12 | ≥83% | 89% | Identification with PAMS, AST with KB and phenotypic detection of ESBLs and carbapenemases. | [39] | |
| Rwanda | La Scola and Raoult 2004 | 10 | NA | NA | API for identification and recA genotyping | [40] |
| Heiden et al., 2020 | 1 | 100% | 0% | MALDI-TOF MS for identification, AST with VITEK 2, phenotypic detection of ESBLs and carbapenemases, and WGS | [41] | |
| Burundi | La Scola and Raoult 2004 | 3 | NA | NA | API for identification and recA genotyping | [40] |
| Mali | Doumbia-Singare et al., 2014 | 1 | NA | NA | Not mentioned | [42] |
| Sierra Leone | Lakoh et al., 2020 | 14 | ≥40% | 10% | Identification and AST with VITEK 2 | [43] |
| Somalia | Mohamed et al., 2020 | 7 | 100% | 100% | AST with KB | [44] |
| Niger | Louni et al., 2018 | 29 | NA | NA | qPCR and rpoB PCR for identification and phylogenetic analysis | [45] |
| Central African Republic | No Reports | |||||
| Chad | No Reports | |||||
| Eritrea | No Reports | |||||
| Gambia | No Reports | |||||
| Guinea | No Reports | |||||
| Guinea-Bissau | No Reports | |||||
| Liberia | No Reports | |||||
| South Sudan | No Reports | |||||
| Togo | No Reports | |||||
| Middle East and North Africa | ||||||
| Syria | Hamzeh et al., 2012 | 260 | ≥65% | 65% | Identification and AST with PAMS | [46] |
| Teicher et al., 2014 | 6 | 100% | 80% | API for identification and AST with MicroScan Walk-Away System | [47] | |
| Peretz et al., 2014 | 5 | 100% | NA | Not mentioned | [50] | |
| Rafei et al., 2014 | 4 | 100% | 100% | Identification with rpoB sequencing and blaoxa51, PCR, AST with KB and Etest, PCR for: blaoxa23-like, blaoxa24-like, blaoxa58-like, and blaNDM, and PFGE for genotyping | [53] | |
| Heydari et al., 2015 | 1 | 100% | 100% | Identification and AST with VITEK 2, phenotypic detection of ESBLs and carbapenemases, PCR for the blaNDM, PFGE and MLST for typing | [51] | |
| Rafei et al., 2015 | 59 | Yes | 74% | Identification with MALDI-TOF MS, rpoB sequencing and blaoxa51 PCR, AST with KB and Etest, PCR for detecting: blaoxa23, blaoxa24, blaoxa58, blaNDM-1, blaVIM, blaoxa143, and blaIMP, and MLST for typing | [54] | |
| Herard and Fakhri 2017 | 38 | NA | NA | Not mentioned | [48] | |
| Salloum et al., 2018 | 2 | 100% | 100% | AST with KB and Etest, PCR for blaoxa58 and blaNDM, plasmid typing with PBRT, MLST, and WGS | [55] | |
| Fily et al., 2019 | 6 | NA | 67% | AST with KB | [49] | |
| Hasde et al., 2019 | 5 | NA | NA | Not mentioned | [52] | |
| Yemen | Bakour et al., 2014 | 3 | 100% | 100% | API and MALDI-TOF MS for identification, AST with KB and E-test, phenotypic detection of carbapenemases, PCR detection of: blaoxa23, blaoxa24, blaoxa58, blaNDM, blaVIM, blaSIM, blaoxa48-like, blaIMP and others, and MLST | [56] |
| Fily et al., 2019 | 1 | NA | 100% | AST with KB | [49] | |
| South Asia | ||||||
| Afghanistan | Sutter et al., 2011 | 57 ¥ | ≥75% | 76% | Identification and AST with MicroScan autoSCAN-4 | [57] |
| Latin America and The Caribbean | ||||||
| Haiti | Potron et al., 2011 | 3 | 66.7% | 0% | API and 16sRNA for identification, AST with KB and E-test, phenotypic detection of ESBLs, PCR for detection of: blaTEM, blaSHV, blaPER-1, blaVEB-1, blaGES-1, and blaCTX-M | [58] |
| Marra et al., 2012 | 1 | 100% | 0% | Identification and AST with VITEK 2 | [59] | |
| Murphy et al., 2016 | 4 | ≥25% | 25% | AST but the method was not indicated | [60] | |
| Chaintarli et al., 2018 | 2 | 0% | 0% | Identification and AST with VITEK 2 and phenotypic detection of ESBLs | [61] | |
| Roy et al., 2018 | 0 ϕ | NA | NA | Metagenomic analyses of water samples | [62] | |
| Europe and Central Asia | ||||||
| Tajikistan | No Reports | |||||
| East Asia and Pacific | ||||||
| Democratic People’s Republic of Korea | No Reports | |||||
This entry is adapted from the peer-reviewed paper 10.3390/antibiotics10070764
References
- Vazquez-Lopez, R.; Solano-Galvez, S.G.; Juarez Vignon-Whaley, J.J.; Abello Vaamonde, J.A.; Padro Alonzo, L.A.; Rivera Resendiz, A.; Muleiro Alvarez, M.; Vega Lopez, E.N.; Franyuti-Kelly, G.; Alvarez-Hernandez, D.A.; et al. Acinetobacter baumannii Resistance: A Real Challenge for Clinicians. Antibiotics 2020, 9, 205.
- Howard, A.; O’Donoghue, M.; Feeney, A.; Sleator, R.D. Acinetobacter baumannii: An emerging opportunistic pathogen. Virulence 2012, 3, 243–250.
- Peleg, A.Y.; Seifert, H.; Paterson, D.L. Acinetobacter baumannii: Emergence of a successful pathogen. Clin. Microbiol. Rev. 2008, 21, 538–582.
- Jawad, A.; Seifert, H.; Snelling, A.M.; Heritage, J.; Hawkey, P.M. Survival of Acinetobacter baumannii on dry surfaces: Comparison of outbreak and sporadic isolates. J. Clin. Microbiol. 1998, 36, 1938–1941.
- Espinal, P.; Marti, S.; Vila, J. Effect of biofilm formation on the survival of Acinetobacter baumannii on dry surfaces. J. Hosp. Infect. 2012, 80, 56–60.
- Wagenvoort, J.H.; Joosten, E.J. An outbreak Acinetobacter baumannii that mimics MRSA in its environmental longevity. J. Hosp. Infect. 2002, 52, 226–227.
- Asif, M.; Alvi, I.A.; Rehman, S.U. Insight into Acinetobacter baumannii: Pathogenesis, global resistance, mechanisms of resistance, treatment options, and alternative modalities. Infect. Drug Resist. 2018, 11, 1249.
- Vaara, M. Polymyxins and Their Potential Next Generation as Therapeutic Antibiotics. Front. Microbiol. 2019, 10, 1689.
- World-Bank. World Bank Country and Lending Groups-Country Classification; The World Bank Group: Washington, DC, USA, 2020.
- Kempf, M.; Abdissa, A.; Diatta, G.; Trape, J.-F.; Angelakis, E.; Mediannikov, O.; La Scola, B.; Raoult, D. Detection of Acinetobacter baumannii in human head and body lice from Ethiopia and identification of new genotypes. Int. J. Infect. Dis. 2012, 16, e680–e683.
- Lema, T.; Woldeamanuel, Y.; Asrat, D.; Hunegnaw, M.; Baraki, A.; Kebede, Y.; Yamuah, L.; Aseffa, A. The pattern of bacterial isolates and drug sensitivities of infected ulcers in patients with leprosy in ALERT, Kuyera and Gambo hospitals, Ethiopia. Lepr. Rev. 2012, 83, 40–51.
- Pritsch, M.; Zeynudin, A.; Messerer, M.; Baumer, S.; Liegl, G.; Schubert, S.; Löscher, T.; Hoelscher, M.; Belachew, T.; Rachow, A. First report on blaNDM-1-producing Acinetobacter baumannii in three clinical isolates from Ethiopia. BMC Infect. Dis. 2017, 17, 180.
- Solomon, F.B.; Wadilo, F.; Tufa, E.G.; Mitiku, M. Extended spectrum and metalo beta-lactamase producing airborne Pseudomonas aeruginosa and Acinetobacter baumanii in restricted settings of a referral hospital: A neglected condition. Antimicrob. Resist. Infect. Control 2017, 6, 106.
- Moges, F.; Eshetie, S.; Abebe, W.; Mekonnen, F.; Dagnew, M.; Endale, A.; Amare, A.; Feleke, T.; Gizachew, M.; Tiruneh, M. High prevalence of extended-spectrum beta-lactamase-producing Gram-negative pathogens from patients attending Felege Hiwot Comprehensive Specialized Hospital, Bahir Dar, Amhara region. PLoS ONE 2019, 14, e0215177.
- Bitew, A.; Molalign, T.; Chanie, M. Species distribution and antibiotic susceptibility profile of bacterial uropathogens among patients complaining urinary tract infections. BMC Infect. Dis. 2017, 17, 654.
- Admas, A.; Gelaw, B.; Worku, A.; Melese, A. Proportion of bacterial isolates, their antimicrobial susceptibility profile and factors associated with puerperal sepsis among post-partum/aborted women at a referral Hospital in Bahir Dar, Northwest Ethiopia. Antimicrob. Resist. Infect. Control 2020, 9, 14.
- Motbainor, H.; Bereded, F.; Mulu, W. Multi-drug resistance of blood stream, urinary tract and surgical site nosocomial infections of Acinetobacter baumannii and Pseudomonas aeruginosa among patients hospitalized at Felegehiwot referral hospital, Northwest Ethiopia: A cross-sectional study. BMC Infect. Dis. 2020, 20, 92.
- Demoz, G.T.; Alebachew, M.; Legesse, Y.; Ayalneh, B. Treatment of ventriculoperitoneal shunt infection and ventriculitis caused by Acinetobacter baumannii: A case report. J. Med. Case Rep. 2018, 12, 141.
- Gashaw, M.; Berhane, M.; Bekele, S.; Kibru, G.; Teshager, L.; Yilma, Y.; Ahmed, Y.; Fentahun, N.; Assefa, H.; Wieser, A. Emergence of high drug resistant bacterial isolates from patients with health care associated infections at Jimma University medical center: A cross sectional study. Antimicrob. Resist. Infect. Control 2018, 7, 138.
- Randrianirina, F.; Vaillant, L.; Ramarokoto, C.E.; Rakotoarijaona, A.; Andriamanarivo, M.L.; Razafimahandry, H.C.; Randrianomenjanahary, J.; Raveloson, J.R.; Hariniana, E.R.; Carod, J.-F. Antimicrobial resistance in pathogens causing nosocomial infections in surgery and intensive care units of two hospitals in Antananarivo, Madagascar. J. Infect. Dev. Ctries. 2010, 4, 074–082.
- Andriamanantena, T.S.; Ratsima, E.; Rakotonirina, H.C.; Randrianirina, F.; Ramparany, L.; Carod, J.-F.; Richard, V.; Talarmin, A. Dissemination of multidrug resistant Acinetobacter baumannii in various hospitals of Antananarivo Madagascar. Ann. Clin. Microbiol. Antimicrob. 2010, 9, 17.
- Rasamiravaka, T.; Sheila, H.S.; Rakotomavojaona, T.; Rakoto-Alson, A.; Rasamindrakotroka, A. Changing profile and increasing antimicrobial resistance of uropathogenic bacteria in Madagascar. Med. Mal. Infect. 2015, 45, 173–176.
- Tchuinte, P.L.S.; Rabenandrasana, M.A.N.; Kowalewicz, C.; Andrianoelina, V.H.; Rakotondrasoa, A.; Andrianirina, Z.Z.; Enouf, V.; Ratsima, E.H.; Randrianirina, F.; Collard, J.-M. Phenotypic and molecular characterisations of carbapenem-resistant Acinetobacter baumannii strains isolated in Madagascar. Antimicrob. Resist. Infect. Control 2019, 8, 31.
- Eremeeva, M.E.; Warang, S.S.; Anderson, M.L.; Capps, D.; Zohdy, S.; Durden, L.A. Molecular Survey for Pathogens and Markers of Permethrin Resistance in Human Head Lice (Phthiraptera: Pediculidae) from Madagascar. J. Parasitol. 2019, 105, 459–468.
- Moore, C.C.; Jacob, S.T.; Banura, P.; Zhang, J.; Stroup, S.; Boulware, D.R.; Scheld, W.M.; Houpt, E.R.; Liu, J. Etiology of sepsis in Uganda using a quantitative polymerase chain reaction-based TaqMan array card. Clin. Infect. Dis. 2019, 68, 266–272.
- Kateete, D.P.; Nakanjako, R.; Namugenyi, J.; Erume, J.; Joloba, M.L.; Najjuka, C.F. Carbapenem resistant Pseudomonas aeruginosa and Acinetobacter baumannii at Mulago Hospital in Kampala, Uganda (2007–2009). Springerplus 2016, 5, 1308.
- Kateete, D.P.; Nakanjako, R.; Okee, M.; Joloba, M.L.; Najjuka, C.F. Genotypic diversity among multidrug resistant Pseudomonas aeruginosa and Acinetobacter species at Mulago Hospital in Kampala, Uganda. BMC Res. Notes 2017, 10, 1–10.
- Aruhomukama, D.; Najjuka, C.F.; Kajumbula, H.; Okee, M.; Mboowa, G.; Sserwadda, I.; Mayanja, R.; Joloba, M.L.; Kateete, D.P. blaVIM-and blaOXA-mediated carbapenem resistance among Acinetobacter baumannii and Pseudomonas aeruginosa isolates from the Mulago hospital intensive care unit in Kampala, Uganda. BMC Infect. Dis. 2019, 19, 853.
- Kaboré, W.A.; Konaté, A.; Bako, E.; Bagré, T.S.; Boisramé, S.; Chandad, F.; Traoré, A.S.; Barro, N.; Sangaré, L. Détection d’Acinetobacter baumannii, agent pathogène opportuniste et multirésistant dans les infections bucco-dentaires à Ouagadougou, Burkina Faso. Médecine Buccale Chir. Buccale 2016, 22, 105–112.
- Sanou, S.; Ouedraogo, A.S.; Aberkane, S.; Vendrell, J.; Ouchar, O.; Bouzimbi, N.; Hema, A.; Poda, A.; Zoungrana, J.; Ouedraogo, G.A. Prevalence and Molecular Characterization of Extended Spectrum β-Lactamase, Plasmid-Mediated Quinolone Resistance, and Carbapenemase-Producing Gram-Negative Bacilli in Burkina Faso. Microb. Drug Resist. 2021, 27, 18–24.
- El Far, M.Y.; El-Mahallawy, H.A.; Attia, A.S. Tracing the dissemination of the international clones of multidrug-resistant Acinetobacter baumannii among cancer patients in Egypt using the PCR-based open reading frame typing (POT) method. J. Glob. Antimicrob. Resist. 2019, 19, 210–215.
- Lukuke, H.M.; Kasamba, E.; Mahuridi, A.; Ngatu, N.R.; Narufumi, S.; Mukengeshayi, A.N.; Malou, V.; Makoutode, M.; Kaj, F.M. L’incidence des infections nosocomiales urinaires et des sites opératoires dans la maternité de l’Hôpital Général de Référence de Katuba à Lubumbashi en République Démocratique du Congo. Pan Afr. Med. J. 2017, 28, 57.
- Koyo, C.S.B.; Amanzougaghene, N.; Davoust, B.; Tshilolo, L.; Lekana-Douki, J.B.; Raoult, D.; Mediannikov, O.; Fenollar, F. Genetic diversity of human head lice and molecular detection of associated bacterial pathogens in Democratic Republic of Congo. Parasites Vectors 2019, 12, 290.
- Bedell, R.A.; Anderson, S.T.; Van Lettow, M.; Åkesson, A.; Corbett, E.L.; Kumwenda, M.; Chan, A.K.; Heyderman, R.S.; Zachariah, R.; Harries, A.D. High prevalence of tuberculosis and serious bloodstream infections in ambulatory individuals presenting for antiretroviral therapy in Malawi. PLoS ONE 2012, 7, e39347.
- Iroh Tam, P.-Y.; Musicha, P.; Kawaza, K.; Cornick, J.; Denis, B.; Freyne, B.; Everett, D.; Dube, Q.; French, N.; Feasey, N. Emerging resistance to empiric antimicrobial regimens for pediatric bloodstream infections in Malawi (1998–2017). Clin. Infect. Dis. 2019, 69, 61–68.
- Hurtado, J.C.; Carrilho, C.; Mandomando, I.; Martínez, M.J.; CaDMIA bacterial study group. Fatal multi-drug-resistant Acinetobacter baumannii pneumonia in Maputo, Mozambique: A case report. Enferm. Infecc. Y Microbiol. Clínica 2019, 37, 485–487.
- Martínez, M.J.; Massora, S.; Mandomando, I.; Ussene, E.; Jordao, D.; Lovane, L.; Muñoz-Almagro, C.; Castillo, P.; Mayor, A.; Rodriguez, C. Infectious cause of death determination using minimally invasive autopsies in developing countries. Diagn. Microbiol. Infect. Dis. 2016, 84, 80–86.
- Mohamed, S.B.; Hassan, M.; Munir, A.; Kambal, S.; Abdalla, N.I.; Hamad, A.; Mohammed, S.; Ahmed, F.; Hamid, O.; Ismail, A. Whole-Genome Sequence of Acinetobacter baumannii Strain NUBRI-A, Isolated from a Hospitalized Patient in Khartoum, Sudan. Microbiol. Resour. Announc. 2019, 8, e00542-19.
- Dirar, M.; Bilal, N.; Ibrahim, M.E.; Hamid, M. Resistance Patterns and Phenotypic Detection of β-lactamase Enzymes among Enterobacteriaceae Isolates from Referral Hospitals in Khartoum State, Sudan. Cureus 2020, 12, e7260.
- La Scola, B.; Raoult, D. Acinetobacter baumannii in human body louse. Emerg. Infect. Dis. 2004, 10, 1671–1673.
- Heiden, S.E.; Kurz, M.S.; Bohnert, J.; Bayingana, C.; Ndoli, J.M.; Sendegeya, A.; Gahutu, J.B.; Eger, E.; Mockenhaupt, F.P.; Schaufler, K. Flies from a tertiary hospital in Rwanda carry multidrug-resistant Gram-negative pathogens including extended-spectrum beta-lactamase-producing E. coli sequence type 131. Antimicrob. Resist. Infect. Control 2020, 9, 1–4.
- Doumbia-Singare, K.; Timbo, S.; Keita, M.; Mohamed, A.A.; Guindo, B.; Soumaoro, S. Cellulite cervico-faciale au cours de la grossesse. À propos d’une série de 10 cas au Mali. Bull. De La Société De Pathol. Exot. 2014, 107, 312–316.
- Lakoh, S.; Li, L.; Sevalie, S.; Guo, X.; Adekanmbi, O.; Yang, G.; Adebayo, O.; Yi, L.; Coker, J.M.; Wang, S. Antibiotic resistance in patients with clinical features of healthcare-associated infections in an urban tertiary hospital in Sierra Leone: A cross-sectional study. Antimicrob. Resist. Infect. Control 2020, 9, 1–10.
- Mohamed, A.H.; Mohamud, M.F.Y.; Mohamud, H.A. Epidemiology and Antimicrobial Susceptibility Pattern of Uropathogens in Patients with the Community-and Hospital-Acquired Urinary Tract Infections at a Tertiary Hospital in Somalia. Jundishapur J. Microbiol. 2020, 13, e107453.
- Louni, M.; Amanzougaghene, N.; Mana, N.; Fenollar, F.; Raoult, D.; Bitam, I.; Mediannikov, O. Detection of bacterial pathogens in clade E head lice collected from Niger’s refugees in Algeria. Parasites Vectors 2018, 11, 348.
- Hamzeh, A.R.; Al Najjar, M.; Mahfoud, M. Prevalence of antibiotic resistance among Acinetobacter baumannii isolates from Aleppo, Syria. Am. J. Infect. Control 2012, 40, 776–777.
- Teicher, C.L.; Ronat, J.-B.; Fakhri, R.M.; Basel, M.; Labar, A.S.; Herard, P.; Murphy, R.A. Antimicrobial drug–resistant bacteria isolated from Syrian war–injured patients, August 2011–March 2013. Emerg. Infect. Dis. 2014, 20, 1949–1951.
- Herard, P.; Boillot, F.; Fakhri, R.M. Bone cultures from war-wounded civilians in the Middle East: A surgical prospective. Int. Orthop. 2017, 41, 1291–1294.
- Fily, F.; Ronat, J.-B.; Malou, N.; Kanapathipillai, R.; Seguin, C.; Hussein, N.; Fakhri, R.M.; Langendorf, C. Post-traumatic osteomyelitis in Middle East war-wounded civilians: Resistance to first-line antibiotics in selected bacteria over the decade 2006–2016. BMC Infect. Dis. 2019, 19, 103.
- Peretz, A.; Labay, K.; Zonis, Z.; Glikman, D. Disengagement does not apply to bacteria: A high carriage rate of antibiotic-resistant pathogens among Syrian civilians treated in Israeli hospitals. Clin. Infect. Dis. 2014, 59, 753–754.
- Heydari, F.; Mammina, C.; Koksal, F. NDM-1-producing Acinetobacter baumannii ST85 now in Turkey, including one isolate from a Syrian refugee. J. Med. Microbiol. 2015, 64, 1027–1029.
- Hasde, A.İ.; Baran, Ç.; Gümüş, F.; Kış, M.; Özçınar, E.; Çakıcı, M.; Yazıcıoğlu, L.; Kaya, B. Effect of temporary vascular shunting as a previous intervention on lower extremity arterial injury: Single center experiences in the Syrian Civil War. Turk. J. Trauma Emerg. Surg. 2019, 25, 389–395.
- Rafei, R.; Dabboussi, F.; Hamze, M.; Eveillard, M.; Lemarié, C.; Mallat, H.; Rolain, J.-M.; Joly-Guillou, M.-L.; Kempf, M. First report of blaNDM-1-producing Acinetobacter baumannii isolated in Lebanon from civilians wounded during the Syrian war. Int. J. Infect. Dis. 2014, 21, 21–23.
- Rafei, R.; Pailhoriès, H.; Hamze, M.; Eveillard, M.; Mallat, H.; Dabboussi, F.; Joly-Guillou, M.-L.; Kempf, M. Molecular epidemiology of Acinetobacter baumannii in different hospitals in Tripoli, Lebanon using blaOXA-51-like sequence based typing. BMC Microbiol. 2015, 15, 103.
- Salloum, T.; Tannous, E.; Alousi, S.; Arabaghian, H.; Rafei, R.; Hamze, M.; Tokajian, S. Genomic mapping of ST85 blaNDM-1 and blaOXA-94 producing Acinetobacter baumannii isolates from Syrian Civil War Victims. Int. J. Infect. Dis. 2018, 74, 100–108.
- Bakour, S.; Alsharapy, S.A.; Touati, A.; Rolain, J.-M. Characterization of Acinetobacter baumannii clinical isolates carrying blaOXA-23 carbapenemase and 16S rRNA methylase armA genes in Yemen. Microb. Drug Resist. 2014, 20, 604–609.
- Sutter, D.E.; Bradshaw, L.U.; Simkins, L.H.; Summers, A.M.; Atha, M.; Elwood, R.L.; Robertson, J.L.; Murray, C.K.; Wortmann, G.W.; Hospenthal, D.R. High incidence of multidrug-resistant gram-negative bacteria recovered from Afghan patients at a deployed US military hospital. Infect. Control Hosp. Epidemiol. 2011, 32, 854–860.
- Potron, A.; Munoz-Price, L.S.; Nordmann, P.; Cleary, T.; Poirel, L. Genetic features of CTX-M-15-producing Acinetobacter baumannii from Haiti. Antimicrob. Agents Chemother. 2011, 55, 5946–5948.
- Marra, A.R.; Martino, M.D.V.; Ribas, M.R.; Rodriguez-Taveras, C.; dos Santos, O.F.P. Microbiological findings from the Haiti disaster. Travel Med. Infect. Dis. 2012, 10, 157–161.
- Murphy, R.A.; Nisenbaum, L.; Labar, A.S.; Sheridan, R.L.; Ronat, J.-B.; Dilworth, K.; Pena, J.; Kilborn, E.; Teicher, C. Invasive infection and outcomes in a humanitarian surgical burn program in Haiti. World J. Surg. 2016, 40, 1550–1557.
- Chaintarli, K.; Lenglet, A.; Beauzile, B.D.; Senat-Delva, R.; Mabou, M.-M.; Martino, C.; Berthet, M.; Wong, S.; Hopman, J. High prevalence of ESBL-positive bacteria in an obstetrics emergency hospital and neonatal care unit—Haiti, 2016. Infect. Control Hosp. Epidemiol. 2018, 39, 1381–1383.
- Roy, M.A.; Arnaud, J.M.; Jasmin, P.M.; Hamner, S.; Hasan, N.A.; Colwell, R.R.; Ford, T.E. A metagenomic approach to evaluating surface water quality in Haiti. Int. J. Environ. Res. Public Health 2018, 15, 2211.
- Zarrilli, R.; Pournaras, S.; Giannouli, M.; Tsakris, A. Global evolution of multidrug-resistant Acinetobacter baumannii clonal lineages. Int. J. Antimicrob. Agents 2013, 41, 11–19.
- Yum, J.H.; Yi, K.; Lee, H.; Yong, D.; Lee, K.; Kim, J.M.; Rossolini, G.M.; Chong, Y. Molecular characterization of metallo-β-lactamase-producing Acinetobacter baumannii and Acinetobacter genomospecies 3 from Korea: Identification of two new integrons carrying the blaVIM-2 gene cassettes. J. Antimicrob. Chemother. 2002, 49, 837–840.
