Bone material strength is determined by several factors, such as bone mass, matrix composition, mineralization, architecture and shape. From a clinical perspective, bone fragility is classified as primary (i.e., genetic and rare) or secondary (i.e., acquired and common) osteoporosis
- bone fragility
- type I collagen
- post-translational modifications
1. Introduction
Developing bones consist of cartilaginous joints, the epiphysis, the growth plate cartilage with adjacent osteogenesis and the cortical and cancellous bone mineralized structure. Bone tissue contains three distinct cell types: (i) the osteoblasts, derived from mesenchymal cells, which deposit new bone tissue; (ii) osteoclasts, derived from bone marrow hematopoietic precursor cells, which break down bone matrix; and (iii) osteocytes (former osteoblasts) which orchestrate the activity of osteoblasts and osteoclasts as a response to mechanical strain [1]. The extracellular matrix of bone tissue is composed of inorganic minerals, collagen, water, non-collagenous proteins and lipids. Bone fragility can originate from alterations in all these components, be it the genetic blueprint, mechanical loading, insufficient remodeling at old age, estrogen deficiency or chronic medical conditions affecting bone accrual, structure or composition.
Traditionally, bone fragility is understood as resulting from reduced bone mass, or from defects in bone matrix composition or mineralization. Medical research has unveiled many monogenic bone fragility conditions, yet their mechanisms of disease remain incompletely understood. In fact, they continue holding secrets which may open doors for drug developments in rare and common osteoporosis.
2. Genetic Causes of Bone Fragility
2.1. Primary Osteoporosis Affecting Collagen (Osteogenesis Imperfecta)
2.1.1. Clinical Symptoms and Classification
Recurrent limb and vertebral fractures are typical for patients with osteogenesis imperfecta (OI). OI, also known as brittle bone disease, is an inherited disorder of connective tissue that has a wide clinical and genetic heterogeneity. OI is a rare disorder, with an incidence of one in 10,000–20,000 births [2]. From a bone material perspective, OI is characterized by low bone mass and increased bone mineralization density, which causes brittleness (Figure 1), recurrent fractures and skeletal deformities, but also extra-skeletal manifestations. The latter include blue sclerae, dentinogenesis imperfecta, joint laxity, hearing loss as well as cranial malformations (i.e., basilar invagination) and pulmonary hypoplasia with reduced lung capacity in severe cases. Depending on clinical severity, mobility is mildly to severely impaired. Fractures are particularly common in childhood but increased fracture risk persists throughout life.
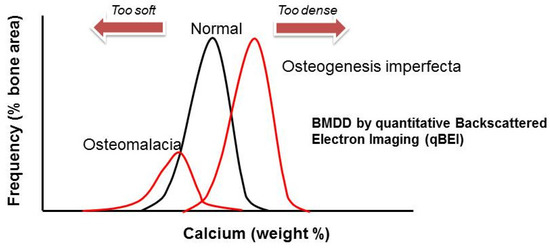
Figure 1. The extremes of bone mineralization density distribution (BMDD): bone tissue in patients with OI has increased mineralization density. This is due to the irregular collagen fibers and their wider spatial distribution which allows more hydroxyapatite deposition. Also depicted is the opposite: low tissue mineralization density in patients with osteomalacia.
2.1.2. Genetic Classification and Protein Function in OI
One recent classification [2] proposes five groups of subtypes for OI (Group A–E). Group A subtypes of OI (Type I–IV, XIII), which are caused by defects in collagen synthesis, structure or processing of COL1A1 and COL1A2, including their C-terminal propeptide cleavage by BMP1. Group B (type VII, VIII, IX and XIV) contains the genes that play a key role in post-translational modification of type I collagen, which are CRTAP, LEPRE1, PPIB and TMEM38B. Group C (type X, XI) includes the genes that are responsible for collagen folding or cross linking, which are SERPINH1, FKBP10, PLOD2 and P4HB. Group D (type V and VI) encompasses the genes IFITM5 and SERPINF1 that modify mineralization. Group E (type XII, XV, XVI) involves genes that cause defects in osteoblast differentiation, which are SP7, WNT1 and CREB3L1. Figure 2 illustrates the molecular target of all known OI types, their locations in or out of the cells, and which protein products interact with collagen. The figure also depicts the affected cellular and extracellular process, including collagen synthesis, structure and assembly, collagen post-translational modification and processing.
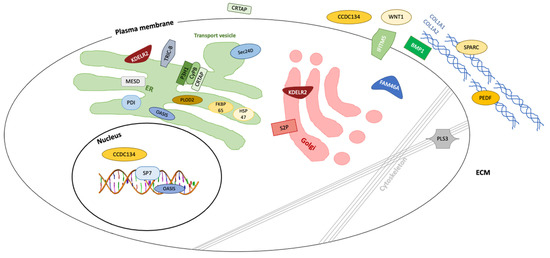
Figure 2. Osteogenesis imperfecta is caused by gene mutations encoding proteins involved in collagen biosynthesis, bone homeostasis and maintenance.
All 24 OI-causing genetic entities have in common that they, by various mechanisms, affect the quality or quantity of type I collagen [2]. Since type I collagen is the main structure protein (94%) of pre-mineralized bone matrix (osteoid), abnormal type I collagen synthesis decreases bone mass and increases susceptibility to fracture. Type I collagen consists of three modified chains that form a triple helical fibril with two identical α1 (I) chains and one structurally similar but genetically different α2 (I) chain. The α chain is characterized by a strict pattern which contains a multiple triple sequence of a Gly (glycine)-X (normally proline)-Y (usually hydroxyproline). Notably, about a third of the residues are prolines that get hydroxylated. Each chain is characterized by amino- and carboxy-propeptides, which are critical in preventing self-assembly of collagen fibrils within the cells.
In the cells, type I collagen is synthesized as a soluble precursor molecule, procollagen with N-terminal and C-terminal propeptides that flank the helical domain. The biosynthesis of type I procollagen is a multistep process that involves an ensemble of proteins for post-translational modifications, folding, transport, secretion and quality control [3]. Procollagen synthesis is started up in the nucleus of collagen producing cells, such as osteoblasts and fibroblasts. The DNA segment is transcribed to precursor RNA that is spliced to mRNA and then transported to the endoplasmic reticulum (rER), via the N-terminal signal peptide, where it is translated to propolypeptide chains termed pro-α chains. In the rER, these propolypeptide chains undergo a series of post-translational modifications described in great detail elsewhere [3][4][5]. The C-terminal propeptide of each chain is attached to the rER membrane and folds into a structure that is stabilized by intra-chain disulfide bonds, which enables the selection and assembly of the correct chains into a triple helix.
2.1.3. Pathway-Specific Therapy
Neutralizing antibody against sclerostin (Scl-AB), a molecule produced by osteocytes to inhibit osteoblasts-mediated bone formation via the WNT signaling cascade, increased bone formation rate and bone mass in an OI preclinical mouse model [6].
Recently, preclinical studies demonstrated the potential of MSC transplantation before and after birth for severe types of OI (types II/III, severe type IV); this treatment approach is currently tested in the ongoing multicenter clinical trial boost brittle bones before birth (BOOSTB4) [7]. However, one major hurdle to such a cellular therapy is low engraftment of cells in all skeletal elements.
2.2. Primary Osteoporosis Caused by WNT-Signaling Pathway Defects
The Wingless-type mouse mammary tumor virus (MMTV) integration site family 1 (WNT1) protein belongs to a family of 19 secreted signaling glycoproteins. WNTs bind with the frizzled receptor and the co-receptors’ low-density lipoprotein receptor-related protein (LRP)-5 and -6 and activate the β catenin signal transduction pathway in various tissues, including bone [8][9]. This in turn leads to the translocation of β catenin into the nucleus, where it induces the expression of genes that regulate osteoblast differentiation [10]. Thus, the WNT signaling pathway controls mature osteoblast differentiation [11], bone development and bone maintenance [12]. Several groups have shown in patients from different countries that bi-allelic WNT1 mutations are associated with moderate to severe cases of recessive OI type XV [13][14]. In addition, heterozygous pathogenic mutations in WNT1 can cause the clinical picture of primary osteoporosis.
Patients that harbor WNT1 bi-allelic nonsense, missense, splice site substitutions, deletion and frameshift mutations display severe bone fragility, reduced bone mass, multiple fractures and growth delay [14][15]. The structural bone phenotype includes low bone turnover with an imbalance between formation and resorption. This imbalance may be mediated through WNT1 function in osteocytes [16]. Similar to the human nonsense WNT1 mutation in exon 3 (c.565G > T, p.Glu189*), mice harbor a single nucleotide deletion wnt1 mutation in exon 3 (c.565delG, p.Glu189Argfs*10) [13][17]. This mouse model shows typical OI features including severe osteopenia, spontaneous fractures, reduced bone strength and impaired matrix mineralization, mimicking the human disease. Notably, bone forming osteoblast function is impaired, while bone resorbing osteoclast function is not changed in this mouse model [18].
WNT1 mutations arrest the downstream intracellular signaling cascade and nuclear translocation of β catenin and the expression of the regulated genes [10][13]. Loss of function of β catenin results in osteochondroprogenitor cells differentiating into chondrocytes instead of osteoblasts. On the other hand, gain of function of WNT signaling result in enhancement of osteoblast differentiation in vitro [19]. In addition, the WNT signaling cascade in osteocytes plays a critical role in regulating bone cell homeostasis [20]. Mice deficient with β catenin in osteocytes show bone loss [20].
Similar to WNT1, biallelic loss of function mutations in its co-receptor LRP5 cause osteoporosis-pseudoglioma syndrome, which is characterized by severe bone fragility and ocular manifestations [21], and monoallelic LRP5 mutations, which cause primary, autosomal dominant osteoporosis [22]. Hence, there appears to be a gene dosing effect in defective WNT signaling. Moreover, patients with gain of function and loss of function mutations in the LRP5 have high or low bone mass disorders resulting from constitutive activation or decreased osteoblast activity, respectively [14][23]. This fact further supports the notion of a gene dosing effect and makes elements of the WNT signaling pathway attractive drug targets. Trials have been conducted in adult osteoporosis, which led to the market approval of antibody therapy against sclerostin, an inhibitor of the WNT signaling pathway [24]. Trials in children are awaited.
2.3. Primary Osteoporosis Caused by Defects in the TGF-β Pathway
Polypeptides in the transforming growth factor β (TGF-β) family are involved in controlling cell activity and metabolism in bone and cartilage tissues. After TGF-β release from the ECM, it interacts with a receptor complex containing type I (TβRI, TGFBR1) and type II (TβRII, TGFBR2) subunits. The highly complex pathway involves intracellular signal transduction through cytoplasmic proteins, belonging to transcription factors from the SMAD family. A variety of heritable skeletal conditions is associated with dysregulated TGF-β signaling, including Camurati-Engelmann disease [MIM: 131300] [25][26][27][67], Marfan syndrome [MIM: 154700] [26] and Loeys-Dietz syndrome [MIM: 613795] [27] but also OI [28]. Bone fragility is a distinct phenotypic feature of Loeys-Dietz syndrome, caused by mutations in SMAD3.
2.4. Primary Osteoporosis Caused by RANKL/RANK/OPG Defects: TNFRSF11B (Juvenile Paget Disease) and TNFRSF11A (Familial Expansile Osteolysis)
The Receptor Activator of Nuclear Factor Kappa B (RANK, TNFRSF11A) and its ligand RANKL play a key role in osteoclast activation and differentiation [29]. RANKL is a cytokine expressed by osteoblast lineage cells, including osteoblasts and osteocytes. RANKL binds with its cognate receptor RANK on the surface of osteoclast precursors, which activates cell differentiation. Hence, RANKL is essential for formation and activation of osteoclasts. Mice and humans lacking RANKL display complete abrogation of osteoclastogenesis. Osteoblast also express a decoy receptor osteoprotegerin (OPG, TNFRSF11B) that competitively binds with the RANK receptor and inhibits the interaction with RANKL. Hence, osteoblasts and osteocytes regulate activation and differentiation of osteoclasts by the RANKL-RANK axis signaling pathway [29][30].
A recessive deletion mutation in TNFRSF11B encoding the decoy receptor OPG results in unopposed RANK activation and permanently elevated bone turnover, a condition called Juvenile Paget’s disease (JPD; [MIM: 602080]) [31]. JPD is a rare autosomal recessive disease which develops during infancy and early childhood, and worsens in adolescence with pain from debilitating fractures and deformities caused by highly accelerated bone turnover of the entire skeleton [32]. JPD also has extra-skeletal manifestations, including bowing deformities and fractures, contractures as well as short stature [33]. Patients demonstrate histopathological evidence of high bone turnover and weak, disorganized woven bone [34][35].
Dominant gain of function mutations in TNFRSF11A cause familial expansile osteolysis [FEO; MIM: 174810], leading to permanent activation of RANK. The ensuing progressive osteoclastic resorption is associated with medullar expansion and severe pain, disabling deformities and pathological fractures. Characteristically, FEO is accompanied by deafness and loss of dentition as a result of middle ear and jaw bone abnormalities, and variably raised serum alkaline phosphatase levels. FEO cases present with osteolytic lesions in long bones, whereas JPD patients tend to present with trunk and skull lesions [36].
2.5. Bone Fragility in Hajdu Cheney Syndrome
In bone, NOTCH signaling is involved in the regulation of bone formation and bone resorption [37]. In vitro studies have suggest that the NOTCH signaling cascade modulates signaling downstream of RANK, hence activating NOTCH2 enhances osteoclasts maturation [38]. NOTCH2 plays a key role in skeletal development [39]. Homozygous deletion of mouse Notch2 results in early embryonic lethality [40]. Heterozygous gain of function mutations in NOTCH2 lead to Hajdu-Cheney syndrome (HCS) [41]. HCS [MIM: 102500] is a rare autosomal dominant disease characterized by severe osteoporosis associated with craniofacial dysmorphism, acroosteolysis and Wormian bones [42]. Nonsense or short deletion mutations in NOTCH2 exon 34 (the last exon) result in an early termination upstream of the Proline-Glutamic acid-Serine-Threonine (PEST) domain, at the end of the protein, which is required for the NOTCH2 receptor ubiquitination and degradation [43]. Hence, these mutations produce a truncated protein lacking the proteolytic degradation domain of NOTCH2 receptors, leading to sustained NOTCH2 activation with increased osteoclastogenesis [44]. Histomorphometric analysis in affected individuals demonstrate increased bone resorption, increased heterogeneity of mineralization and woven bone. Typical features are reduced cortical thickness and low bone mass. Given the increased osteoclast numbers and turnover, affected individuals respond well to bisphosphonate therapy [45].
This entry is adapted from the peer-reviewed paper 10.3390/ijms22020625
References
- Florencio-Silva, R.; Sasso, G.R.; Sasso-Cerri, E.; Simoes, M.J.; Cerri, P.S. Biology of Bone Tissue: Structure, Function, and Factors That Influence Bone Cells. BioMed Res. Int. 2015, 2015, 421746.
- Forlino, A.; Marini, J.C. Osteogenesis imperfecta. Lancet 2016, 387, 1657–1671.
- Marini, J.C.; Forlino, A.; Bachinger, H.P.; Bishop, N.J.; Byers, P.H.; Paepe, A.; Fassier, F.; Fratzl-Zelman, N.; Kozloff, K.M.; Krakow, D.; et al. Osteogenesis imperfecta. Nat. Rev. Dis Primers 2017, 3, 17052.
- Alten, E.D.; Chaturvedi, A.; Cullimore, M.; Fallon, A.A.; Habben, L.; Hughes, I.; O’Malley, N.T.; Rahimi, H.; Renodin-Mead, D.; Schmidt, B.L.; et al. No longer a historical ailment: Two cases of childhood scurvy with recommendations for bone health providers. Osteoporos Int. 2020, 31, 1001–1005.
- Pozzer, D.; Invernizzi, R.W.; Blaauw, B.; Cantoni, O.; Zito, E. Ascorbic Acid Route to the Endoplasmic Reticulum: Function and Role in Disease. Antioxid Redox Signal. 2020.
- Marini, J.C.; Dang Do, A.N. Osteogenesis Imperfecta. In Endotext; Feingold, K.R., Anawalt, B., Boyce, A., Chrousos, G., de Herder, W.W., Dungan, K., Grossman, A., Hershman, J.M., Hofland, H.J., Kaltsas, G., et al., Eds.; MDText.xom, Inc.: South Dartmouth, MA, USA, 2020.
- Gotherstrom, C.; Walther-Jallow, L. Stem Cell Therapy as a Treatment for Osteogenesis Imperfecta. Curr. Osteoporos Rep. 2020, 18, 337–343.
- Baron, R.; Kneissel, M. WNT signaling in bone homeostasis and disease: From human mutations to treatments. Nat. Med. 2013, 19, 179–192.
- Komiya, Y.; Habas, R. Wnt signal transduction pathways. Organogenesis 2008, 4, 68–75.
- Laine, C.M.; Joeng, K.S.; Campeau, P.M.; Kiviranta, R.; Tarkkonen, K.; Grover, M.; Lu, J.T.; Pekkinen, M.; Wessman, M.; Heino, T.J.; et al. WNT1 mutations in early-onset osteoporosis and osteogenesis imperfecta. N. Engl. J. Med. 2013, 368, 1809–1816.
- Westendorf, J.J.; Kahler, R.A.; Schroeder, T.M. Wnt signaling in osteoblasts and bone diseases. Gene 2004, 341, 19–39.
- Nusse, R.; Clevers, H. Wnt/beta-Catenin Signaling, Disease, and Emerging Therapeutic Modalities. Cell 2017, 169, 985–999.
- Keupp, K.; Beleggia, F.; Kayserili, H.; Barnes, A.M.; Steiner, M.; Semler, O.; Fischer, B.; Yigit, G.; Janda, C.Y.; Becker, J.; et al. Mutations in WNT1 cause different forms of bone fragility. Am. J. Hum. Genet. 2013, 92, 565–574.
- Pyott, S.M.; Tran, T.T.; Leistritz, D.F.; Pepin, M.G.; Mendelsohn, N.J.; Temme, R.T.; Fernandez, B.A.; Elsayed, S.M.; Elsobky, E.; Verma, I.; et al. WNT1 mutations in families affected by moderately severe and progressive recessive osteogenesis imperfecta. Am. J. Hum. Genet. 2013, 92, 590–597.
- Makitie, R.E.; Kampe, A.J.; Taylan, F.; Makitie, O. Recent Discoveries in Monogenic Disorders of Childhood Bone Fragility. Curr. Osteoporos Rep. 2017, 15, 303–310.
- Joeng, K.S.; Lee, Y.C.; Lim, J.; Chen, Y.; Jiang, M.M.; Munivez, E.; Ambrose, C.; Lee, B.H. Osteocyte-specific WNT1 regulates osteoblast function during bone homeostasis. J. Clin. Investig. 2017, 127, 2678–2688.
- Thomas, K.R.; Musci, T.S.; Neumann, P.E.; Capecchi, M.R. Swaying is a mutant allele of the proto-oncogene Wnt-1. Cell 1991, 67, 969–976.
- Joeng, K.S.; Lee, Y.C.; Jiang, M.M.; Bertin, T.K.; Chen, Y.; Abraham, A.M.; Ding, H.; Bi, X.; Ambrose, C.G.; Lee, B.H. The swaying mouse as a model of osteogenesis imperfecta caused by WNT1 mutations. Hum. Mol. Genet. 2014, 23, 4035–4042.
- Day, T.F.; Guo, X.; Garrett-Beal, L.; Yang, Y. Wnt/beta-catenin signaling in mesenchymal progenitors controls osteoblast and chondrocyte differentiation during vertebrate skeletogenesis. Dev. Cell 2005, 8, 739–750.
- Kramer, I.; Halleux, C.; Keller, H.; Pegurri, M.; Gooi, J.H.; Weber, P.B.; Feng, J.Q.; Bonewald, L.F.; Kneissel, M. Osteocyte Wnt/beta-catenin signaling is required for normal bone homeostasis. Mol. Cell Biol. 2010, 30, 3071–3085.
- Gong, Y.; Slee, R.B.; Fukai, N.; Rawadi, G.; Roman-Roman, S.; Reginato, A.M.; Wang, H.; Cundy, T.; Glorieux, F.H.; Lev, D.; et al. LDL receptor-related protein 5 (LRP5) affects bone accrual and eye development. Cell 2001, 107, 513–523.
- Korvala, J.; Juppner, H.; Makitie, O.; Sochett, E.; Schnabel, D.; Mora, S.; Bartels, C.F.; Warman, M.L.; Deraska, D.; Cole, W.G.; et al. Mutations in LRP5 cause primary osteoporosis without features of OI by reducing Wnt signaling activity. BMC Med. Genet. 2012, 13, 26.
- Boyden, L.M.; Mao, J.; Belsky, J.; Mitzner, L.; Farhi, A.; Mitnick, M.A.; Wu, D.; Insogna, K.; Lifton, R.P. High bone density due to a mutation in LDL-receptor-related protein 5. N. Engl. J. Med. 2002, 346, 1513–1521.
- Kaveh, S.; Hosseinifard, H.; Ghadimi, N.; Vojdanian, M.; Aryankhesal, A. Efficacy and safety of Romosozumab in treatment for low bone mineral density: A systematic review and meta-analysis. Clin. Rheumatol. 2020, 39, 3261–3276.
- Van Hul, W.; Boudin, E.; Vanhoenacker, F.M.; Mortier, G. Camurati-Engelmann Disease. Calcif. Tissue Int. 2019, 104, 554–560.
- Verstraeten, A.; Alaerts, M.; Van Laer, L.; Loeys, B. Marfan Syndrome and Related Disorders: 25 Years of Gene Discovery. Hum. Mutat. 2016, 37, 524–531.
- Tan, E.W.; Offoha, R.U.; Oswald, G.L.; Skolasky, R.L.; Dewan, A.K.; Zhen, G.; Shapiro, J.R.; Dietz, H.C.; Cao, X.; Sponseller, P.D. Increased fracture risk and low bone mineral density in patients with loeys-dietz syndrome. Am. J. Med. Genet. A 2013, 161A, 1910–1914.
- Grafe, I.; Yang, T.; Alexander, S.; Homan, E.P.; Lietman, C.; Jiang, M.M.; Bertin, T.; Munivez, E.; Chen, Y.; Dawson, B.; et al. Excessive transforming growth factor-beta signaling is a common mechanism in osteogenesis imperfecta. Nat. Med. 2014, 20, 670–675.
- Ming, J.; Cronin, S.J.F.; Penninger, J.M. Targeting the RANKL/RANK/OPG Axis for Cancer Therapy. Front. Oncol. 2020, 10, 1283.
- Boyce, B.F.; Xing, L. Functions of RANKL/RANK/OPG in bone modeling and remodeling. Arch. Biochem. Biophys. 2008, 473, 139–146.
- Whyte, M.P.; Obrecht, S.E.; Finnegan, P.M.; Jones, J.L.; Podgornik, M.N.; McAlister, W.H.; Mumm, S. Osteoprotegerin deficiency and juvenile Paget’s disease. N. Engl. J. Med. 2002, 347, 175–184.
- Polyzos, S.A.; Cundy, T.; Mantzoros, C.S. Juvenile Paget disease. Metabolism 2018, 80, 15–26.
- Grasemann, C.; Unger, N.; Hovel, M.; Arweiler-Harbeck, D.; Herrmann, R.; Schundeln, M.M.; Muller, O.; Schweiger, B.; Lausch, E.; Meissner, T.; et al. Loss of Functional Osteoprotegerin: More Than a Skeletal Problem. J. Clin. Endocrinol. Metab. 2017, 102, 210–219.
- Caffey, J. Familial hyperphosphatasemia with ateliosis and hypermetabolism of growing membranous bone; review of the clinical, radiographic and chemical features. Bull. Hosp. Joint Dis. 1972, 33, 81–110.
- Golob, D.S.; McAlister, W.H.; Mills, B.G.; Fedde, K.N.; Reinus, W.R.; Teitelbaum, S.L.; Beeki, S.; Whyte, M.P. Juvenile Paget disease: Life-long features of a mildly affected young woman. J. Bone Miner. Res. 1996, 11, 132–142.
- Hughes, A.E.; Ralston, S.H.; Marken, J.; Bell, C.; MacPherson, H.; Wallace, R.G.; van Hul, W.; Whyte, M.P.; Nakatsuka, K.; Hovy, L.; et al. Mutations in TNFRSF11A, affecting the signal peptide of RANK, cause familial expansile osteolysis. Nat. Genet. 2000, 24, 45–48.
- Hilton, M.J.; Tu, X.; Wu, X.; Bai, S.; Zhao, H.; Kobayashi, T.; Kronenberg, H.M.; Teitelbaum, S.L.; Ross, F.P.; Kopan, R.; et al. Notch signaling maintains bone marrow mesenchymal progenitors by suppressing osteoblast differentiation. Nat. Med. 2008, 14, 306–314.
- Fukushima, H.; Nakao, A.; Okamoto, F.; Shin, M.; Kajiya, H.; Sakano, S.; Bigas, A.; Jimi, E.; Okabe, K. The association of Notch2 and NF-kappaB accelerates RANKL-induced osteoclastogenesis. Mol. Cell Biol. 2008, 28, 6402–6412.
- Simpson, M.A.; Irving, M.D.; Asilmaz, E.; Gray, M.J.; Dafou, D.; Elmslie, F.V.; Mansour, S.; Holder, S.E.; Brain, C.E.; Burton, B.K.; et al. Mutations in NOTCH2 cause Hajdu-Cheney syndrome, a disorder of severe and progressive bone loss. Nat. Genet. 2011, 43, 303–305.
- Hamada, Y.; Kadokawa, Y.; Okabe, M.; Ikawa, M.; Coleman, J.R.; Tsujimoto, Y. Mutation in ankyrin repeats of the mouse Notch2 gene induces early embryonic lethality. Development 1999, 126, 3415–3424.
- Canalis, E. Clinical and experimental aspects of notch receptor signaling: Hajdu-Cheney syndrome and related disorders. Metabolism 2018, 80, 48–56.
- Narumi, Y.; Min, B.J.; Shimizu, K.; Kazukawa, I.; Sameshima, K.; Nakamura, K.; Kosho, T.; Rhee, Y.; Chung, Y.S.; Kim, O.H.; et al. Clinical consequences in truncating mutations in exon 34 of NOTCH2: Report of six patients with Hajdu-Cheney syndrome and a patient with serpentine fibula polycystic kidney syndrome. Am. J. Med. Genet. A 2013, 161A, 518–526.
- Descartes, M.; Rojnueangnit, K.; Cole, L.; Sutton, A.; Morgan, S.L.; Patry, L.; Samuels, M.E. Hajdu-Cheney syndrome: Phenotypical progression with de-novo NOTCH2 mutation. Clin. Dysmorphol. 2014, 23, 88–94.
- Isidor, B.; Lindenbaum, P.; Pichon, O.; Bezieau, S.; Dina, C.; Jacquemont, S.; Martin-Coignard, D.; Thauvin-Robinet, C.; Le Merrer, M.; Mandel, J.L.; et al. Truncating mutations in the last exon of NOTCH2 cause a rare skeletal disorder with osteoporosis. Nat. Genet. 2011, 43, 306–308.
- Sakka, S.; Gafni, R.I.; Davies, J.H.; Clarke, B.; Tebben, P.; Samuels, M.; Saraff, V.; Klaushofer, K.; Fratzl-Zelman, N.; Roschger, P.; et al. Bone Structural Characteristics and Response to Bisphosphonate Treatment in Children With Hajdu-Cheney Syndrome. J. Clin. Endocrinol. Metab. 2017, 102, 4163–4172.
