Metabolomics is used to reduce the complexity of plants and to understand the underlying pathways of the plant phenotype. The metabolic profile of plants can be obtained by mass spectrometry or liquid-state NMR. Extraction of metabolites from the sample is necessary for both techniques to obtain the metabolic profile. This extraction step can be eliminated by making use of high-resolution magic angle spinning (HR-MAS) NMR which allows to get metabolic profile directly in intact plant tissues such as intact leaves. An HR-MAS NMR-based metabolomics workflow is thus established that provide a novel platform for obtaining important information of regular metabolic network non-invasively.
- metabolomics
- plants
- HR-MAS NMR
- multivariate analysis
1. Introduction
To understand the biological pathway underlying the phenotype of plants, a systems biology approach can be used [1,2,3]. In systems biology, the information and interaction of the functional physical structure and the genetic information are integrated to provide a comprehensive model of the organism (Figure 1). Different high-throughput technologies are used to study the genetic program of the various -omics fields: genomics, transcriptomics, proteomics, and metabolomics.
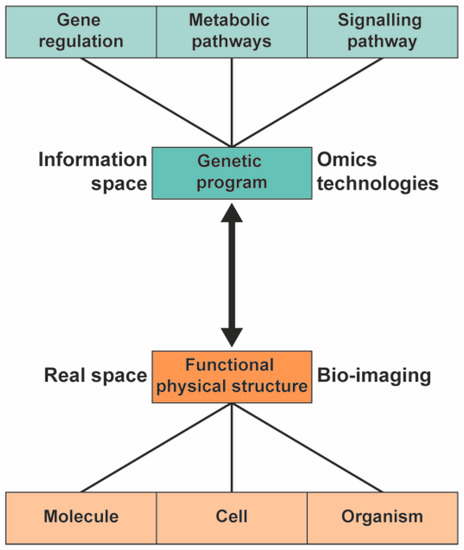
Figure 1. In systems biology, the information from the genetic program is integrated with information from functional physical structures to provide a comprehensive model of plants.
Metabolomics was the newest field added to the systems biology toolbox at the beginning of the 21st century. Metabolomics gives a quantitative and qualitative overview of all the metabolites, small molecules with a molecular weight of 30–3000 Da, present in an organism with various properties and functions [4]. There are approximately 1,000,000 different metabolites available in the plant kingdom, which makes metabolomics a challenging field [5]. Moreover, the metabolome changes quite quickly due to circadian rhythm [6,7,8] and environmental stresses [9,10] and differs between organs, tissues and even for single cells [11,12]. The metabolome is most closely related to the phenotype of a plant since metabolites are the end products of cellular processes [13]. Metabolomics is used to study development under normal and abiotic conditions (temperature, light, salt) [14] and biotic stress conditions (fungal, insects) [15,16], the safety assessment of genetically modified crops [17], speed up crop improvements [18], the effect of fruit storage [19] and the detection of food fraud [20,21].
The link between the gene regulatory network and the functional physical structure (the double arrow in Figure 1) is generally considered highly complex, with many pathways and pathway nodes interacting in what are often considered multifactorial processes. While it is undoubtedly flexible and adaptable to environmental constraints, the underlying links for a specific phenotype may turn out to be monofactorial, particularly in plants that can be grown under highly controlled conditions. The ultimate goal is to understand the complexity of organisms using metabolomics and to understand the underlying pathways of the phenotype of the organisms in a general framework [22].
2. Analytical Techniques in Metabolomics
To study the metabolic profile of a plant, mass spectrometry (MS) or liquid-state nuclear magnetic resonance (NMR) spectroscopy are the most common techniques in metabolomics. Both techniques have their own advantages and limitations, as shown in Table 1. NMR spectroscopy is a method which is non-destructive, with a high reproducibility and allows to quantify metabolites. On the other hand, while MS is more sensitive, allowing to detect more metabolites in a sample, it needs different chromatography techniques such as gas chromatography (GC) or liquid chromatography (LC) for different classes of metabolites [1,2].
Table 1. The advantages and limitations of NMR spectroscopy and mass spectrometry for metabolic profiling [2,3,4,5].
| NMR spectroscopy | Mass Spectroscopy | |
|---|---|---|
| Sensitivity | Low sensitivity, but can be improved with higher field strength and cryo- or microprobes | High sensitivity, can reach the detection limit of attomolar (10–18) concentrations |
| Sample measurement | In one measurement with a detectable concentration can be detected | Need chromatography techniques for different classes of metabolites |
| Sample recovery | Non-destructive technique Several analyses can be performed on the same extracted sample |
Destructive technique |
| Reproducibility | Very high | Moderate |
| Quantification | Absolute quantitation of metabolites possible by adding one standard with known concentration | Quantification is possible with authentic standards, which are not available for newly identified compounds. Ionisation efficiencies, ion suppression and matrix effects have influences on the concentration. |
| Targeted or untargeted approach | Untargeted and targeted approach | Untargeted and targeted approach, mainly used for targeted analysis |
For both techniques, the extraction of metabolites from the sample is necessary to obtain the metabolic profile. The drawback of this extraction is that it is not only time-consuming, but also that metabolites might be lost or degraded during extraction [23]. One way to eliminate the extraction procedure is to use high-resolution magic angle spinning (HR-MAS) NMR, which allows using intact tissue samples [24,25,26].
2. HR-MAS NMR-Based Workflow
Here, authors will explain in more detail an HR-MAS NMR-based workflow and apply the workflow to plant material. The HR-MAS NMR-based workflow is shown in Figure 3. The workflow starts with the harvesting of the leaves from plants for the preparation of a sample in the rotor, followed by performing the HR-MAS NMR experiments. The pulse sequences which can be used in metabolomics are described in Section 5. The data are pre-processed and reduced by bucketing (Section 6). Multivariate analysis is executed in three steps: the detection of outliers, investigation of the variation between different samples, and the selection of potential biomarker candidates (Section 7). Finally, the biomarkers quantification and biological interpretation is explained in a comprehensive systems biology approach by using available information from the literature. This workflow is based on a recently established liquid-state NMR approach [10]. The information about pulse sequences, the pre-processing of the data and multivariate analysis is also applicable to liquid-state NMR data. The advantage of using HR-MAS NMR spectroscopy on leaves is that experiments can be genuinely performed in vivo, which will be illustrated with selected plant metabolomics applications (Section 8). As suggested recently, sample preparations and instrumental setup protocols need to be carefully standardized in order to obtain highly reproducible and reliable data [11].
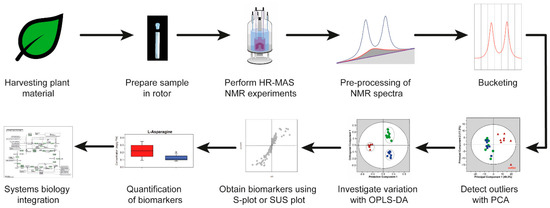
Figure 3. A typical high-resolution magic angle spinning (HR-MAS) NMR-based workflow. OPLS-DA, orthogonal partial least squares discriminant analysis; PCA, principal component analysis; SUS plot, Shared and unique (SUS) plot.
Reference (Editors will rearrange the references after the entry is submitted)
- Kumar, R.; Bohra, A.; Pandey, A.K.; Pandey, M.K.; Kumar, A. Metabolomics for Plant Improvement: Status and Prospects. Front. Plant. Sci. 2017, 8, 1302.
- Emwas, A.H. The strengths and weaknesses of NMR spectroscopy and mass spectrometry with particular focus on metabolomics research. Methods Mol. Biol. 2015, 1277, 161–193.
- Emwas, A.-H.M.; Salek, R.M.; Griffin, J.L.; Merzaban, J. NMR-based metabolomics in human disease diagnosis: Applications, limitations, and recommendations. Metabolomics 2013, 9, 1048–1072.
- Markley, J.L.; Brüschweiler, R.; Edison, A.S.; Eghbalnia, H.R.; Powers, R.; Raftery, D.; Wishart, D.S. The future of NMR-based metabolomics. Curr. Opin. Biotechnol. 2017, 43, 34–40.
- Matsuda, F. Technical Challenges in Mass Spectrometry-Based Metabolomics. Mass Spectrom. 2016, 5, S0052.
- Beckonert, O.; Coen, M.; Keun, H.C.; Wang, Y.; Ebbels, T.M.; Holmes, E.; Lindon, J.C.; Nicholson, J.K. High-resolution magic-angle-spinning NMR spectroscopy for metabolic profiling of intact tissues. Nat. Protoc 2010, 5, 1019–1032.
- Alia, A.; Ganapathy, S.; de Groot, H.J. Magic Angle Spinning (MAS) NMR: A new tool to study the spatial and electronic structure of photosynthetic complexes. Photosynth Res. 2009, 102, 415–425.
- Mazzei, P.; Piccolo, A. HRMAS NMR spectroscopy applications in agriculture. Chem. Biol. Technol. Agric. 2017, 4, 11.
- Vermathen, M.; Marzorati, M.; Vermathen, P. Exploring high-resolution magic angle spinning (HR-MAS) NMR spectroscopy for metabonomic analysis of apples. Chimisty 2012, 66, 747–751.
- Deborde, C.; Moing, A.; Roch, L.; Jacob, D.; Rolin, D.; Giraudeau, P. Plant metabolism as studied by NMR spectroscopy. Prog Nucl Magn Reson Spectrosc 2017, 102, 61–97.
- Flores, I.S.; Martinelli, B.C.B.; Pinto, V.S.; Queiroz, L.H.K., Jr.; Lião, L.A.-O. Important issues in plant tissues analyses by HR-MAS NMR. Phytochem. Anal. 2019, 30, 5–13.
- Kitazaki, K.; Fukushima, A.; Nakabayashi, R.; Okazaki, Y.; Kobayashi, M.; Mori, T.; Nishizawa, T.; Reyes-Chin-Wo, S.; Michelmore, R.W.; Saito, K.; et al. Metabolic Reprogramming in Leaf Lettuce Grown Under Different Light Quality and Intensity Conditions Using Narrow-Band LEDs. Sci. Rep. 2018, 8, 7914.
- Qin, Z.; Liao, D.; Chen, Y.; Zhang, C.; An, R.; Zeng, Q.; Li, X.e. A Widely Metabolomic Analysis Revealed Metabolic Alterations of Epimedium Pubescens Leaves at Different Growth Stages. Molecules 2019, 25, 137.
- Augustijn, D.; Roy, U.; van Schadewijk, R.; de Groot, H.J.; Alia, A. Metabolic Profiling of Intact Arabidopsis thaliana Leaves during Circadian Cycle Using 1H High Resolution Magic Angle Spinning NMR. PLoS ONE 2016, 11, e0163258.
- Wang, L.; Ma, K.-B.; Lu, Z.-G.; Ren, S.-X.; Jiang, H.-R.; Cui, J.-W.; Chen, G.; Teng, N.-J.; Lam, H.-M.; Jin, B. Differential physiological, transcriptomic and metabolomic responses of Arabidopsis leaves under prolonged warming and heat shock. BMC Plant. Biol. 2020, 20, 86.
- Augustijn, D.; Tol, N.V.; van der Zaal, B.J.; de Groot, H.J.M.; Alia, A. High-resolution magic angle spinning NMR studies for metabolic characterization of Arabidopsis thaliana mutants with enhanced growth characteristics. PLoS ONE 2018, 13, e0209695.
- Augustijn, D.; de Groot, H.J.M.; Alia, A. A robust circadian rhythm of metabolites in Arabidopsis thaliana mutants with enhanced growth characteristics. PLoS ONE 2019, 14, e0218219.
- Lucas-Torres, C.; Bernard, T.; Huber, G.; Berthault, P.; Nishiyama, Y.; Kandiyal, P.S.; Elena-Herrmann, B.; Molin, L.; Solari, F.; Bouzier-Sore, A.-K.; et al. General Guidelines for Sample Preparation Strategies in HR-µMAS NMR-based Metabolomics of Microscopic Specimens. Metabolites 2020, 10, 54.
- Kruk, J.; Doskocz, M.; Jodlowska, E.; Zacharzewska, A.; Lakomiec, J.; Czaja, K.; Kujawski, J. NMR Techniques in Metabolomic Studies: A Quick Overview on Examples of Utilization. Appl. Magn. Reson. 2017, 48, 1–21.
- Elena-Herrmann, B. CHAPTER 2 NMR Pulse Sequences for Metabolomics. In NMR-Based Metabolomics; The Royal Society of Chemistry: Cambridge, UK, 2018; pp. 22–38.
- Le Guennec, A.; Tayyari, F.; Edison, A.S. Alternatives to Nuclear Overhauser Enhancement Spectroscopy Presat and Carr-Purcell-Meiboom-Gill Presat for NMR-Based Metabolomics. Anal. Chem. 2017, 89, 8582–8588.
- Dona, A.C.; Kyriakides, M.; Scott, F.; Shephard, E.A.; Varshavi, D.; Veselkov, K.; Everett, J.R. A guide to the identification of metabolites in NMR-based metabonomics/metabolomics experiments. Comput. Struct. Biotechnol. J. 2016, 14, 135–153.
- Vu, T.N.; Laukens, K. Getting your peaks in line: A review of alignment methods for NMR spectral data. Metabolites 2013, 3, 259–276.
- Ludwig, C.; Viant, M.R. Two-dimensional J-resolved NMR spectroscopy: Review of a key methodology in the metabolomics toolbox. Phytochem. Anal. 2010, 21, 22–32.
- Emwas, A.-H.; Saccenti, E.; Gao, X.; McKay, R.T.; Dos Santos, V.A.P.M.; Roy, R.; Wishart, D.S. Recommended strategies for spectral processing and post-processing of 1D (1)H-NMR data of biofluids with a particular focus on urine. Metab. Off. J. Metab. Soc. 2018, 14, 31.
- Liland, K.H. Multivariate methods in metabolomics—From pre-processing to dimension reduction and statistical analysis. Trends Anal. Chem. 2011, 30, 827–841.
This entry is adapted from the peer-reviewed paper 10.3390/molecules26040931
