Nerve growth factor (NGF) is a protein essential to neurons survival, which interacts with its receptor as a non-covalent dimer. Copper affects biological activity of NGF and conversely NGF may regulates copper trafficking in synaptic cleft.
- copper
- NGF
- Ctr-1
- Alzheimer’s
- protein
- peptidomimetic
1. Introduction
Copper performs several essential functions as a cofactor in many enzymes in the living systems and it is required for the development and function of the human brain [1,2,3]. In recent years, it has been shown that copper, analogously to zinc, plays a role as modulator of cellular signal transduction pathways [4,5,6]. Copper is stored in synaptic vesicles and is released by electrical depolarization in the synaptic cleft of glutamatergic synapses at concentration values that can reach 100 μM [7]. Copper may have inhibitory or stimulating effect on synaptic plasticity, affecting memory and learning processes although the mechanisms by which the metal performs these functions remain largely undefined [8,9]. The dual effect may be related to the metal binding by different proteins expressed in neurons and released in the synapses as beta-amyloid (Aβ) [10].
The actual biological function of Aβ is still unknown. The polypeptide may control copper efflux in the synapses and it has been demonstrated that the polypeptide improves memory formation [11], synaptic plasticity, and neuronal survival [12].
In particular, the monomeric form of Aβ activates the cyclic AMP response element-binding protein (CREB), which in turn promotes the transcription and release of the brain-derived neurotrophic factor (BDNF) a neurotrophin (NT) strongly involved in long term potentiation (LTP) and memory formation [13].
It is worth noting that it has been demonstrated that copper ions may modulate kinase signaling networks of neuronal tissues induced by neurotrophins [14]. The nerve growth factor (NGF) is the first discovered member of NTs family [15]. NGF is essential for the development, survival, and activity of neurons [16,17]. NGF initiates the signaling pathways through the binding to tropomyosin receptor kinase A (TrkA) triggering a signaling cascade up to the activation of CREB [18,19].
Copper ions enhance NGF functionality and the effect seems related to the presence of metal binding site in the N-terminus domain of NGF [20]. NGF(1-14), a peptide encompassing the first 14 residue of NGF, is able to bind copper ions and mimic whole protein signal transduction activating CREB [21]. The NGF interacts with TrkA as a non-covalent dimer, so a dimeric form of the peptide NGF(1–14) has been tested, showing higher activity than monomers in the release of BDNF [14].
NGF signaling pathways control also the post-translational modifications of the amyloid precursor protein (APP) and then Aβ production in neurons [22,23,24]. Moreover, the deprivation in NGF determines Aβ aggregation and tau hyperphosphorylation in Alzheimer’s disease (AD) [25] while NGF addition protects against cell death and toxicity triggered by Aβ but the underlying mechanism remains unclear [26]. On the other hand, astrocytes activated by Aβ stimulate NGF secretion, whose excess in turn causes the death of hippocampal neurons [27].
In the dynamic environment present at the synaptic cleft, a potential interconnection among copper, Aβ, and NGF, could be one of the key components of the memory formation process.
Therefore, the dyshomeostasis of copper could be at the basis of Aβ and/or NGF dysfunctions; conversely, a malfunction in the metabolic pathways could negatively affect the normal influx of copper into neurons [28]. The consequences could lead to the development of the progressive neurodegenerative disorder such as Alzheimer’s disease (AD) [29,30].
In the brains of patients affected by AD, a high copper concentration, up to 0.4 mM, is localized in senile plaques composed of amyloid aggregates [31]; however, the involvement of copper in AD is still controversial [32]. Studies show copper deficiency in patients affected by AD, and consequently a need to enhance copper levels so to restore normal metal concentration [33,34]; differently, other experiments point to metal overload and consequent demand to reduce copper concentration [35,36].
Copper dyshomeostasis with an increase in the labile pool of metal and a parallel decrease in the copper bound to proteins or peptides is the main interpretation [37,38,39]. A local imbalance of copper between extra- and intra-cellular spaces may drive its binding to Aβ, which may result in formation of oligomeric fibrils, amyloid, or amorphous aggregates, depending on the metal ion/peptide molar ratio.
Aβ may play the role of handling copper cellular influx/efflux via binding metal and transfer it to human copper transporter 1 (hCtr1), the main protein responsible for Cu import into cells [40]. Studies carried out with the peptide model showed that a peptide encompassing the first 14 residues of hCtr1 binds copper ions with higher affinity than N-terminal Aβ peptides [41]. For this reason, it is relevant to know the affinity of Aβ towards copper ions and many studies have been carried out with different techniques to determine the coordination environment of copper complexes formed with Aβ [42,43].
The pH value is another environmental condition that influences Aβ aggregation and Aβ copper binding. The brain pH is imbalanced towards mild acidic condition in aging and patients with AD [44]. A lowering of pH favors Aβ aggregation and strongly alters the release of pro-inflammatory cytokine affecting the uptake of Aβ by microglial cells [45].
Pathologies, such as ischemic stroke, induce a local decrease of extracellular pH due to an inflammatory insult that is as supposed to be an early event in AD progression [46]. Furthermore, the pH influences copper complex speciation with Aβ and mild acidosis can promote conformational changes or reactive oxygen species production promoting Aβ aggregate precipitation [47].
NGF has been postulated as potential therapeutic agent for the treatment of AD and its side effects, as loss of memory, and taking into account that NGF can bind copper ions in the same spaces shared by Aβ, it may be of interest to determine its affinity for copper(II).
Potentiometry is a technique that determines the stability constant values with a certain accuracy and permits also detection of minor species. This experimental technique may be the ideal choice when conditions allow for its application that can be limited by some factors such as poor solubility of longer or hydrophobic peptides. For this reason, potentiometric measurements have been carried out on shorter and more soluble Aβ fragments, in particular the N-terminal ones where the copper binding sites are found [43].
2. Protonation Constants
| Species | NGF(1-14) b | rNGF(14-1) | sNGF(1-14) | dNGF(1-15) |
|---|---|---|---|---|
| LH | 7.56 | 7.82 (2) | 7.85 (3) | 7.65 (5) |
| LH2 | 14.13 | 14.51 (2) | 14.65 (3) | 15.35 (2) |
| LH3 | 20.14 | 20.60 (2) | 20.87 (3) | - |
| LH4 | 24.44 | 24.88 (2) | 25.11 (4) | 28.77 (6) |
| LH5 | - | 35.12 (4) | ||
| LH6 | - | 41.31 (4) | ||
| LH7 | - | 46.62 (4) | ||
| LH8 | - | 50.75 (4) | ||
| pK COO- | 4.13 | 4.28 | 4.23 | 4.13 |
| pK COO- | - | - | - | 5.28 |
| pK His | 6.01 | 6.09 | 6.22 | 6.21 |
| pK His | 6.57 | 6.69 | 6.81 | 6.35 |
| pK His (×2) | - | - | - | (6.71 × 2) |
| pK NH2 | 7.56 | 7.82 | 7.85 | 7.65 |
| pK NH2 | - | 7.70 |
3. Speciation and Characterization of Copper-Peptide Complexes
| Species (pqr) b | logβpqr NGF(1-14)c |
logβpqr rNGF(14-1) |
logβpqr sNGF(1-14) |
logβpqr dNGF(1-15) |
|---|---|---|---|---|
| CuLH5 | - | - | - | 41.18 (3) |
| CuLH3 | - | - | - | 31.56 (2) |
| CuLH2 | - | - | - | 25.91 (5) |
| CuLH | 14.09 | 14.15 (1) | 13.97 (4) | 17.79 (5) |
| CuL | 8.72 | 8.77 (1) | 8.32 (5) | 8.21 (4) |
| CuLH-1 | 3.27 | 2.33 (2) | −2.74 (3) | - |
| CuLH-2 | −4.02 | −6.15 (4) | −5.58 (8) | - |
| CuLH-3 | −13.34 | −14.67 (2) | −13.27 (4) | - |
| pK (n/m) | ||||
| pK (5/3) | - | - | - | 4.81 × 2 |
| pK (3/2) | - | - | - | 5.65 |
| pK (2/1) | - | - | - | 8.12 |
| pK (1/0) | 5.37 | 5.38 | 5.65 | 9.57 |
| pK (0/−1) | 5.45 | 6.43 | 5.57 | - |
| pK (−1/−2) | 7.29 | 8.48 | 8.33 | - |
| pK (−2/−3) | 9.30 | 8.52 | 7.69 | - |
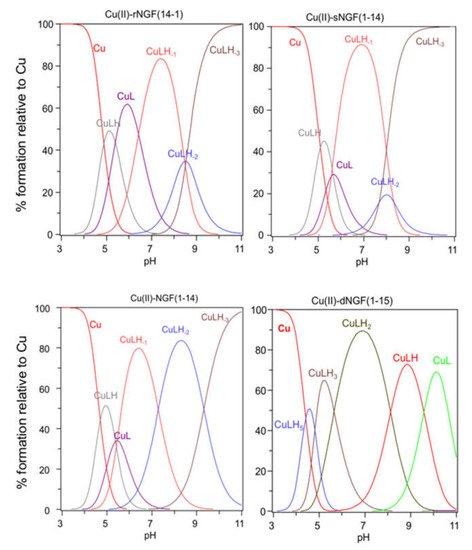
[CuLH] is the first complex species formed by rNGF(14-1) and sNGF(1-14) and reaches its maximum percentage of formation at pH 5 (Figure 2). The calculated stability constant values (logK(111) = logβ(111) − logβ(011)) are 6.33 and 6.12, for rNGF(14-1) and sNGF(1-14) respectively; the value obtained for reverse sequence is more similar to that of NGF(1-14) (logK(111) = 6.53). The logK(111) values are in the range 6.0–6.5, then higher than those of similar Cu2+ complexes of peptide fragments in which the metal ion is bound to one imidazole nitrogen and one carboxylate [48,51,52]. Indeed, these values are in good agreement with those reported for analogous complex species formed by Aβ peptide fragments for which metal ions have been shown to form a macrochelate with a 2Nim, COO coordination mode (log K = 6.18) [53,54].
Spectroscopic parameters measured at pH = 5, confirm the 2NIm,COO− coordination mode for copper ion and the higher stability of metal complex formed by reverse (λmax = 640 nm, ε = 64 M−1 cm−1) compared to scrambled peptide (λmax = 670 nm, ε = 40 M−1 cm−1) (Table 3). However, it must be underlined that the parameters are partly influenced by the presence of other species other than free copper.
| Peptide | pH | UV-vis λ (nm) (ε (M−1 cm−1)) |
CD λ (nm) (Δε (M−1 cm−1) |
|---|---|---|---|
| rNGF(14-1) | 5 | 640 (64) | 280 (−0.30); 316 (+0.20); 670 (−0.27) |
| 6 | 625 (144) | 280 (−0.40); 323 (+0.33); 671 (−0.44) | |
| 7.4 | 609 (178) | 280 (−0.30); 314 (−0.22); 352 (+0.07); 508 (+0.31); 625 (−0.60) | |
| 9 | 532 (194) | 280 (−1.30); 484 (+0.46); 589 (−0.79) | |
| 10 | 522 (232) | 280 (−1.80); 483 (+0.54); 577 (−1.00) | |
| sNGF(1-14) | 5 | 670 (40) | 289 (−0.20); 328 (+0.12); 647 (−0.04) |
| 6 | 617 (94) | 287 (−1.34); 328 (+0.68); 617 (−0.59) | |
| 7.4 | 603 (102) | 288 (−1.38); 329 (+0.82); 603 (−0.72) | |
| 9-10 | 522 (141) | 306 (+1.47); 544 (−1.13) | |
| dNGF(1-15) | 5 | 635 (50) | 280 (−0.34); 321 (+0.19); 669 (−0.24) |
| 6 | 612 (70) | 280 (−0.40); 323 (+0.33); 671 (−0.44) | |
| 7.4 | 569 (94) | 280 (−1.80); 324 (+0.92); 504 (+0.08); 611 (−0.30) | |
| 8 | 561 (104) | 280 (−2.04); 323 (+0.98); 496 (+0.11); 586 (−0.34) | |
| 9 | 554 (119) | 280 (−2.15); 324 (+0.98); 496 (+0.08); 587 (−0.38) | |
| 10 | 530 (138) | 280 (−2.14); 324 (+0.71); 563 (−0.46) |
Increasing the pH, [CuL] complex species is formed. This species is a minor one but the obtained logβ value suggest the involvement of a further donor atom as N-terminal group for both peptides. The reverse peptide also shows for this species a higher value than scrambled one and very similar to that reported for NGF(1-14).
The contemporary presence of an isomer in which a deprotonated amide is bound to metal instead of an amino group, that remains still protonated, cannot be ruled out.
In particular the Cu-rNGF(14-1) system shows a CD signal centered around 320 nm, that is diagnostic of a charge transfer from a deprotonated amide nitrogen to copper ion [55] (Figure 3).
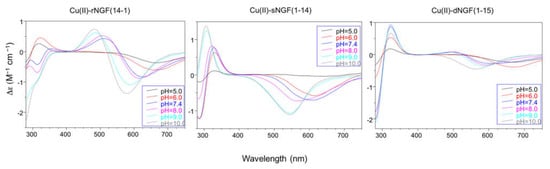
The next species formed, [CuLH-1], is the predominant complex in the pH range 6.5–8.0. The stepwise constant values logK(11-1) (logK(11-1) = logβ(110) − logβ(11-1)) are 6.43 for rNGF(14-1) and 5.57 for sNGF(1-14). Both values indicate the deprotonation of an amide nitrogen atom but this step is more favored for sNGF(1-14). The UV–vis parameters are similar suggesting an analogous coordination environment of metal ion with three nitrogen atoms involved in the binding. CD spectra show the diagnostic signal of deprotonated amide nitrogen bound to copper ion even though the conformational features of two peptides are different due to distinctive primary sequence (Figure 3).
As the pH increases, the second deprotonation occurs but it is less favored than the third nitrogen amide deprotonation step. Indeed, the [CuLH-2] complex species is a minor one for both peptides, suggesting that the coordination of successive amide nitrogen deprotonation reactions are accompanied by the rearrangement of the peptide metal binding sites [56]. Namely, the histidine imidazole moiety and the subsequent amide are the primary binding sites below pH 8.5, but they are partly replaced by the amino group and preceding amide functions at higher pH values. This effect is supported by significant blue shift of the absorption spectra as well as of the CD band characteristic of peptide amino-bonded copper(II) complexes [57].
The first species formed by dimeric peptide dNGF(1-15) is [CuLH5] that reaches its maximum percentage at pH 4.5 (Figure 2).
The stability constant (logK(115) = logβ(115) -logβ(015) = 41.18 − 35.12 = 6.06) is indicative of a 2Nim, COO− coordination mode analogous to monomeric peptides and Aβ as above reported. This is further confirmed by UV–vis parameters similar to that of rNGF(14-1) copper complex. It is to note that in the [CuLH5] species there are only three deprotonated centers and therefore the involvement of 2Nim requires that one carboxylate group is still protonated. The unusual high pK value of one carboxylate moiety (pK = 5.28) and the pH range of species existence, between 4.0–5.5, makes this hypothesis plausible.
[CuLH3] is the next species formed with a maximum percentage of formation around pH 5.5 (Figure 2). It is not possible to calculate the stepwise stability constant value as performed for other species because the protonation constant of the [LH3] species was not experimentally obtained. However, the difference with the stability constant of [CuLH5] (logK(112) = logβ(115) − logβ(113) = 9.62) suggests the deprotonation of the carboxylate not coordinated to copper ion and the binding of another nitrogen atom (imidazole or amino) to the metal ion.
[CuLH2] is the predominant species at physiological pH; the stability constant value logK (logK(112) = logβ(112) − logβ(012) = 10.56) is high and can be related with the relevant blue shift in the UV–vis maximum absorption (λmax = 569 nm, ε = 94 M−1 cm−1). All these data are indicative of a strong ligand field around copper ion, determined by four nitrogen atoms coordination mode in a planar arrangement. It can be assumed that the (NH2, N−) five-membered chelate is assisted by the macrochelation with the NIm donor of the His-4 residue of the same chain and the other histidine belonging to the other chain of dimeric peptide.
The CD spectra show an increase in the wide band centered at 324 nm, that include charge transfer signals of both imidazole and deprotonated amide to metal ion, and a d–d transition band with a relatively low intensity compared to the analogous species formed by monomeric peptides. These CD features are in agreement with the involvement of more imidazole side chain in the metal binding (Figure 3).
The next species [CuLH] and [CuL] show similar spectroscopic parameters to those of [CuLH2], suggesting the deprotonation of amino/imidazole not bound to metal ion and afterwards, at strongly basic pH, the deprotonation of an amide nitrogen atom that substitutes one imidazole in the coordination to metal ion.
This entry is adapted from the peer-reviewed paper 10.3390/ijms22105085
