AKR1B10 is a human nicotinamide adenine dinucleotide phosphate (NADPH)-dependent reductase belonging to the aldo-keto reductase (AKR) 1B subfamily. It catalyzes the reduction of aldehydes, some ketones and quinones, and interacts with acetyl-CoA carboxylase and heat shock protein 90α. The enzyme is highly expressed in epithelial cells of the stomach and intestine, but down-regulated in gastrointestinal cancers and inflammatory bowel diseases. In contrast, AKR1B10 expression is low in other tissues, where the enzyme is upregulated in cancers, as well as in non-alcoholic fatty liver disease and several skin diseases. In addition, the enzyme’s expression is elevated in cancer cells resistant to clinical anti-cancer drugs. Thus, growing evidence supports AKR1B10 as a potential target for diagnosing and treating these diseases. Herein, we reviewed the literature on the roles of AKR1B10 in a healthy gastrointestinal tract, the development and progression of cancers and acquired chemoresistance, in addition to its gene regulation, functions, and inhibitors.
- aldo-keto reductases
- AKR1B10
- biomarkers
1. Introduction
2. Gene Regulation of AKR1B10
2.1. Factors Regulating AKR1B10 Expression

| Agent * | Signal Molecule | Cell ** | References |
|---|---|---|---|
| Up-regulation | |||
| Ethoxyquin | Nrf2 | Lung cancer A549, H23 | [14] |
| MG-132, bortezomib | Nrf2 | CRC SW-480, HT29 | [17] |
| Doxorubicin | Nrf2 | Gastric cancer MKN45 | [18] |
| EGF, insulin | AP-1, ERK | HCC HepG2, Hep3B | [19] |
| Lipopolysaccharide | Blood mononuclear cells | [20] | |
| BMP, IBMX | Mesenchymal stem cells | [21] | |
| 9,10-Phenanthrenequinone | Nrf2, ERK | Lung cancer A549 | [22] |
| Cigarette smoke extract | Airway epithelium | [23] | |
| Carnosic acid, t-BHQ | Nrf2 | Astrocytoma U373MG | [15] |
| 5-FU, L-OHP | p53 | CRC HT116 | [16] |
| Down-regulation | |||
| TPA | c-Jun, ERK | Lung cancer A549 | [15] |
| 5-FU, L-OHP | CRC HT29 | [16] |
2.2. Contribution of Nrf2 to AKR1B10 Induction
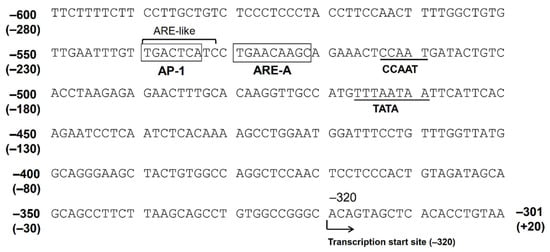
2.3. The Function of AP-1 Protein in AKR1B10 Gene Regulation
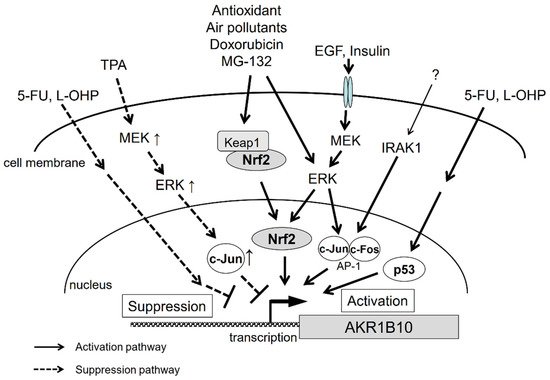
2.4. Signal Transduction
3. Conclusions
This entry is adapted from the peer-reviewed paper 10.3390/metabo11060332
