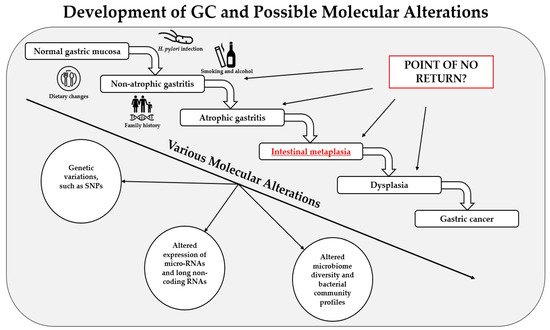
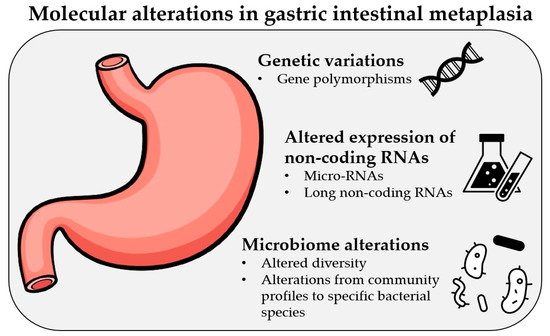
Gastric cancer (GC) remains one of the most common causes of mortality worldwide. Intestinal metaplasia (IM) is one of the preneoplastic gastric lesions and is considered an essential predisposing factor in GC development. Here we present a review of recent most relevant papers to summarize major findings on the molecular alterations in gastric IM. The latest progress in novel diagnostic methods allows scientists to identify various types of molecular alterations in IM, such as polymorphisms in various genes, changes in the expression of micro-RNAs and long noncoding RNAs, and altered microbiome profiles. The results have shown that some of these alterations have strong associations with IM and a potential to be used for screening, treatment, and prognostic purposes; however, one of the most important limiting factors is the inhomogeneity of the studies. Therefore, further large-scale studies and clinical trials with standardized methods designed by multicenter consortiums are needed. As of today, various molecular alterations in IM could become a part of personalized medicine in the near future, which would help us deliver a personalized approach for each patient and identify those at risk of progression to GC.
- intestinal metaplasia
- molecular alterations
- genetic variations
- micro-RNAs
- microbiome
1. Introduction
2. Genetic Variations in IM
3. MicroRNAs in IM
4. Long-Non-Coding RNAs in IM
5. Microbiome Alterations in IM
6. Conclusions
This entry is adapted from the peer-reviewed paper 10.3390/ijms22115758
References
- Karimi, P.; Islami, F.; Anandasabapathy, S.; Freedman, N.D.; Kamangar, F. Gastric Cancer: Descriptive Epidemiology, Risk Factors, Screening, and Prevention. Cancer Epidemiol. Biomark. Prev. 2014, 23, 700–713.
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2021.
- Smyth, E.C.; Nilsson, M.; Grabsch, H.I.; van Grieken, N.C.; Lordick, F. Gastric cancer. Lancet (Lond. Engl.) 2020, 396, 635–648.
- Oue, N.; Sentani, K.; Sakamoto, N.; Uraoka, N.; Yasui, W. Molecular carcinogenesis of gastric cancer: Lauren classification, mucin phenotype expression, and cancer stem cells. Int. J. Clin. Oncol. 2019, 24, 771–778.
- Berlth, F.; Bollschweiler, E.; Drebber, U.; Hoelscher, A.H.; Moenig, S. Pathohistological classification systems in gastric cancer: Diagnostic relevance and prognostic value. World J. Gastroenterol. 2014, 20, 5679–5684.
- Malfertheiner, P.; Megraud, F.; O’morain, C.A.; Gisbert, J.P.; Kuipers, E.J.; Axon, A.T.; Bazzoli, F.; Gasbarrini, A.; Atherton, J.; Graham, D.Y.; et al. Management of Helicobacter pylori infection-the Maastricht V/Florence Consensus Report. Gut 2017, 66, 6–30.
- Goh, K.-L.; Chan, W.-K.; Shiota, S.; Yamaoka, Y. Epidemiology of Helicobacter pylori Infection and Public Health Implications. Helicobacter 2011, 16, 1–9.
- Sjomina, O.; Pavlova, J.; Niv, Y.; Leja, M. Epidemiology of Helicobacter pylori infection. Helicobacter 2018, 23, e12514.
- Jonaitis, L.; Pellicano, R.; Kupcinskas, L. Helicobacter pylori and nonmalignant upper gastrointestinal diseases. Helicobacter 2018, 23 (Suppl. 1), e12522.
- Moss, S.F. The Clinical Evidence Linking Helicobacter pylori to Gastric Cancer. Cell. Mol. Gastroenterol. Hepatol. 2017, 3, 183–191.
- Takahashi-Kanemitsu, A.; Knight, C.T.; Hatakeyama, M. Molecular anatomy and pathogenic actions of Helicobacter pylori CagA that underpin gastric carcinogenesis. Cell. Mol. Immunol. 2020, 17, 50–63.
- Tohidpour, A. CagA-mediated pathogenesis of Helicobacter pylori. Microb. Pathog. 2016, 93, 44–55.
- Leung, W.K.; Lin, S.-R.; Ching, J.Y.L.; To, K.-F.; Ng, E.K.W.; Chan, F.K.L.; Lau, J.Y.W.; Sung, J.J.Y. Factors predicting progression of gastric intestinal metaplasia: Results of a randomised trial on Helicobacter pylori eradication. Gut 2004, 53, 1244–1249.
- Correa, P.; Piazuelo, M.B. The gastric precancerous cascade. J. Dig. Dis. 2011, 13, 2–9.
- Correa, P.; Houghton, J. Carcinogenesis of Helicobacter pylori. Gastroenterology 2007, 133, 659–672.
- Correa, P.; Piazuelo, B.M.; Wilson, K.T. Pathology of Gastric Intestinal Metaplasia: Clinical Implications. Am. J. Gastroenterol. 2010, 105, 493–498.
- Kinoshita, H.; Hayakawa, Y.; Koike, K. Metaplasia in the Stomach—Precursor of Gastric Cancer? Int. J. Mol. Sci. 2017, 18, 2063.
- Hwang, Y.-J.; Kim, N.; Lee, H.S.; Lee, J.B.; Choi, Y.J.; Yoon, H.; Shin, C.M.; Park, Y.S.; Lee, D.H. Reversibility of atrophic gastritis and intestinal metaplasia after Helicobacter pylori eradication-a prospective study for up to 10 years. Aliment. Pharmacol. Ther. 2017, 47, 380–390.
- Walker, M.M. Is intestinal metaplasia of the stomach reversible? Gut 2003, 52, 1–4.
- Lam, S.K.; Lau, G. Novel treatment for gastric intestinal metaplasia, a precursor to cancer. JGH Open 2020, 4, 569–573.
- Sánchez Cuén, J.A.; Irineo Cabrales, A.B.; Bernal Magaña, G.; Peraza Garay, F. Regression of gastric intestinal metaplasia after the eradication of Helicobacter pylori infection in a hospital in Mexico. Rev. Esp. Enferm. Dig. Organo Soc. Esp. Patol. Dig. 2016, 108, 770–775.
- Liu, K.S.-H.; Wong, I.O.-L.; Leung, W.K. Helicobacter pylori associated gastric intestinal metaplasia: Treatment and surveillance. World J. Gastroenterol. 2016, 22, 1311–1320.
- Huang, R.J.; Choi, A.Y.; Truong, C.D.; Yeh, M.M.; Hwang, J.H. Diagnosis and Management of Gastric Intestinal Metaplasia: Current Status and Future Directions. Gut Liver 2019, 13, 596–603.
- Jencks, D.S.; Adam, J.D.; Borum, M.L.; Koh, J.M.; Stephen, S.; Doman, D.B. Overview of Current Concepts in Gastric Intestinal Metaplasia and Gastric Cancer. Gastroenterol. Hepatol. 2018, 14, 92–101.
- Gómez Zuleta, M.A.; Torres, K.E.; Falduto, M.T.; Magnuson, S.R. Identification of Blood Biomarkers for Detecting Premalignant Lesions and Gastric Cancer. Rev. Colomb. Gastroenterol. Scieloco 2017, 32, 7–19.
- Leja, M.; Kupcinskas, L.; Funka, K.; Sudraba, A.; Jonaitis, L.; Ivanauskas, A.; Janciauskas, D.; Kiudelis, G.; Chiu, H.M.; Lin, J.T. The validity of a biomarker method for indirect detection of gastric mucosal atrophy versus standard histopathology. Dig. Dis. Sci. 2009, 54, 2377–2384.
- Suh, Y.-S.; Lee, H.-J.; Jung, E.-J.; Kim, M.-A.; Nam, K.T.; Goldenring, J.R.; Yang, H.-K.; Kim, W.H. The Combined Expression of Metaplasia Biomarkers Predicts the Prognosis Of Gastric Cancer. Ann. Surg. Oncol. 2011, 19, 1240–1249.
- Novelli, G.; Ciccacci, C.; Borgiani, P.; Amati, M.P.; Abadie, E. Genetic tests and genomic biomarkers: Regulation, qualification and validation. Clin. Cases Miner. Bone Metab. 2008, 5, 149–154.
- Goetz, L.H.; Schork, N.J. Personalized medicine: Motivation, challenges, and progress. Fertil. Steril. 2018, 109, 952–963.
- Elemento, O. The future of precision medicine: Towards a more predictive personalized medicine. Emerg. Top. Life Sci. 2020, 4, 175–177.
- Jonaitis, P.; Kiudelis, V.; Streleckiene, G.; Gedgaudas, R.; Skieceviciene, J.; Kupcinskas, J. Novel Biomarkers in the Diagnosis of Benign and Malignant GI Diseases. Dig. Dis. 2021.
- Seeneevassen, L.; Bessède, E.; Mégraud, F.; Lehours, P.; Dubus, P.; Varon, C. Gastric Cancer: Advances in Carcinogenesis Research and New Therapeutic Strategies. Int. J. Mol. Sci. 2021, 22, 3418.
- Dargiene, G.; Streleckiene, G.; Skieceviciene, J.; Leja, M.; Link, A.; Wex, T.; Kupcinskas, L.; Malfertheiner, P.; Kupcinskas, J. TLR1 and PRKAA1 Gene Polymorphisms in the Development of Atrophic Gastritis and Gastric Cancer. J. Gastrointest. Liver Dis. 2018, 27, 363–369.
- Kupcinskas, J.; Wex, T.; Link, A.; Leja, M.; Bruzaite, I.; Steponaitiene, R.; Juzenas, S.; Gyvyte, U.; Ivanauskas, A.; Ancans, G.; et al. Gene Polymorphisms of Micrornas in Helicobacter pylori-Induced High Risk Atrophic Gastritis and Gastric Cancer. PLoS ONE 2014, 9, e87467.
- Kupcinskas, J.; Wex, T.; Bornschein, J.; Selgrad, M.; Leja, M.; Juozaitytė, E.; Kiudelis, G.; Jonaitis, L.; Malfertheiner, P. Lack of association between gene polymorphisms of Angiotensin converting enzyme, Nod-like receptor 1, Toll-like receptor 4, FAS/FASL and the presence of Helicobacter pylori-induced premalignant gastric lesions and gastric cancer in Caucasians. BMC Med. Genet. 2011, 12, 112.
- Kupcinskas, J.; Wex, T.; Link, A.; Bartuseviciute, R.; Dedelaite, M.; Kevalaite, G.; Leja, M.; Skieceviciene, J.; Kiudelis, G.; Jonaitis, L.; et al. PSCA and MUC1 gene polymorphisms are associated with gastric cancer and pre-malignant gastric conditions [corrected]. Anticancer Res. 2014, 34, 7167–7175.
- Petkevicius, V.; Salteniene, V.; Juzenas, S.; Wex, T.; Link, A.; Leja, M.; Steponaitienė, R.; Skiecevičienė, J.; Kupcinskas, L.; Jonaitis, L.; et al. Polymorphisms of microRNA target genes IL12B, INSR, CCND1 and IL10 in gastric cancer. World J. Gastroenterol. 2017, 23, 3480–3487.
- Kupcinskas, L.; Wex, T.; Kupcinskas, J.; Leja, M.; Ivanauskas, A.; Jonaitis, L.V.; Janciauskas, D.; Kiudelis, G.; Funka, K.; Sudraba, A.; et al. Interleukin-1B and interleukin-1 receptor antagonist gene polymorphisms are not associated with premalignant gastric conditions: A combined haplotype analysis. Eur. J. Gastroenterol. Hepatol. 2010, 22, 1189–1195.
- Nelson, M.R.; Marnellos, G.; Kämmerer, S.; Hoyal, C.R.; Shi, M.M.; Cantor, C.R.; Braun, A. Large-Scale Validation of Single Nucleotide Polymorphisms in Gene Regions. Genome Res. 2004, 14, 1664–1668.
- Köberle, B.; Koch, B.; Fischer, B.M.; Hartwig, A. Single nucleotide polymorphisms in DNA repair genes and putative cancer risk. Arch. Toxicol. 2016, 90, 2369–2388.
- Matsuda, K. PCR-Based Detection Methods for Single-Nucleotide Polymorphism or Mutation: Real-Time PCR and Its Substantial Contribution Toward Technological Refinement. Adv. Clin. Chem. 2017, 80, 45–72.
- Nogales, A.; LDeDiego, M. Host Single Nucleotide Polymorphisms Modulating Influenza A Virus Disease in Humans. Pathogens 2019, 8, 168.
- Erichsen, H.C.; Chanock, S.J. SNPs in cancer research and treatment. Br. J. Cancer 2004, 90, 747–751.
- Fagny, M.; Platig, J.; Kuijjer, M.; Lin, X.; Quackenbush, J. Nongenic cancer-risk SNPs affect oncogenes, tumour-suppressor genes, and immune function. Br. J. Cancer 2020, 122, 569–577.
- Kara, B.; Akkiz, H.; Doran, F.; Bayram, S.; Erken, E.; Gumurdullu, Y.; Sandikci, M.; Akkız, H. The significance of E266K polymorphism in the NOD1 gene on Helicobacter pylori infection: An effective force on pathogenesis? Clin. Exp. Med. 2009, 10, 107–112.
- Nieuwenburg, S.A.V.; Mommersteeg, M.C.; Eikenboom, E.L.; Yu, B.; den Hollander, W.J.; Holster, I.L.; den Hoed, C.M.; Capelle, L.G.; Tang, T.J.; Anten, M.P.; et al. Factors associated with the progression of gastric intestinal metaplasia: A multicenter, prospective cohort study. Endosc. Int. Open 2021, 9, E297–E305.
- Leung, W.K.; Chan, M.C.; To, K.-F.; Man, E.P.S.; Ng, E.K.W.; Chu, E.S.H.; Lau, J.Y.W.; Lin, S.-R.; Sung, J.J.Y. H. pylori Genotypes and Cytokine Gene Polymorphisms Influence the Development of Gastric Intestinal Metaplasia in a Chinese Population. Am. J. Gastroenterol. 2006, 101, 714–720.
- Wang, Y.-M.; Li, Z.-X.; Tang, F.-B.; Zhang, Y.; Zhou, T.; Zhang, L.; Ma, J.-L.; You, W.-C.; Pan, K.-F. Association of genetic polymorphisms of interleukins with gastric cancer and precancerous gastric lesions in a high-risk Chinese population. Tumor Biol. 2015, 37, 2233–2242.
- Mansour-Ghanaei, F.; Joukar, F.; Baghaee, M.; Sepehrimanesh, M.; Hojati, A. Only serum pepsinogen I and pepsinogen I/II ratio are specific and sensitive biomarkers for screening of gastric cancer. Biomol. Concepts 2019, 10, 82–90.
- Lee, S.-Y. Endoscopic gastritis, serum pepsinogen assay, and Helicobacter pylori infection. Korean J. Intern. Med. 2016, 31, 835–844.
- Correa, P. Serum Pepsinogens in Gastric Cancer Screening. Dig. Dis. Sci. 2010, 55, 2123–2125.
- Con, S.A.; Con-Wong, R.; Con-Chin, G.R.; Con-Chin, V.G.; Takeuchi, H.; Valerín, A.L.; Echandi, G.; Mena, F.; Brenes, F.; Yasuda, N.; et al. Serum Pepsinogen Levels, Helicobacter pylori CagA Status, and Cytokine Gene Polymorphisms Associated with Gastric Premalignant Lesions in Costa Rica. Cancer Epidemiol. Biomark. Prev. 2007, 16, 2631–2636.
- Zabaleta, J.; Camargo, M.C.; Ritchie, M.D.; Piazuelo, M.B.; Sierra, R.A.; Turner, S.D.; Delgado, A.; Fontham, E.T.H.; Schneider, B.G.; Correa, P.; et al. Association of haplotypes of inflammation-related genes with gastric preneoplastic lesions in African Americans and Caucasians. Int. J. Cancer 2011, 128, 668–675.
- Kato, I.; Canzian, F.; Franceschi, S.; Plummer, M.; Van Doorn, L.-J.; Lu, Y.; Gioia-Patricola, L.; Vivas, J.; Lopez, G.; Severson, R.K.; et al. Genetic polymorphisms in anti-inflammatory cytokine signaling and the prevalence of gastric precancerous lesions in Venezuela. Cancer Causes Control. 2006, 17, 1183–1191.
- Li, Z.-W.; Wu, Y.; Sun, Y.; Liu, L.-Y.; Tian, M.-M.; Feng, G.-S.; You, W.-C.; Li, J.-Y. Inflammatory cytokine gene polymorphisms increase the risk of atrophic gastritis and intestinal metaplasia. World J. Gastroenterol. 2010, 16, 1788–1794.
- Negovan, A.; Iancu, M.; Fülöp, E.; Bănescu, C. Helicobacter pylori and cytokine gene variants as predictors of premalignant gastric lesions. World J. Gastroenterol. 2019, 25, 4105–4124.
- Jiang, F.; Shen, X.B. miRNA and mRNA expression profiles in gastric cancer patients and the relationship with circRNA. Neoplasma 2019, 66, 879–886.
- Jung, J.; Jeong, S.; Jeong, H.; Oh, H.E.; Choi, J.-W.; Lee, E.S.; Kim, Y.-S.; Kwak, Y.; Kim, W.H.; Lee, J.-H. Increased HOXC6 mRNA expression is a novel biomarker of gastric cancer. PLoS ONE 2020, 15, e0236811.
- Bhat, S.A.; Mir, M.U.R.; Majid, S.; Hassan, T.; Rehman, M.U.; Kuchy, S. Diagnostic utility of glycosyltransferase mRNA expression in gastric cancer. Hematol. Stem Cell Ther. 2018, 11, 158–168.
- Joo, M.K.; Park, J.-J.; Chun, H.J. Impact of homeobox genes in gastrointestinal cancer. World J. Gastroenterol. 2016, 22, 8247–8256.
- Mizoshita, T.; Inada, K.-I.; Tsukamoto, T.; Kodera, Y.; Yamamura, Y.; Hirai, T.; Kato, T.; Joh, T.; Itoh, M.; Tatematsu, M. Expression of Cdx1 and Cdx2 mRNAs and relevance of this expression to differentiation in human gastrointestinal mucosa—with special emphasis on participation in intestinal metaplasia of the human stomach. Gastric Cancer 2001, 4, 185–191.
- Lin, R.; Li, C.; Liu, Z.; Wu, R.; Lu, J. Genome-wide DNA methylation profiling identifies epigenetic signatures of gastric cardiac intestinal metaplasia. J. Transl. Med. 2020, 18, 1–11.
- Duarte, M.; Babeto, E.; Leite, K.; Miyazaki, K.; Borim, A.; Rahal, P.; Silva, A. Expression of TERT in precancerous gastric lesions compared to gastric cancer. Braz. J. Med. Biol. Res. 2011, 44, 100–104.
- Tatemichi, M.; Nomura, S.; Ogura, T.; Sone, H.; Nagata, H.; Esumi, H. Mutagenic activation of environmental carcinogens by microsomes of gastric mucosa with intestinal metaplasia. Cancer Res. 1999, 59, 3893–3898.
- Mattick, J.S.; Makunin, I.V. Non-coding RNA. Hum. Mol. Genet. 2006, 15 (Suppl. 1), R17–R29.
- Panni, S.; Lovering, R.C.; Porras, P.; Orchard, S. Non-coding RNA regulatory networks. Biochim. et Biophys. Acta (BBA) Bioenerg. 2020, 1863, 194417.
- Zhang, P.; Wu, W.; Chen, Q.; Chen, M. Non-Coding RNAs and their Integrated Networks. J. Integr. Bioinform. 2019, 16.
- Wang, J.; Zhu, S.; Meng, N.; He, Y.; Lu, R.; Yan, G.-R. ncRNA-Encoded Peptides or Proteins and Cancer. Mol. Ther. 2019, 27, 1718–1725.
- Fassan, M.; Croce, C.M.; Rugge, M. miRNAs in precancerous lesions of the gastrointestinal tract. World J. Gastroenterol. 2011, 17, 5231–5239.
- Liu, N.; Wang, Z.-Z.; Zhao, M.; Zhang, Y.; Chen, N.-H. Role of non-coding RNA in the pathogenesis of depression. Gene 2020, 735, 144276.
- Link, A.; Kupcinskas, J.; Wex, T.; Malfertheiner, P. Macro-Role of MicroRNA in Gastric Cancer. Dig. Dis. 2012, 30, 255–267.
- Link, A.; Kupcinskas, J. MicroRNAs as non-invasive diagnostic biomarkers for gastric cancer: Current insights and future perspectives. World J. Gastroenterol. 2018, 24, 3313–3329.
- Nishizawa, T.; Suzuki, H. The Role of microRNA in Gastric Malignancy. Int. J. Mol. Sci. 2013, 14, 9487–9496.
- Cortés-Márquez, A.C.; Mendoza-Elizalde, S.; Arenas-Huertero, F.; Trillo-Tinoco, J.; Valencia-Mayoral, P.; Consuelo-Sánchez, A.; Zarate-Franco, J.; Dionicio-Avendaño, A.R.; Herrera-Esquivel, J.D.J.; Recinos-Carrera, E.G.; et al. Differential expression of miRNA-146a and miRNA-155 in gastritis induced by Helicobacter pylori infection in paediatric patients, adults, and an animal model. BMC Infect. Dis. 2018, 18, 463.
- Li, T.; Guo, H.; Li, H.; Jiang, Y.; Zhuang, K.; Lei, C.; Wu, J.; Zhou, H.; Zhu, R.; Zhao, X.; et al. MicroRNA-92a-1–5p increases CDX2 by targeting FOXD1 in bile acids-induced gastric intestinal metaplasia. Gut 2019, 68, 1751–1763.
- Li, H.; Wu, Q.; Li, T.; Liu, C.; Xue, L.; Ding, J.; Shi, Y.; Fan, D. The miR-17-92 cluster as a potential biomarker for the early diagnosis of gastric cancer: Evidence and literature review. Oncotarget 2017, 8, 45060–45071.
- Zhu, Y.; Jiang, Q.; Lou, X.; Ji, X.; Wen, Z.; Wu, J.; Tao, H.; Jiang, T.; He, W.; Wang, C.; et al. MicroRNAs Up-Regulated by CagA of Helicobacter pylori Induce Intestinal Metaplasia of Gastric Epithelial Cells. PLoS ONE 2012, 7, e35147.
- Qu, M.; Li, L.; Zheng, W.-C. Reduced miR-490-3p expression is associated with poor prognosis of Helicobacter pylori induced gastric cancer. Eur. Rev. Med. Pharmacol. Sci. 2017, 21, 3384–3388.
- Shen, J.; Xiao, Z.; Wu, W.K.; Wang, M.H.; To, K.F.; Chen, Y.; Yang, W.; Li, M.S.; Shin, V.Y.; Tong, J.H.; et al. Epigenetic Silencing of miR-490-3p Reactivates the Chromatin Remodeler SMARCD1 to Promote Helicobacter pylori–Induced Gastric Carcinogenesis. Cancer Res. 2015, 75, 754–765.
- Min, J.; Han, T.-S.; Sohn, Y.; Shimizu, T.; Choi, B.; Bae, S.-W.; Hur, K.; Kong, S.-H.; Suh, Y.-S.; Lee, H.-J.; et al. microRNA-30a arbitrates intestinal-type early gastric carcinogenesis by directly targeting ITGA2. Gastric Cancer 2020, 23, 600–613.
- Ma, M.; Zhang, Y.; Weng, M.; Hu, Y.; Xuan, Y.; Hu, Y.; Lv, K. lncRNA GCAWKR Promotes Gastric Cancer Development by Scaffolding the Chromatin Modification Factors WDR5 and KAT2A. Mol. Ther. 2018, 26, 2658–2668.
- Yang, L.; Long, Y.; Li, C.; Cao, L.; Gan, H.; Huang, K.; Jia, Y. Genome-Wide Analysis of Long Noncoding RNA Profile in Human Gastric Epithelial Cell Response to Helicobacter pylori. Jpn. J. Infect. Dis. 2015, 68, 63–66.
- Sun, T.-T.; He, J.; Liang, Q.; Ren, L.-L.; Yan, T.-T.; Yu, T.-C.; Tang, J.-Y.; Bao, Y.-J.; Hu, Y.; Lin, Y.; et al. LncRNA GClnc1 Promotes Gastric Carcinogenesis and May Act as a Modular Scaffold of WDR5 and KAT2A Complexes to Specify the Histone Modification Pattern. Cancer Discov. 2016, 6, 784–801.
- Weiss, G.A.; Hennet, T. Mechanisms and consequences of intestinal dysbiosis. Cell. Mol. Life Sci. 2017, 74, 2959–2977.
- Barko, P.; McMichael, M.; Swanson, K.; Williams, D. The Gastrointestinal Microbiome: A Review. J. Veter Intern. Med. 2017, 32, 9–25.
- Morkūnas, E.; Skiecevičienė, J.; Kupčinskas, J. The impact of modulating the gastrointestinal microbiota in cancer patients. Best Pr. Res. Clin. Gastroenterol. 2020, 48-49, 101700.
- Toor, D.; Wasson, M.K.; Kumar, P.; Karthikeyan, G.; Kaushik, N.K.; Goel, C.; Singh, S.; Kumar, A.; Prakash, H. Dysbiosis Disrupts Gut Immune Homeostasis and Promotes Gastric Diseases. Int. J. Mol. Sci. 2019, 20, 2432.
- Kupcinskas, J.; Hold, G.L. Other Helicobacters and the gastric microbiome. Helicobacter 2018, 23, e12521.
- Schulz, C.; Kupčinskas, J. Review-Helicobacter pylori and non-malignant upper gastro-intestinal diseases. Helicobacter 2020, 25 (Suppl. 1), e12738.
- Vinasco, K.; Mitchell, H.M.; Kaakoush, N.O.; Castaño-Rodríguez, N. Microbial carcinogenesis: Lactic acid bacteria in gastric cancer. Biochim. Biophys. Acta (BBA) Bioenerg. 2019, 1872, 188309.
- Watanabe, T.; Nadatani, Y.; Suda, W.; Higashimori, A.; Otani, K.; Fukunaga, S.; Hosomi, S.; Tanaka, F.; Nagami, Y.; Taira, K.; et al. Long-term persistence of gastric dysbiosis after eradication of Helicobacter pylori in patients who underwent endoscopic submucosal dissection for early gastric cancer. Gastric Cancer 2021, 24, 710–720.
- Spiegelhauer, M.R.; Kupcinskas, J.; Johannesen, T.B.; Urba, M.; Skieceviciene, J.; Jonaitis, L.; Frandsen, T.H.; Kupcinskas, L.; Fuursted, K.; Andersen, L.P. Transient and Persistent Gastric Microbiome: Adherence of Bacteria in Gastric Cancer and Dyspeptic Patient Biopsies after Washing. J. Clin. Med. 2020, 9, 1882.
- Rajilic-Stojanovic, M.; Figueiredo, C.; Smet, A.; Hansen, R.; Kupcinskas, J.; Rokkas, T.; Andersen, L.; Machado, J.C.; Ianiro, G.; Gasbarrini, A.; et al. Systematic review: Gastric microbiota in health and disease. Aliment. Pharmacol. Ther. 2020, 51, 582–602.
- Alarcón, T.; Llorca, L.; Perez-Perez, G. Impact of the Microbiota and Gastric Disease Development by Helicobacter pylori. Curr. Top. Microbiol. Immunol. 2017, 400, 253–275.
- Castaño-Rodríguez, N.; Goh, K.-L.; Fock, K.M.; Mitchell, H.M.; Kaakoush, N.O. Dysbiosis of the microbiome in gastric carcinogenesis. Sci. Rep. 2017, 7, 1–9.
- Yu, C.; Su, Z.; Li, Y.; Li, Y.; Liu, K.; Chu, F.; Liu, T.; Chen, R.; Ding, X. Dysbiosis of gut microbiota is associated with gastric carcinogenesis in rats. Biomed. Pharmacother. 2020, 126, 110036.
- Gong, J.; Li, L.; Zuo, X.; Li, Y. Change of the duodenal mucosa-associated microbiota is related to intestinal metaplasia. BMC Microbiol. 2019, 19, 1–7.
- Jiménez, F.A.; Vazquezjimenez, F.E.; Medrano-Guzman, R.; Mantilla, A.; Torres, J. Stomach microbiota composition varies between patients with non-atrophic gastritis and patients with intestinal type of gastric cancer. Sci. Rep. 2015, 4, 4202.
- Sung, J.J.Y.; Coker, O.O.; Chu, E.; Szeto, C.H.; Luk, S.T.Y.; Lau, H.C.H.; Yu, J. Gastric microbes associated with gastric inflammation, atrophy and intestinal metaplasia 1 year after Helicobacter pylori eradication. Gut 2020, 69, 1572–1581.
- Eun, C.S.; Kim, B.K.; Han, D.S.; Kim, S.Y.; Kim, K.M.; Choi, B.Y.; Song, K.S.; Kim, Y.S.; Kim, J.F. Differences in Gastric Mucosal Microbiota Profiling in Patients with Chronic Gastritis, Intestinal Metaplasia, and Gastric Cancer Using Pyrosequencing Methods. Helicobacter 2014, 19, 407–416.
- Park, C.H.; Lee, A.-R.; Lee, Y.-R.; Eun, C.S.; Kil Lee, S.; Han, D.S. Evaluation of gastric microbiome and metagenomic function in patients with intestinal metaplasia using 16S rRNA gene sequencing. Helicobacter 2019, 24, e12547.
- Wang, Z.; Gao, X.; Zeng, R.; Wu, Q.; Sun, H.; Wu, W.; Zhang, X.; Sun, G.; Yan, B.; Wu, L.; et al. Changes of the Gastric Mucosal Microbiome Associated With Histological Stages of Gastric Carcinogenesis. Front. Microbiol. 2020, 11, 997.
- Gao, J.-J.; Zhang, Y.; Gerhard, M.; Mejias-Luque, R.; Zhang, L.; Vieth, M.; Ma, J.-L.; Bajbouj, M.; Suchanek, S.; Liu, W.-D.; et al. Association Between Gut Microbiota and Helicobacter pylori-Related Gastric Lesions in a High-Risk Population of Gastric Cancer. Front. Cell. Infect. Microbiol. 2018, 8, 202.
