Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is an old version of this entry, which may differ significantly from the current revision.
Subjects:
Nanoscience & Nanotechnology
Using DNA self-assembly, materials can be controlled at the nano scale to achieve atomic- or nano-scaled fabrication. The programmability and addressability of DNA molecules can be applied to realize the self-assembly of materials from the bottom-up, which is called DNA nanotechnology. DNA nanotechnology does not focus on the biological functions of DNA molecules, but combines them into motifs, and then assembles these motifs to form ordered two-dimensional (2D) or three-dimensional (3D) lattices. These lattices can serve as general templates to regulate the assembly of guest materials.
- self-assembly
- bottom-up
- DNA tile
- DNA brick
- nanoparticles
1. Self-Assembly Based on DNA Tile
In 1982, Seeman proposed using DNA to construct a 3D periodic network, which is considered to be the origin of “DNA nanotechnology” (Figure 1a) [1]. In the early years, DNA branched junctions were selected as basic building blocks for constructing 3D periodic networks, as shown in Figure 1b. In 1983, Seeman et al. successfully synthesized the four-arm junction structure of DNA [16]. DNA building blocks are required to have a certain rigidity for assembling into a larger structure, but DNA branched junctions obviously fail to meet this feature. By introducing a crossover between two DNA double helices, Seeman et al. designed the “DNA double-crossover molecule” (DX) with sufficient rigidity to form a larger structure [17]. In the mathematical theory of tiling, rectangular tiles with programmable interactions, known as Wang tiles, can be tiled into 2D spaces. DX molecules can be arranged in a 2D space imitating a Wang tile, and single-stranded DNA sequences can be extended on both sides of the DX molecule as sticky ends to connect other DX molecules. Reasonably designed sticky ends can assemble DX molecules and DX + J molecules into a 2D array (Figure 1c) [6,18]. The structure of a single DNA tile is relatively simple, consisting of several stoichiometric single oligonucleotide strands. Simple small units are repeatedly arranged to form a large 2D array through the connection of sticky ends. However, this method has relatively low control over the size and shape of the product. Li et al. proposed to restrict the size and shape of DNA tile growth with a prescribed DNA origami frame [19]. A 2D array consisting of a large number of simple repeating DNA tiles is filled in a hollow origami frame, which have faster growth kinetics when in an origami frame than when there is no frame. The design of DX molecules has been modified to obtain different DNA building blocks, such as paranemic crossover (PX) DNA, topoisomer (JX) DNA, and triple crossover complex. Through the strand displacement reaction, PX and JX can be converted into one another, that is, one end of a DNA strand rotates 180° relative to the other end. Seeman used this conformational change between PX and JX to confirm that a rotary nanomechanical device can be recycled [20].
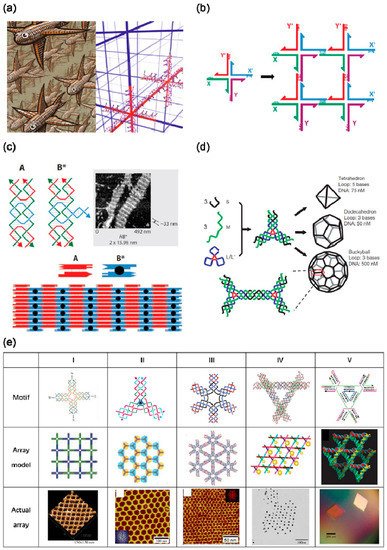
Figure 1. Self-assembly based on DNA tiles (a) Escher’s woodcut depth (left) and the prototype of DNA nanotechnology inspired by it (right). Reproduced from [21]. (b) A schematic diagram of a two-dimensional (2D) DNA lattice assembled by four-arm holiday junctions connected by sticky ends. Reproduced with permission [22]. Copyright Materials Research Society, 2017. (c) A 2D lattice assembled by DNA double-crossover molecule (DX) and DX + J tiles. Reproduced with permission of [22]. Copyright Materials Research Society, 2017. (d) The structure of the DNA polyhedron assembled by three-pointed star DNA motifs. Reproduced with permission of [11]. Copyright Springer Nature, 2008. (e) DX-based complex DNA tile motif library and its assembly results. (I) 4 × 4 tile. Reproduced with permission of [9]. Copyright American Association for the Advancement of Science, 2003; (II) three-pointed star. Reproduced with permission of [23]. Copyright American Chemical Society, 2005; (III) six-pointed star. Reproduced with permission of [24]. Copyright American Chemical Society, 2006; cop; (IV) DX-based tensegrity triangle. Reproduced with permission of [25]. Copyright American Chemical Society, 2006; (V) a tensegrity triangle with complete triple symmetry. Reproduced from [10].
A simple DX structure can be expanded into a high-order DNA motif, which can be assembled into a more complex 2D array, as shown in Figure 1e. Yan et al. designed a 4 × 4 DNA tile that can self-assemble into uniform-width nanoribbons, 2D nano−grids and ladder-like grids [10]. These grids can be subsequently used for the periodic arrangement of protein molecules and gold nanoparticles [26,27,28]. In addition, they also confirmed that sophisticated 2D and 3D tessellation patterns can be formed by using three- and four-arm DNA junction tiles with specifically designed arm lengths and inter-tile sticky-end interactions [29]. Mao et al. designed a series of n-pointed-stars and assembled them into 2D arrays with different topologies [23,24,30,31,32]. In addition, five- and six-pointed-stars have been co-crystallized to a 2D array similar to a quasi-crystal arrangement [33]. By controlling the flexibility and concentration of motifs, a DNA n-pointed-star motif can be assembled into a complex polyhedron framework structure (Figure 1d) [11,31], which can then be used as a nanocage to encapsulate nanoparticles in the frame to form clusters with a specific conformation [34,35,36,37]. Another complex motif formed by simple DNA tiles is tensegrity triangle. Mao et al. used three four-arm junctions to construct a tensegrity triangle with a double helix on each side [30] with sticky ends at each vertex, allowing it to self-assemble into one-dimensional (1D) or 2D ordered arrays by selectively using the vertices. Seeman et al. designed a tensegrity triangle composed of DX molecules, and used their assembled 1D and 2D arrays to complete the regular arrangement of gold nanoparticles (Figure 1e) [25]. Subsequently, Seeman et al. designed a tensegrity triangle with triple rotational symmetry in sequence [10]. The structure was assembled with sticky ends to obtain a single crystal with a rhombohedral lattice (Figure 1e). By adjusting the length of the sides of the tensegrity triangle, single crystals with different lattice parameters were obtained with cavities of different sizes, and the cavities could be used to accommodate guest particles. This work is considered to be a major breakthrough in the field of DNA nanotechnology. Since then, a lot of work has been carried out around the tensegrity triangle, such as controlling the crystal nucleation and growth process [38], improving the quality and stability of the single crystal [39,40], and completing dynamic changes [41].
The self-assembly product of DNA tiles can be used for the manipulation of nanoparticles. However, due to the limitation of DNA tile assembly products, nanoparticles can only be manipulated on a linear or planar template. The particle spacing and arrangement can be well controlled and, the polyhedral frame assembled by tiles can be used as a nanocage for nanoparticle encapsulation or as a restriction frame template to guide nanoparticle assembly. Thus, clusters of nanoparticles can be obtained, but it is difficult to expand them into a 3D space to form ordered crystals.
2. Self-Assembly Based on DNA Brick
The results of electron microscopy characterization indicate that the size and shape of a 2D array assembled by the DNA tile mentioned above cannot be controlled. Wei et al. proposed a “single-stranded DNA tile” (SST) that consists of a 42-base strand of DNA composed entirely of concatenated sticky ends. Therefore, each SST has the ability to bind with four local neighbors during self-assembly (Figure 2a) [42].
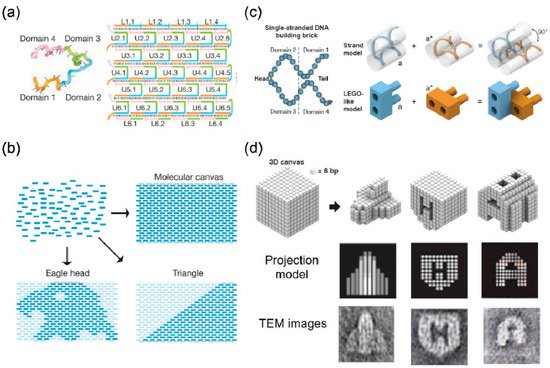
Figure 2. Single-stranded DNA tiles and single-stranded DNA bricks. (a) Rectangular 2D molecular canvas assembled by single-stranded DNA tiles. Reproduced from [42]. (b) Drawing the desired shape on the 2D molecular canvas. Reproduced from [42]. (c) LEGO model analogue of single-stranded DNA. Reproduced from [7]. (d) “Carving” the 3D canvas into any shape. Reproduced from [7].
SST tiles can be assembled into a rectangular 2D array as a molecular canvas, where each SST DNA sequence is different and corresponds to a particular pixel in the molecular canvas (Figure 2a). The expected shape is drawn on the molecular canvas to find the target SST corresponding to all pixels in the shape, and then the target SST can be annealed in one pot to obtain the desired shape (Figure 2b). In this article, 107 unique and complex 2D shapes were synthesized in this way. DNA bricks are an extension of the concept of SST, extending the modular-assembly from 2D to 3D. Ke et al. first proposed the concept of DNA brick and used this method to construct more than 100 different 3D structures, as shown in Figure 2b,c [7]. Each brick is a single-stranded DNA with a distinct nucleic acid sequence. Utilizing the twist of the DNA helix, the complementary base pairing between adjacent bricks produces a dihedral angle close to 90 degrees (Figure 2c). Many bricks self-assemble into a 3D cubic molecular canvas onto which can be “carved” a variety of different 3D shapes (Figure 2d). Through one-pot annealing, the DNA strands corresponding to the bricks self-assemble to form the pre-designed 3D shape. DNA bricks provide a simple and modular method to assemble simple short DNA strands into complex 3D shapes. Later, Ke et al. constructed a 2D crystal with complex 3D geometric features (with the thickness dimension much smaller than the other two dimensions) and used it for the regular arrangement of gold nanoparticles [43]. The team later designed a second-generation brick with a longer bonding domain—because of which, it can self-assemble to form a larger and more complex 3D structure [12].
The largest advantage of DNA bricks is that the same batch of DNA bricks can be carved into any shape. In addition, since the DNA sequence of each brick is different, the composed 3D structure can be used as an addressable template to place the guest particles with a finer degree of control. Ke et al. also realized the single-layer and dispersed arrangement of gold nanoparticles on the 2D materials assembled by bricks [43]; however, the formation of multi-layer patterns of gold nanoparticles remains a challenge.
This entry is adapted from the peer-reviewed paper 10.3390/nano10102047
This entry is offline, you can click here to edit this entry!
