Muellerian-type tumors of the urinary tract (MTUT) were implemented in the 2016 WHO classification as an entity comprising clear cell adenocarcinomas (CCAs) and endometrioid adenocarcinomas (EAs) of the bladder/urinary tract [1]. The histologically clear cell tumors were initially described as mesonephric adenocarcinomas by Konnak in 1973 [2] and re-named as clear cell adenocarcinomas in 1985 by Young and Scully [3].
1. Introduction
MTUT are rare malignancies and occur preferably in women (female-to-male ratio 2:1) or in case of endometrioid adenocarcinomas exclusively in women [1]. However, recently, a series of n = 15 CCAs in men was also reported [4]. Most of the CCAs are located in the bladder neck and trigone, less frequently in the lower posterior bladder wall, whereas EAs are located in the trigone and the posterior wall, where endometriosis is most common [1]. Due to lack of information, prognosis of EAs is not assessable [1]. Prognosis of CCAs depends on tumor growth/stage (exophytic tumors are more favorable) and aggressive treatment [1], since theses tumors respond poorly to chemotherapy or radiotherapy and show the best long term survival after radical cystectomy [5].
Diagnostically striking features are hobnail-like tubulocystic, papillary, or diffusely arranged cells with either clear glycogen-rich or eosinophilic cytoplasm (clear cell adenocarcinomas) or typically endometrioid features with squamoid nests (endometrioid adenocarcinomas) [
1]. Secondary bladder infiltration from tumors of neighbor organs must be excluded. The most helpful immunohistochemical markers are PAX2 or PAX8, which are positive in all cases [
6], but keratin7, EMA, HNF1ß, and CA125 are usually positive as well [
1]. The immunohistochemical profile of endometrioid adenocarcinomas is similar to the female genital counterparts, including hormone receptor expression [
1].
Due to the low incidence rates, the biological background and implications for clinical management are not yet well understood, and first reports are now available [
7]. In the latest study of Lin et al., they analyzed four CCAs by targeted panel sequencing and identified BK virus-mediated oncogenesis (one of four cases) and PI3K/AKT/mTOR pathway activation (three of four cases) as underlying tumorigenic events, waiting for further validation in lager cohorts [
7]. Due to these findings, we hypothesized that clear cell adenocarcinomas might be closer related to genital clear cells and nephrogenic cells than urothelial cells. Therefore, in the presented work, we collected Muellerian-type tumors and analyzed their molecular alterations on SNV, CNV, and gene fusion level and compared it with available TCGA data from bladder, endometrioid, and clear cell kidney cancers in order to understand tumorigenesis and identify new treatment options.
2.Histomorphological and Immunohistochemical Diagnostic Evaluation
In total, 10 cases showed typical clear cell morphology with a predominantly papillary architecture in 50% (5/10) of cases, followed by 30% (3/10) tubulo-cystic pattern, and only 20% (2/10) of cases exhibited a predominant solid clear cell tumor mass.
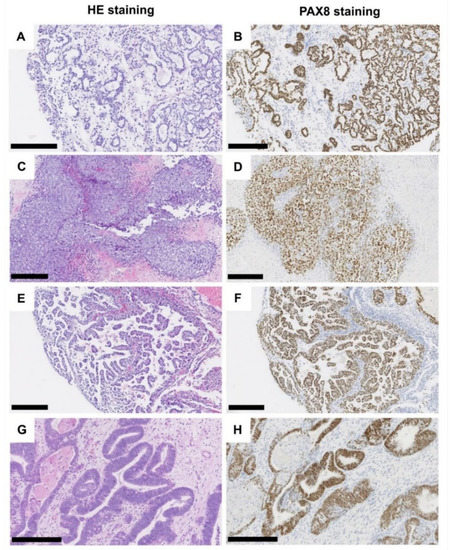
One case showed the typical endometrioid histology with a glandular pattern and more or less distinct squamous areas. All cases presented with invasive growth and revealed a strong nuclear positivity for PAX8, medium to strong cytoplasmatic and membranous positivity for keratin7, and only weak to absent staining for GATA3 (see ). illustrates the different histomorphological phenotypes of MTUT.
Figure 1. Histological characteristics of tumors of the Muellerian type. Clear cell adenocarcinoma, tubule-cystic growth pattern (A), HE, (B) PAX8 immunohistochemistry. Clear cell adenocarcinoma, solid growth pattern (C), HE, (D) PAX8 immunohistochemistry. Clear cell adenocarcinoma, (pseudo-/micro-) papillary growth pattern (E), HE, (F) PAX8 immunohistochemistry. Endometrioid adenocarcinoma (G), HE, (H) PAX8 immunohistochemistry. Black scale bar: 250 µM.
3. Mutational Analysis
DNA NGS panel analysis was successfully performed for all 11 samples. Region-of-interest coverage was at least 91% (up to 98%) at 500× coverage (average 95%). A total of 32 variants of pathogenic or at least uncertain significance emerged after filtering (B;
Table S1). The identified mutations were 25 missense mutations (78%), six truncating mutations (19%), and one in-frame deletion (3%). A total of 22 variants were detected in genes classified as tumor suppressors according to the OncoKB database (69%), whereas nine variants were found in corresponding oncogenes (28%). For one variant of the
NOTCH3 gene, both functions were described according to OncoKB (3%).
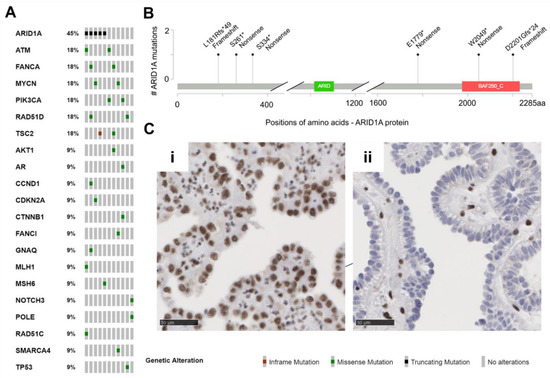
Figure 2. Muellerian tumors of the urinary tract (MTUT) are characterized by frequent mutations in ARID1A. (A) Oncoprint image illustrates single nucleotide variations (SNVs) including in-frame mutations, missense mutations, truncating mutations. (B) Positions and type of ARID1A mutations identified in MTUT. (C) ARID1A protein expression/loss in exemplary MTUT tissues. (i) MTUT tissue with ARID1A wild type shows strong nuclear staining of ARID1A protein. (ii) ARID1A protein loss in MTUT tissue with p.Ser334* ARID1A mutation. Black scale bar: 50 µM.
No identical recurrent mutations were detected. Except for one ARID1A mutated case, in none of the tumor suppressor genes were two mutations of the same gene detected (patient MT-10). The most frequently mutated gene (5/11 patients, 45%, 6/32 variants 19%) in our cohort was ARID1A (B). Interestingly, all six detected variants were clearly truncating mutations, which resulted in loss of protein in immunohistochemical analysis in only one patient (MT-6: p.Ser334*, Cii). Two further ARID1A mutations led to a partial protein loss (MT-4: p.Ser261*, MT-5: p.Asp2201Glyfs*24), and the residual mutations (MT-7: p.Leu181Argfs*49) showed retained ARID1A protein expression or immunohistochemistry was not possible (MT-10: p.Glu1779*, p.Trp2049*). Further, mutations of ATM, FANCA, MYCN, PIK3CA, RAD51D, and TSC2 genes were observed two times each, while single changes in other genes were detected only once (AKT1, AR, CCND1, CDKN2A, CTNNB1, FANCI, GNAQ, MLH1, MSH6, NOTCH3, POLE, RAD51C, SMARCA4, TP53).
To reliably exclude
TERT promoter mutations, in particular, the known hotspot mutations c.-146C>T (“C250T”) c.-124C>T (“C228T”), we performed a separate coverage analysis with respect to
TERT [
14]. For all samples, a minimum coverage of 100× in the range of the two mentioned mutations was achieved (90.9% of cases >300×, 63.6% of cases >500×). A
TERT promoter mutation could not be identified in any of the 11 patients.
4. Copy Number Analysis
All 11 analyzed samples passed our internal CNV quality assessment (quality scores ranged from 97%–100%, mean value: 99%). In total, we identified 13 copy number alterations (only amplifications) in seven samples of our cohort. Nine of the 13 detected CNVs affected oncogenes (69%), whereas four of the detected amplifications involved tumor suppressor genes (31%).
In one patient, four CNVs were detected. However, two of the CNVs were located on chromosome 3p25.2–3p25.3 in spatial proximity to each other, therefore, a larger chromosomal event seems to be likely in this case (patient MT-7: 3p25.2–3p25.3: 1×
FANCD2, 1×
PPARG; additionally: 1×
MET, 1×
NBN). Another patient harbored three CNVs, with two of the affected genes chromosomally adjacent (patient MT-8: 1× ERBB2, 1×
CDK12: chromosome 17q12; also: 1×
FGFR1). Additionally, in the last case with multiple CNVs, amplifications of two genes in spatial proximity were detected (patient MT-2: 1×
NTRK1, 1×
DDR1; chromosome 1q23.1–1q23.3). The remaining four cases showed one amplification event each (1×
FGFR1, 1×
CDK6, 1×
ERBB3, 1×
NF1). Therapeutically relevant amplifications were further evaluated by FISH.
MET-FISH revealed a polysomy with four to six copies of chromosome 7 (ratio
MET signals (132)/CEN 7 (104) = 1.27 and ratio
MET signals (132)/number of cells (20) = 6.6).
ERBB2-FISH proved a true amplification of
ERBB2 (ratio
ERBB2 signals (324)/CEN 17 (132) = 2.45) with one to three copies of chromosome 17 and immunohistochemically strong baso-lateral Her2neu staining. A summary of the CNV data is shown in ,
Table S2.
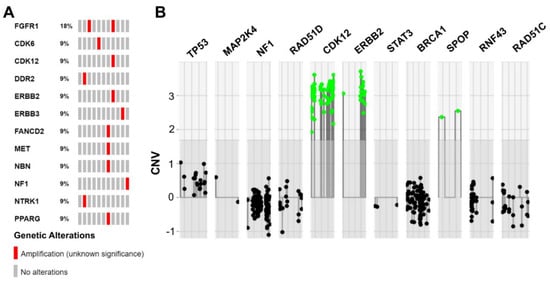
Figure 3. Summarized copy number variations (CNVs) identified in MTUT. (
A) Oncoprint image illustrates detected CNVs. (
B) Illustration extracted from the ACopy tool [
8] showing the co-occurring amplification of
ERBB2 and
CDK12 in patient MT-8 (x-axis: 0 corresponds to two copies).
5. Fusion Gene Analysis
Analysis of fusion genes on RNA level was successfully performed for all 11 patients. At least 230,000 single RNA reads were obtained per case (range: 235,710–4,770,440, mean value: 1,127,386 reads). A total of 10 of the cases did not show gene fusions, however, in one patient with the endometrioid adenocarcinoma (MT-2), an
RSPO2 rearrangement was detected, which was already described as functionally relevant in hepatocellular adenoma (
n = 3/17, [
15]). The fusion connects an intergenic short interspersed nuclear element (SINE) upstream of
RSPO2 (chr8:109141374) to exon 2 of the
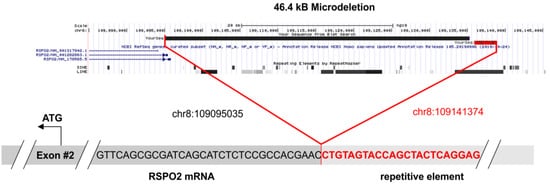
RSPO2 gene (chr8:109095035). Ultimately, 16,750 supporting reads were obtained. Analysis of the fastq files confirmed the breakpoints of the 46.4 kb microdeletion on chromosome 8q23.1 (,
Table S3).
Figure 4. The 46.4 kB microdeletion of patient MT-2. In the upper panel, the deleted sequence is demonstrated. The chromosomal region was extracted from the UCSC genome browser (
https://genome.ucsc.edu/, accessed on 15 April 2021) and the exact breakpoints are listed. The lower panel shows the fusion of the intronic repetitive element to the first (non-coding) base of exon 2. The sequence data were extracted from the original fastq file and were aligned to the reference genome GRCh37/hg19.
6. Comparison with Publicly Available TCGA Data of Bladder, Endometrial, and Kidney Cancer
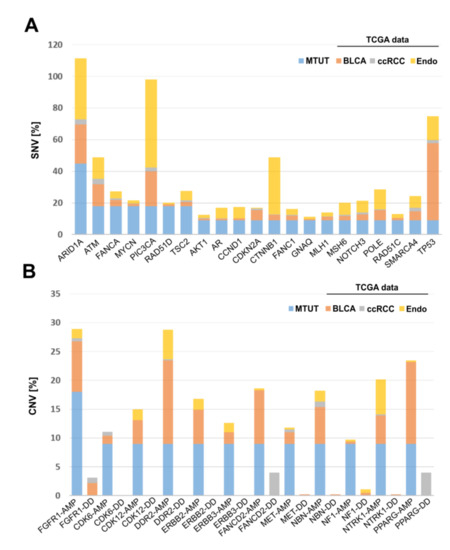
Finally, the identified SNV and CNV alterations in Muellerian-type tumors of the urinary tract were compared with cancers of other origin, i.e., with bladder (BLCA, n = 412), endometrial (n = 205), and clear cell kidney (ccRCC, n = 426) cancer of the TCGA platform. SNVs are illustrated in A and revealed the lowest similarity between MTUT and clear cell kidney cancers, showing overall very low mutational frequency of analyzed genes. Contrary to that, we identified distinct gene alterations such as ATM and POLE present in MTUT with comparable frequencies in both ±3%, i.e., bladder (ATM: 13.8, POLE: 6.6) and endometrial cancers (ATM: 13.7, POLE: 12.7), while genes such as PIC3CA or TP53 showed only parallels with one of the two entities. Interestingly, we also observed gene alterations which seemed to be more specific for MTUT such as MYCN and RAD51D.
Figure 5. Comparison of genetic alterations found in MTUT with TCGA data sets of bladder, endometrial, and clear cell kidney cancers. (A) SNVs are shown. (B) CNVs are illustrated. AMP: amplification; DD: deep deletion, BLCA: bladder cancer, ccRCC: clear cell renal cell carcinoma.
Focusing on CNVs (B), ccRCC exhibited amplifications of analyzed genes only in rare cases, whereas deep deletions were much more frequent for PPARG and FANCD2. No similarities were observed when comparing MTUT and endometrial cancers, showing only an increased amplified status for DDR2. In turn, CNV data revealed parallels between bladder cancers and MTUT for most of analyzed genes, such as FGFR1, according to both frequency and type of CNV, i.e., amplifications were mainly present, whereas deep deletions occurred
rarely.
This entry is adapted from the peer-reviewed paper 10.3390/genes12060880