Haemophilus influenzae is an obligate commensal of the upper respiratory tract in humans and may be responsible for upper respiratory tract infections and even meningitis. Seven biologically active H. influenzae dsDNA phages have been currently described: HP1, HP2, HP3, S2A, S2B, S2C, N3 and Mu-like phage φflu. The most studied is the group of HP1/S2 phages. The temperate H. influenzae phage HP1 belongs to the Myoviridae family of phages and infects Rd strains of H. influenzae. HP1 has a head-tail structure. Its genome is about 32 kb long and encodes 41 potential proteins. Early promoters, which control the lysis-versus-lysogeny decision seem to be located toward the 5’ end of the genome. The late promoter is located between orf16 and orf17, and regulates the expression of late genes, including the lys and hol genes, encoding a SAR-endolysin and a pinholin. Endolysin alone is responsible for cell lysis. The control of endolysin activity seems to be related to conformational changes in holin structure. HP1 phages encodes Dam methyltransferase, which role in HP1 life cycle remains unknown.
- HP1 phage, cox protein, temparate phage, lytic sys
HP1 is a temperate bacteriophage, which specifically infects and may lysogenize a human respiratory tract pathogen-Haemophilus influenzae Rd [1]. Historically HP1 phage was used to elucidate the mechanism of natural transformation in H. influenzae [1,2]. HP1 phage is classified in Caudovirales order, Myoviridae family and Peduovirinae subfamily. Electron microscopy has established that HP1 has a head-tail structure [3]. The particle consists of a small icosahedral head (with elongated icosahedral symmetry about 50 × 52 nm, T=7, composed of 72 capsomers), a relatively long and complex contractile tail (about 122 nm). The tail terminates in tail plate with extended tail fibers [3]. The phage particles are fragile and are readily sheared into filled heads or headless tail assemblies [3]. HP1c1 is a derivative of HP1, that gives larger an clearer plaques, but is also able to induce lysogeny [1].
Genome organization
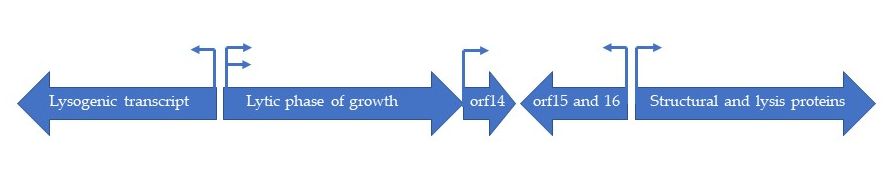
The genome of HP1 is represented by linear, double stranded DNA [4]. It is 32 355 nucleotides long and encodes 41 potential proteins [5]. The GC content is 39 %, which is identical to the GC content of H. influenzae. The complete HP1 genome sequence has been deposited in GenBank under the accession no. U24159. HP1 displays significant similarity to P2 and 186 coliphages concerning gene organization and sequences of the encoded proteins. The HP1 genome contains four strong promoter sequences of the σ70 type and three ρ-independent transcription terminator sequences were found [5]. The genome is organized into five plausible transcriptional units. Esposito et al. (1996) identified regions encoding proteins involved in (i) recombination (lysogenic transcript), (ii) replication (lytic phase of growth), (iii) maturation and packaging of phage genome (orf15 and orf16), (iv) host cell lysis and structural proteins and (v) a transcript encoding for orf14, which product has an unknown function [5] (Figure 1). The most of predicted gene products are similar to those encoded by P2 and 186 coliphages, as well as by the retronphage Ec67 [5].
Figure 1. Schematic organization of five transcriptional subunits in HP1 genome.
Studies on restriction-modification in H. influenzae demonstrated that 3 different types of molecular organization can be detected concerning HP1 phage, both in the viral DNA isolated from the mature phage particles and isolated from prophage [6].
Adsorption and genome ejection
The receptor for HP1 phage is the bacterial lipooligosaccharide and precisely the first two sugars (Glucose-Galactose) in globotetraose extending from the third heptose [7].
Upon contraction, the external contractile envelope of tail (tail sheath) relaxes and slides along the tail core (tail tube), resulting in ejection of the viral genome into host cell cytoplasm. After infection, the HP1 genome circularizes thanks to its cohesive ends (cos). Cos are composed of complementary 5' single-stranded extensions which are seven residues long. The left cos contains only pyrimidines and the right only purines [4].
Lysis-versus-lysogeny decision
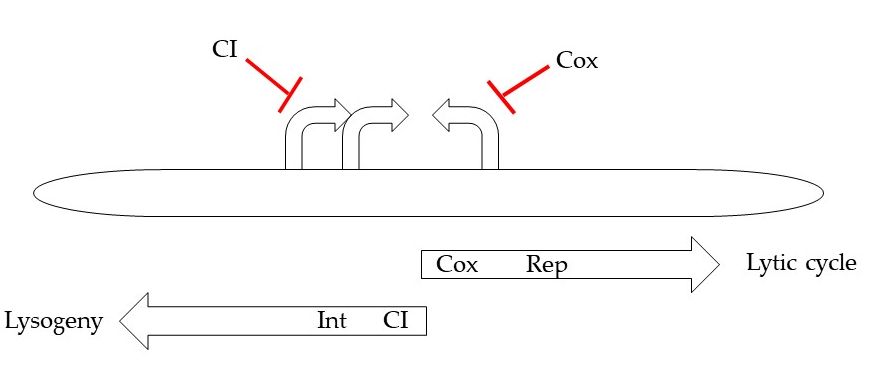
Tight regulation of the switch between lysogenic and lytic pathways is observed. The switch seems to be controlled by the transcription process from three early promoters, which are located near the 5’ end of the HP1 genome (Figure 2). One leftward and two rightward early promoters transcribe HP1 CI and HP1 Cox, which have genetic and functional homology to transcriptional regulators found in λ phage [8]. Face-to face overlapping promoters organized the open reading frames into two operons transcribed in opposite directions: lytic operon from the PR1 and PR2 promoters and lysogenic operon from the PL promoter [5].
Figure 2. Schematic organization of early region involved in lysis/lysogeny decision.
As mentioned, HP1 phage encodes two transcription factors: multifunctional regulatory protein HP1 Cox and lysogenic repressor HP1 CI. The PL promoter controls expression of CI (which have functional homology to transcriptional regulator CI from λ phage), several short ORFs and HP1 integrase, while the PR1 and PR2 promoters allow the expression of Cox regulator and of several other genes, that products are involved in the early lytic pathway. Cox appears to function as an analogue of the λ phage Cro protein [8]. Cox represses the production of transcripts from PL promoter, while CI protein repressed lytic transcription from PR. In the presence of CI protein, transcription from PR is shut down, while PL will continue to function, producing more CI protein and maintaining the lysogenic state. The PL transcript also contains the int gene encoding protein necessary for genome integration that is similar in amino acid sequence to coliphage 186 integrase [9, 10].
In the lysogenic state, the HP1 genome is inserted into the host chromosome by recombination reaction. Two probable attB sites were found within H. influenzae chromosome [11, 12]. One of them coincides with the sequence of a tRNA gene, and HP1 is one of a diverse group of integrating elements targeted to tRNA genes [13]. The integration does not require ATP hydrolysis but is stimulated by bacterial integration host factor (IHF) [9, 10]. The substrates for integration are the 500-bp phage attachment site, attP and an 18-bp host sites, attB [13]. Once recombination takes place, the integrated prophage is flanked by recombinant attL and attR sites, which serve as the substrates for excisive recombination during induction of lytic cycle in bacteria containing prophage [14]. During the integration of viral DNA into bacterial chromosome, the participation of integration host factors (IHF) are suspected [15].
Other probable regulatory proteins were identified, based on their amino-acid sequence similarity to proteins of P2 or 186 phages. For example, the orf5 product of HP1 shares some homology with the 186 CII protein and may be involved in enhancement of lysogeny [5].
Multifunctional Cox regulator
Cox protein not only inhibit PL promoter, but is also involved in DNA recombination during prophage excision from bacterial chromosome. Excisive recombination, like integration, requires HP1 int product and bacterial IHF but also requires the presence of Cox regulator [14]. The formation of properly arranged protein-DNA structures, termed intasomes (in which the two substrate DNA molecules are held together by integrase and accessory proteins), is the fundamental step in site-specific recombination [16]. Cox factor has the capacity to control the direction of DNA recombination reaction: Cox inhibits integrative recombination of attP and attB, but activates excision of the HP1 genome from its site within the host chromosome [14]. In λ phage, the inhibition of phage genome integration and the prophage excision are mediated by the additional λ - encoded protein - Xis [17]. Cox acts as tetramer but may also form octamers by self-assembling. Cox binds two directly repeated sites present in attP sequences. Cox binding interferes with the integrase binding to the DNA. The competition between these two proteins is crucial for control integration and excision of HP1 genome. Cox probably plays a positive role in assembling of the excision-competent nucleoprotein complexes on attL site [8, 16].
DNA replication
It was demonstrated that HP1c1 DNA replication takes place on the cell membrane where a concatemer of phage DNA is first synthesized and subsequently trimmed to phage-equivalent DNA. Phage-equivalent DNA may be released from the membrane for packaging or is packaged into heads while associated to the membrane [18]. DNA replication was studied for HP1-related phages P2 and 186. Their genome replication is unidirectional and follows a modified rolling circle mechanism.
Predicted 90 kDa protein HP1 Rep is involved in phage DNA replication. It initiates the replication probably by inducing the nick at origin of replication that is included in rep gene [5]. The HP1 Rep is similar in size and sequence to the P2 phage A protein, the 186 phage CP87 protein and the Ec67 retronphage ORF2 protein [5].
Promoter elements for late genes
In HP1, the genes encoding structural elements appear to lack a σ70-like promoter. The late promoter is located between orf16 and orf17, and regulates the expression of late genes [5]. Very few intergenic regions exist in these part of genome.
Lysis system
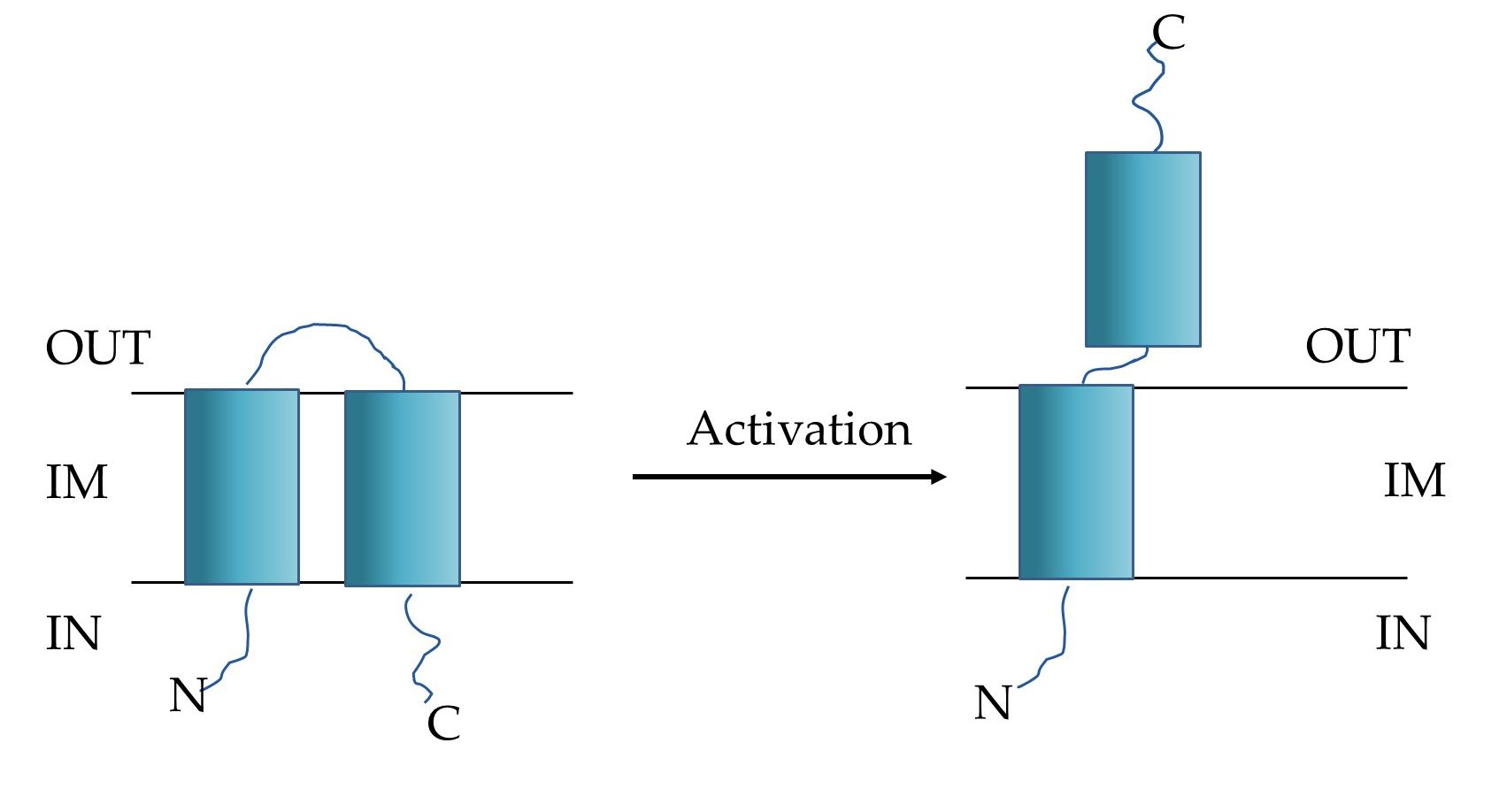
HP1 lytic system seems to be encoded by at least two proteins: products of lys and hol genes. The analysis of the amino acid sequence of the lys gene product suggested that HP1 endolysin belongs to the family of signal-arrest-release (SAR) endolysins, that reach the peptidoglycan without holin presence. Indeed, HP1 endolysin induced lysis on its own when cloned and expressed in Escherichia coli, and the new phage release from H. influenzae cells was suppressed by inhibition of the secretion (sec) pathway with sodium azide [19]. It was also demonstrated that protein encoded by hol gene is a transmembrane and has with unusual for holins C-out and N-in topology, when overexpressed/activated (Figure 3). Studies on pores formation by HP1 holin demonstrated that this protein did not allow the formation of large pores, allowing the leakage of β-galactosidase. However, expression of hol gene in E. coli caused cell death (decrease in viable cell count) without lysis (turbidity remained constant). New phage release from the natural host was inhibited by deletion of the hol gene. The hol gene product is in fact a pinholin and probably regulates the time of lysis [19].
Figure 3. Schematic model of HP1 pinholin (OUT: periplasm, IN: cytoplasm, IM: inner membrane). Left: the inactive, non-lethal form, right: the active-lethal form [19].
Dam methyltransferase
The product of orf13 encodes Dam methyltransferase, that recognizes the 5'-GATC-3' sequence and methylates the adenine residue on ss or dsDNA [20]. The amino-acid sequence of the protein shows some homology to Dam methyltransferase encoded by bacteriophage T1. The order of subdomains in HP1 and T1 Dam enzymes corresponds to the γ-class of m6A methyltransferases, in contrast to the α-class encoded by Bacteria and T-even phages [21]. Moreover, HP1 Dam lacks the motif responsible for binding of S-adenosylmethionine, the donor of methyl group [20].
The host, H. influenzae also encodes a enzyme with Dam activity. Bacterial and phage Dam methyltransferases are distant evolutionary, but a common ancestor could not be excluded [21]. The role of viral Dam methyltransferase in the HP1 life cycle is still unknown, but coliphages P2 and 186, related to HP1, do not encode the Dam activity.
Resistance to phage infection
The HP1 host, H. influenzae modulates the activity of its defense systems against phage infection by phase variation. Variation in sensitivity/resistance to phage infection correlates with changes in lipooligosaccharide (LOS) structure, due to slippage at CAAT repeats, present within a gene involved in LOS synthesis (lic2A). The lack of bacterial Dam methyltransferase increases the frequency of switching from sensitivity to resistance concerning HP1 infections [7].
Similar phages
Several phages infecting H. influenzae are described in the literature (HP1, S2A, S2B, S2C, N3, HP2, HP3 and Mu-like phage φflu) [1,3,5,22].
The HP2 phage appears to be closely related to the HP1 phage. Its genome is 31.5 kb (1 kb smaller than that of HP1), and it contains 36 ORFs, compared to 41 ORFs in HP1 [22]. Genomic comparisons between HP1 and HP2 suggest a recent divergence. However, HP2 does not infect or lysogenize H. influenzae Rd strains and was isolated from invasive H. influenzae NTHI strain [22].
Also S2 phage types A, B and C share homology in genomic sequence to HP1 phage [23]. Closer examination suggests that S2 type C may be the original HP1 [24]. However, it is possible to replicate S2 phage on the lysogen of the HP1 (with reduced efficacity), suggesting that the immunity systems are to some extent similar but not identical [3, 23]. As HP1, S2 phages lysogenize H. influenzae Rd strains.
Literature
- Harm, W. and C. S. Rupert. "Infection of transformable cells of Haemophilus influenzae by bacteriophage and bacteriophage DNA." Z Vererbungsl 94 (1963): 336-48. 10.1007/bf00897593.
- Stuy, J. H. "Mechanism of Haemophilus influenzae transfection by single and double prophage deoxyribonucleic acid." J Bacteriol 144 (1980): 1003-8.
- Boling, M. E., D. P. Allison and J. K. Setlow. "Bacteriophage of Haemophilus influenzae. 3. Morphology, DNA homology, and immunity properties of HPlcl, S2, and the defective bacteriophage from strain Rd." J Virol 11 (1973): 585-91.
- Fitzmaurice, W. P., A. S. Waldman, R. C. Benjamin, P. C. Huang and J. J. Scocca. "Nucleotide sequence and properties of the cohesive DNA termini from bacteriophage HP1c1 of Haemophilus influenzae Rd." Gene 31 (1984): 197-203. 10.1016/0378-1119(84)90210-5.
- Esposito, D., W. P. Fitzmaurice, R. C. Benjamin, S. D. Goodman, A. S. Waldman and J. J. Scocca. "The complete nucleotide sequence of bacteriophage HP1 DNA." Nucleic Acids Res 24 (1996): 2360-68. 10.1093/nar/24.12.2360.
- Piekarowicz, A., R. Brzeziński, M. Smorawińska, L. Kauc, K. Skowronek, M. Lenarczyk, M. Gołembiowska and M. Siwińska. "Major spontaneous genomic rearrangements in Haemophilus influenzae S2 and HP1c1 bacteriophages." Gene 49 (1986): 111-8. 10.1016/0378-1119(86)90390-2.
- Zaleski, P., M. Wojciechowski and A. Piekarowicz. "The role of dam methylation in phase variation of Haemophilus influenzae genes involved in defence against phage infection." Microbiology 151 (2005): 3361-69. 10.1099/mic.0.28184-0.
- Esposito, D., J. C. Wilson and J. J. Scocca. "Reciprocal regulation of the early promoter region of bacteriophage HP1 by the Cox and Cl proteins." Virology 234 (1997): 267-76. 10.1006/viro.1997.8646.
- Goodman, S. D. and J. J. Scocca. "Nucleotide sequence and expression of the gene for the site-specific integration protein from bacteriophage HP1 of Haemophilus influenzae." J Bacteriol 171 (1989): 4232-40. 10.1128/jb.171.8.4232-4240.1989.
- Hakimi, J. M. and J. J. Scocca. "Purification and characterization of the integrase from the Haemophilus influenzae bacteriophage HP1; identification of a four-stranded intermediate and the order of strand exchange." Mol Microbiol 21 (1996): 147-58. 10.1046/j.1365-2958.1996.6311351.x.
- Skowronek, K. "Identification of the second attachment site for HP1 and S2 bacteriophages in Haemophilus influenzae genome." Acta Microbiol Pol 47 (1998): 7-17.
- Kauc, L., K. Skowronek and S. H. Goodgal. "The identification of the bacteriophage HP1c1 and S2 integration sites in Haemophilus influenzae Rd by field-inversion gel electrophoresis of large DNA fragments." Acta Microbiol Pol 40 (1991): 11-26.
- Hauser, M. A. and J. J. Scocca. "Location of the host attachment site for phage HPl within a cluster of Haemophilus influenzae tRNA genes." Nucleic Acids Res 18 (1990): 5305. 10.1093/nar/18.17.5305.
- Esposito, D. and J. J. Scocca. "Identification of an HP1 phage protein required for site-specific excision." Mol Microbiol 13 (1994): 685-95. 10.1111/j.1365-2958.1994.tb00462.x.
- Astumian, J. H., A. S. Waldman and J. J. Scocca. "Site-specific recombination between cloned attp and attb sites from the Haemophilus influenzae bacteriophage HP1 propagated in recombination-deficient Escherichia coli." J Bacteriol 171 (1989): 1747-50. 10.1128/jb.171.3.1747-1750.1989.
- Esposito, D., J. S. Thrower and J. J. Scocca. "Protein and DNA requirements of the bacteriophage HP1 recombination system: A model for intasome formation." Nucleic Acids Res 29 (2001): 3955-64. 10.1093/nar/29.19.3955.
- Better, M., S. Wickner, J. Auerbach and H. Echols. "Role of the xis protein of bacteriophage lambda in a specific reactive complex at the attr prophage attachment site." Cell 32 (1983): 161-8. 10.1016/0092-8674(83)90506-8.
- Barnhart, B. J. and S. H. Cox. "DNA replication of induced prophage in Haemophilus influenzae." J Virol 12 (1973): 165-76.
- Adamczyk-Popławska, M., Z. Tracz-Gaszewska, P. Lasota, A. Kwiatek and A. Piekarowicz. "Haemophilus influenzae HP1 bacteriophage encodes a lytic cassette with a pinholin and a signal-arrest-release endolysin." Int J Mol Sci 21 (2020): 10.3390/ijms21114013.
- Piekarowicz, A. and J. Bujnicki. "Cloning of the dam methyltransferase gene from Haemophilus influenzae bacteriophage HP1." Acta Microbiol Pol 48 (1999): 123-9.
- Bujnicki, J. M., M. Radlinska, P. Zaleski and A. Piekarowicz. "Cloning of the Haemophilus influenzae dam methyltransferase and analysis of its relationship to the dam methyltransferase encoded by the HP1 phage." Acta Biochim Pol 48 (2001): 969-83.
- Williams, B. J., M. Golomb, T. Phillips, J. Brownlee, M. V. Olson and A. L. Smith. "Bacteriophage HP2 of haemophilus influenzae." J Bacteriol 184 (2002): 6893-905. 10.1128/jb.184.24.6893-6905.2002.
- Skowronek, K. and S. Baranowski. "The relationship between HP1 and S2 bacteriophages of Haemophilus influenzae." Gene 196 (1997): 139-44. 10.1016/s0378-1119(97)00219-9.
- Skowronek, K., A. Piekarowicz and L. Kauc. "Comparison of HP1c1 and S2 phages of Haemophilus influenzae." Acta Microbiol Pol 35 (1986): 227-32.
This entry is adapted from the peer-reviewed paper 10.3390/ijms21114013