The tumor microenvironment (TME) comprises a heterogenous assemblage of malignant and non-malignant cells, including infiltrating immune cells and other stromal cells, together with extracellular matrix and a variety of soluble factors. This complex and dynamic milieu strongly affects tumor differentiation, progression, immune evasion, and response to therapy, thus being an important therapeutic target. The phenotypic and functional features of the various cell types present in the TME are largely dependent on their ability to adopt different metabolic programs. Hence, modulating the metabolism of the cells in the TME, and their metabolic crosstalk, has emerged as a promising strategy in the context of anticancer therapies. Natural compounds offer an attractive tool in this respect as their multiple biological activities can potentially be harnessed to ‘(re)-educate’ TME cells towards antitumoral roles.
- tumor microenvironment
- stromal cells
- metabolism
- metabolic modulation
- natural compounds
- phytochemicals
- cancer
1. Introduction
2. Metabolic Modulation of TME Cells by Natural Compounds
2.1. Curcumin
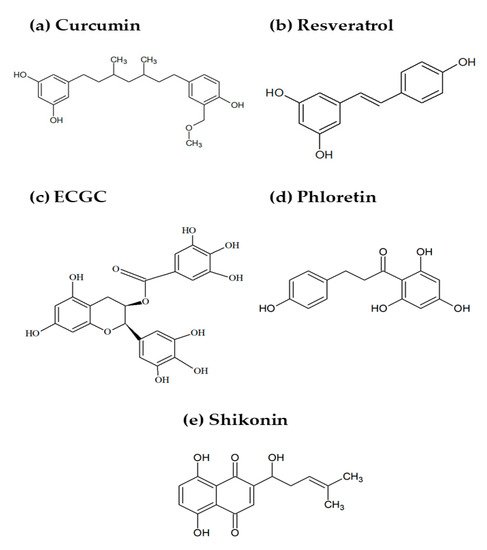
2.2. Resveratrol
2.3. Epigallocatechin Gallate
2.4. Phloretin
2.5. Shikonin
Shikonin (Figure 1e) is a naturally occurring naphthoquinone found in the root of plants from the Boraginaceae family and the first compound to be obtained from large-scale plant cell cultures [45]. It has been used in traditional Chinese medicine for centuries and shown to possess several therapeutic properties, including antimicrobial, wound healing, anti-inflammatory, antioxidant, and anticancer activities [46]. The potential of shikonin and its derivatives in cancer treatment has received increasing attention in recent years, mainly due to its wide spectrum antitumor effects [47]. Repression of glycolysis through specific inhibition of PKM2, the enzyme catalyzing the conversion of phosphoenolpyruvate to pyruvate, is a key mechanism in shikonin’s antitumor activity, as demonstrated in a variety of tumor cells [48][49][50].
Recent data clearly showed that shikonin-mediated metabolic effects impacted the TME by repolarizing TAM and synergizing with PD-1 blockage (mediated by JQ1), thus enhancing the immune response [51]. In that work, mannosylated lactoferrin nanoparticles were loaded with shikonin (1 μM) and JQ1 (3 μM), for targeted codelivery to colon cancer cells (CT26) and TAM. The bioactive NPs reduced lactate production in cancer cells (in association with PKM2-mediated glycolytic inhibition), and skewed macrophages towards a pro- inflammatory phenotype, characterized by higher production of TNF-α and lower secretion of TGF-β. Furthermore, treatment of CT26-tumor-bearing mice with the shikonin/JQ1-loaded nanosystem efficiently decreased tumor growth, suppressed glucose metabolism (as seen by reduced levels of lactate, PKM2 and HIF-1α in the tumor tissue), downregulated intratumoral PD-L1 expression, and remodeled the TME’s immune configuration (e.g., promotion of dendritic cell maturation and CD8+ T cell infiltration, as well as suppression of Treg). Overall, the synergism between metabolic reprogramming and regulation of immune responses was shown to improve antitumor treatment efficacy.
This entry is adapted from the peer-reviewed paper 10.3390/molecules26123494
References
- Labani-Motlagh, A.; Ashja-Mahdavi, M.; Loskog, A. The Tumor Microenvironment: A Milieu Hindering and Obstructing Antitumor Immune Responses. Front. Immunol. 2020, 11, 940.
- Wu, T.; Dai, Y. Tumor microenvironment and therapeutic response. Cancer Lett. 2017, 387, 61–68.
- Hanahan, D.; Weinberg, R.A. Hallmarks of Cancer: The Next Generation. Cell 2011, 144, 646–674.
- Fouad, Y.A.; Aanei, C. Revisiting the hallmarks of cancer. Am. J. Cancer Res. 2017, 7, 1016–1036.
- Romero-Garcia, S.; Lopez-Gonzalez, J.S.; Báez-Viveros, J.L.; Aguilar-Cazares, D.; Prado-Garcia, H. Tumor cell metabolism: An integral view. Cancer Biol. Ther. 2011, 12, 939–948.
- Dias, A.S.; Almeida, C.R.; Helguero, L.; Duarte, I.F. Metabolic crosstalk in the breast cancer microenvironment. Eur. J. Cancer 2019, 121, 154–171.
- De La Cruz-López, K.G.; Castro-Muñoz, L.J.; Reyes-Hernández, D.O.; García-Carrancá, A.; Manzo-Merino, J. Lactate in the Regulation of Tumor Microenvironment and Therapeutic Approaches. Front. Oncol. 2019, 9, 1143.
- Xing, Y.; Zhao, S.; Zhou, B.P.; Mi, J. Metabolic reprogramming of the tumour microenvironment. FEBS J. 2015, 282, 3892–3898.
- Zyad, A.; Leouifoudi, I.; Tilaoui, M.; Mouse, H.A.; Khouchani, M.; Jaafari, A. Natural Products as Cytotoxic Agents in Chemotherapy against Cancer. Cytotoxicity 2018.
- Liskova, A.; Samec, M.; Koklesova, L.; Brockmueller, A.; Zhai, K.; Abdellatif, B.; Siddiqui, M.; Biringer, K.; Kudela, E.; Pec, M.; et al. Flavonoids as an effective sensitizer for anti-cancer therapy: Insights into multi-faceted mechanisms and applicability towards individualized patient profiles. EPMA J. 2021, 17, 1–22.
- Park, S.; Surh, Y. Modulation of tumor microenvironment by chemopreventive natural products. Ann. N. Y. Acad. Sci. 2017, 1401, 65–74.
- Zubair, H.; Khan, M.A.; Anand, S.; Srivastava, S.K.; Singh, S.; Singh, A.P. Modulation of the tumor microenvironment by natural agents: Implications for cancer prevention and therapy. Semin. Cancer Biol. 2020.
- Pan, P.; Huang, Y.-W.; Oshima, K.; Yearsley, M.; Zhang, J.; Arnold, M.; Yu, J.; Wang, L.-S. The immunomodulatory potential of natural compounds in tumor-bearing mice and humans. Crit. Rev. Food Sci. Nutr. 2019, 59, 992–1007.
- Focaccetti, C.; Izzi, V.; Benvenuto, M.; Fazi, S.; Ciuffa, S.; Giganti, M.G.; Potenza, V.; Manzari, V.; Modesti, A.; Bei, R. Polyphenols as Immunomodulatory Compounds in the Tumor Microenvironment: Friends or Foes? Int. J. Mol. Sci. 2019, 20, 1714.
- Samec, M.; Liskova, A.; Koklesova, L.; Samuel, S.M.; Zhai, K.; Buhrmann, C.; Varghese, E.; Abotaleb, M.; Qaradakhi, T.; Zulli, A.; et al. Flavonoids against the Warburg phenotype—concepts of predictive, preventive and personalised medicine to cut the Gordian knot of cancer cell metabolism. EPMA J. 2020, 11, 377–398.
- Guerra, A.R.; Duarte, M.F.; Duarte, I.F. Targeting Tumor Metabolism with Plant-Derived Natural Products: Emerging Trends in Cancer Therapy. J. Agric. Food Chem. 2018, 66, 10663–10685.
- Giordano, A.; Tommonaro, G. Curcumin and Cancer. Nutrients 2019, 11, 2376.
- Li, H.; Du, H.; Zhang, G.; Wu, Y.; Qiu, P.; Liu, J.; Guo, J.; Liu, X.; Sun, L.; Du, B.; et al. Curcumin plays a synergistic role in combination with HSV-TK/GCV in inhibiting growth of murine B16 melanoma cells and melanoma xenografts. PeerJ 2019, 7, e7760.
- Šudomová, M.; Hassan, S. Nutraceutical Curcumin with Promising Protection against Herpesvirus Infections and Their Associated Inflammation: Mechanisms and Pathways. Microorganisms 2021, 9, 292.
- Vishvakarma, N.K. Novel antitumor mechanisms of curcumin: Implication of altered tumor metabolism, reconstituted tumor microenvironment and augmented myelopoiesis. Phytochem. Rev. 2014, 13, 717–724.
- Wang, K.; Fan, H.; Chen, Q.; Ma, G.; Zhu, M.; Zhang, X.; Zhang, Y.; Yu, J. Curcumin inhibits aerobic glycolysis and induces mitochondrial-mediated apoptosis through hexokinase II in human colorectal cancer cells in vitro. Anti-Cancer Drugs 2015, 26, 15–24.
- Siddiqui, F.A.; Prakasam, G.; Chattopadhyay, S.; Rehman, A.U.; Padder, R.A.; Ansari, M.A.; Irshad, R.; Mangalhara, K.; Bamezai, R.N.K.; Husain, M.; et al. Curcumin decreases Warburg effect in cancer cells by down-regulating pyruvate kinase M2 via mTOR-HIF1α inhibition. Sci. Rep. 2018, 8, 1–9.
- Yang, R.; Fang, X.-L.; Zhen, Q.; Chen, Q.-Y.; Feng, C. Mitochondrial targeting nano-curcumin for attenuation on PKM2 and FASN. Colloids Surf. B Biointerfaces 2019, 182, 110405.
- Soni, V.K.; Shukla, D.; Kumar, A.; Vishvakarma, N.K. Curcumin circumvent lactate-induced chemoresistance in hepatic cancer cells through modulation of hydroxycarboxylic acid receptor-1. Int. J. Biochem. Cell Biol. 2020, 123, 105752.
- Fan, H.; Tian, W.; Ma, X. Curcumin induces apoptosis of HepG2 cells via inhibiting fatty acid synthase. Target. Oncol. 2014, 9, 279–286.
- Younesian, O.; Kazerouni, F.; Dehghan-Nayeri, N.; Omrani, D.; Rahimipour, A.; Shanaki, M.; Kalkhoran, M.R.; Cheshmi, F. Effect of Curcumin on Fatty Acid Synthase Expression and Enzyme Activity in Breast Cancer Cell Line SKBR3. Int. J. Cancer Manag. 2017, 10.
- Bianchi, G.; Ravera, S.; Traverso, C.; Amaro, A.; Piaggio, F.; Emionite, L.; Bachetti, T.; Pfeffer, U.; Raffaghello, L. Curcumin induces a fatal energetic impairment in tumor cells in vitro and in vivo by inhibiting ATP-synthase activity. Carcinogenesis 2018, 39, 1141–1150.
- Sordillo, P.P.; Helson, L. Curcumin and cancer stem cells: Curcumin has asymmetrical effects on cancer and normal stem cells. Anticancer Res. 2015, 35, 599–614.
- Huang, Y.-T.; Lin, Y.-W.; Chiu, H.-M.; Chiang, B.-H. Curcumin Induces Apoptosis of Colorectal Cancer Stem Cells by Coupling with CD44 Marker. J. Agric. Food Chem. 2016, 64, 2247–2253.
- Nakagawa, K.; Zingg, J.-M.; Kim, S.H.; Thomas, M.J.; Dolnikowski, G.G.; Azzi, A.; Miyazawa, T.; Meydani, M. Differential cellular uptake and metabolism of curcuminoids in monocytes/macrophages: Regulatory effects on lipid accumulation. Br. J. Nutr. 2014, 112, 8–14.
- Zingg, J.-M.; Hasan, S.T.; Cowan, D.; Ricciarelli, R.; Azzi, A.; Meydani, M. Regulatory effects of curcumin on lipid accumulation in monocytes/macrophages. J. Cell. Biochem. 2012, 113, 833–840.
- Han, Y.; Jo, H.; Cho, J.H.; Dhanasekaran, D.N.; Song, Y.S. Resveratrol as a Tumor-Suppressive Nutraceutical Modulating Tumor Microenvironment and Malignant Behaviors of Cancer. Int. J. Mol. Sci. 2019, 20, 925.
- Jiang, Z.; Chen, K.; Cheng, L.; Yan, B.; Qian, W.; Cao, J.; Liang, C.; Wu, E.; Ma, Q.; Yang, W. Resveratrol and cancer treatment: Updates. Ann. N. Y. Acad. Sci. 2017, 1403, 59–69.
- Brockmueller, A.; Sameri, S.; Liskova, A.; Zhai, K.; Varghese, E.; Samuel, S.M.; Büsselberg, D.; Kubatka, P.; Shakibaei, M. Resveratrol’s Anti-Cancer Effects through the Modulation of Tumor Glucose Metabolism. Cancers 2021, 13, 188.
- Jia, L.; Gao, Y.; Zhou, T.; Zhao, X.-L.; Hu, H.-Y.; Chen, D.-W.; Qiao, M.-X. Enhanced response to PD-L1 silencing by modulation of TME via balancing glucose metabolism and robust co-delivery of siRNA/Resveratrol with dual-responsive polyplexes. Biomaterials 2021, 271, 120711.
- Craveiro, M.; Cretenet, G.; Mongellaz, C.; Matias, M.I.; Caron, O.; De Lima, M.C.P.; Zimmermann, V.S.; Solary, E.; Dardalhon, V.; Dulić, V.; et al. Resveratrol stimulates the metabolic reprogramming of human CD4+T cells to enhance effector function. Sci. Signal. 2017, 10, eaal3024.
- Nakaya, M.; Xiao, Y.; Zhou, X.; Chang, J.-H.; Chang, M.; Cheng, X.; Blonska, M.; Lin, X.; Sun, S.-C. Inflammatory T Cell Responses Rely on Amino Acid Transporter ASCT2 Facilitation of Glutamine Uptake and mTORC1 Kinase Activation. Immunity 2014, 40, 692–705.
- Cheng, Z.; Zhang, Z.; Han, Y.; Wang, J.; Wang, Y.; Chen, X.; Shao, Y.; Cheng, Y.; Zhou, W.; Lu, X.; et al. A review on anti-cancer effect of green tea catechins. J. Funct. Foods 2020, 74, 104172.
- Wei, R.; Mao, L.; Xu, P.; Zheng, X.; Hackman, R.M.; Mackenzie, G.G.; Wang, Y. Suppressing glucose metabolism with epigallocatechin-3-gallate (EGCG) reduces breast cancer cell growth in preclinical models. Food Funct. 2018, 9, 5682–5696.
- Zhang, Z.; Zhang, S.; Yang, J.; Yi, P.; Xu, P.; Yi, M.; Peng, W. Integrated transcriptomic and metabolomic analyses to characterize the anti-cancer effects of (−)-epigallocatechin-3-gallate in human colon cancer cells. Toxicol. Appl. Pharmacol. 2020, 401, 115100.
- Chu, K.O.; Chan, K.P.; Chan, S.O.; Ng, T.K.; Jhanji, V.; Wang, C.-C.; Pang, C.P. Metabolomics of Green-Tea Catechins on Vascular-Endothelial-Growth-Factor-Stimulated Human-Endothelial-Cell Survival. J. Agric. Food Chem. 2018, 66, 12866–12875.
- Wu, K.-H.; Ho, C.-T.; Chen, Z.-F.; Chen, L.-C.; Whang-Peng, J.; Lin, T.-N.; Ho, Y.-S. The apple polyphenol phloretin inhibits breast cancer cell migration and proliferation via inhibition of signals by type 2 glucose transporter. J. Food Drug Anal. 2018, 26, 221–231.
- Lin, S.-T.; Tu, S.-H.; Yang, P.-S.; Hsu, S.-P.; Lee, W.-H.; Ho, C.-T.; Wu, C.-H.; Lai, Y.-H.; Chen, M.-Y.; Chen, L.-C. Apple Polyphenol Phloretin Inhibits Colorectal Cancer Cell Growth via Inhibition of the Type 2 Glucose Transporter and Activation of p53-Mediated Signaling. J. Agric. Food Chem. 2016, 64, 6826–6837.
- Patel, B.B.; Ackerstaff, E.; Serganova, I.S.; Kerrigan, J.E.; Blasberg, R.G.; Koutcher, J.A.; Banerjee, D. Tumor stroma interaction is mediated by monocarboxylate metabolism. Exp. Cell Res. 2017, 352, 20–33.
- Přenosil, J.E.; Kut, Ö.M.; Dunn, I.J.; Heinzle, E. Biocatalysis, 2. Immobilized Biocatalysts. Ullmann’s Encycl. Ind. Chem. 2009, 5, 477–527.
- Guo, C.; He, J.; Song, X.; Tan, L.; Wang, M.; Jiang, P.; Li, Y.; Cao, Z.; Peng, C. Pharmacological properties and derivatives of shikonin—A review in recent years. Pharmacol. Res. 2019, 149.
- Boulos, J.C.; Rahama, M.; Hegazy, M.-E.F.; Efferth, T. Shikonin derivatives for cancer prevention and therapy. Cancer Lett. 2019, 459, 248–267.
- Zhao, X.; Zhu, Y.; Hu, J.; Jiang, L.; Li, L.; Jia, S.; Zen, K. Shikonin Inhibits Tumor Growth in Mice by Suppressing Pyruvate Kinase M2-mediated Aerobic Glycolysis. Sci. Rep. 2018, 8, 1–8.
- Liu, B.; Jin, J.; Zhang, Z.; Zuo, L.; Jiang, M.; Xie, C. Shikonin exerts antitumor activity by causing mitochondrial dysfunction in hepatocellular carcinoma through PKM2–AMPK–PGC1α signaling pathway. Biochem. Cell Biol. 2019, 97, 397–405.
- Tao, T.; Su, Q.; Xu, S.; Deng, J.; Zhou, S.; Zhuang, Y.; Huang, Y.; He, C.; He, S.; Peng, M.; et al. Down-regulation of PKM2 decreases FASN expression in bladder cancer cells through AKT/mTOR/SREBP-1c axis. J. Cell. Physiol. 2019, 234, 3088–3104.
- Wang, H.; Tang, Y.; Fang, Y.; Zhang, M.; Wang, H.; He, Z.; Wang, B.; Xu, Q.; Huang, Y. Reprogramming Tumor Immune Microenvironment (TIME) and Metabolism via Biomimetic Targeting Codelivery of Shikonin/JQ1. Nano Lett. 2019, 19, 2935–2944.
