The first antibiotic-producing actinomycete (Streptomyces antibioticus) was described by Waksman and Woodruff in 1940. This discovery initiated the “actinomycetes era”, in which several species were identified and demonstrated to be a great source of bioactive compounds. However, the remarkable group of microorganisms and their potential for the production of bioactive agents were only partially exploited. This is caused by the fact that the growth of many actinomycetes cannot be reproduced on artificial media at laboratory conditions. In addition, sequencing, genome mining and bioactivity screening disclosed that numerous biosynthetic gene clusters (BGCs), encoded in actinomycetes genomes are not expressed and thus, the respective potential products remain uncharacterized. Therefore, a lot of effort was put into the development of technologies that facilitate the access to actinomycetes genomes and activation of their biosynthetic pathways. In this review (Antibiotics 2020, 9(8), 494; https://doi.org/10.3390/antibiotics9080494), we mainly focus on molecular tools and methods for genetic engineering of actinomycetes that have emerged in the field in the past five years (2015–2020). In addition, we highlight examples of successful application of the recently developed technologies in genetic engineering of actinomycetes for activation and/or improvement of the biosynthesis of secondary metabolites.
- actinomycetes
- natural products
- genetic engineering
1. Introduction
Actinomycetes are Gram-positive, mostly aerobic, filamentous bacteria of the phylum Actinobacteria [1–4]. They were initially regarded as organisms, which share morphological characteristics with both bacteria and fungi [5,6] and thus, they were considered as transitional forms between the two groups. This is reflected in the name “Actinomycetes” that was derived from Greek “atkis” or “aktin” (means ray) and “mykes” (means fungus) [7,8]. Actinomycetes have adapted to different ecological niches: terrestrial [9–18], aquatic [19–25] and artificial (“manmade”) [26,27]. In many cases, their biological function is indispensable. Some species are involved in decomposing complex mixtures of polymers in dead plants, animals and fungal materials contributing to humus formation and the recycling of biomaterials [28,29]. Actinomycetes can also provide a nitrogen-fixing function in symbiotic associations with plants [30,31] or take part in other complex, ecologically relevant interactions [32–34]. The most known example includes interactions with insects, where the antibiotics produced by actinomycetes protect the associated organism (e.g., termites, ants, beetles, wasps or they larvae) against detrimental microorganisms [35–40].
However, many researchers have drawn attention to actinomycetes mainly because of their biosynthetic potential for the production of secondary metabolites [41–43]. The discovery of the first actinomycete-derived antibiotic (actinomycin) [44] boosted the isolation of actinomycetes from diverse sources and the traditional screening involving the cultivation, extraction of metabolites and disk diffusion-based activity assays and/or chromatographic analysis [43,45–47]. Since the “Golden Age” of antibiotic discovery (1940s–1960s), several antimicrobial and other valuable compounds were discovered from actinomycetes. Many of these molecules have been developed to commercial products, including agrochemicals and pharmaceuticals [14,47–51]. Thus, actinomycetes are one of the most prolific and important sources of bioactive compounds. Since the whole genome sequencing of the first actinomycetes, which was the genome of Streptomyces coelicolor A3 [52], hundreds actinomycetes genomes were sequenced and annotated. These data revealed that the genomes encode several biosynthetic gene clusters (BGCs) for the production of diverse secondary metabolites. However, only a few of the BGCs are expressed at standard laboratory conditions and, therefore, the biosynthetic potential of many actinomycetes remains unexploited [53–60]. Numerous strategies were developed to induce the expression of the BGCs and access the corresponding products.
2. Genetic Engineering of Actinomycetes
The currently used genetic engineering strategies for activation of BGC-expression and production of the respective compounds in actinomycetes include the expression of multiple copies of the whole BGC or factors that are limiting the production, expression of activator genes, deletion of genes encoding repressors of the BGC, refactoring of the BGC by substitution or modification of native regulatory elements (e.g., promoters) and/or expression of the BGC in optimized (e.g., genome-minimized, precursor-optimized), native or heterologous hosts [69–73] (Figure 1 and Figure 2). These in vivo engineering approaches require genetic manipulation that depends on molecular tools for assembling of genetic constructs and methods for their introduction into the host. In the past four decades, several barriers were identified that retarded the process of natural product discovery from actinomycetes. Lack of compatible molecular tools, limited cloning and DNA transfer methods, DNA degradation, genetic instability, high guanine-cytosine content (GC-content) of the genomes, and different codon usage of the newly introduced foreign DNA are barriers that often prevent the genetic manipulation of actinomycetes (Figure 1).
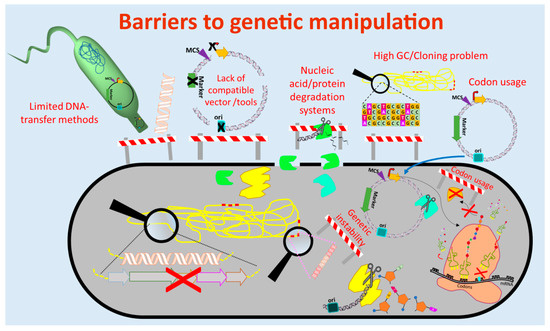
Figure 1. Barriers to genetic manipulation of actinomycetes. Obstacles hindering the genetic manipulation and engineering of actinomycetes include the lack of strain-compatible tools, cloning methods for high guanine-cytosine content (GC-content) DNA sequences as well as transfer methods for introduction of the genetic constructs into the host. Furthermore, nucleic acid degradation systems present in actinomycetes and differences in codon usage may result in fragmentation of the genetic constructs and production of non-functional proteins, respectively. (Single elements of the figure (e.g., ribosome) were re-used from a previous publication [74] with permission from the Royal Society of Chemistry.)
Therefore, enormous efforts have been undertaken to develop tools and methods to overcome these obstacles (Figure 2). Advances in sequencing technics [75] and bioinformatics tools for genome mining [63] facilitate cost-effective sequencing and a fast identification of BGCs and other potential targets for genetic engineering. Molecular biology enzymes [76–78], fast cloning strategies and synthetic biology which involves vectors and genetic parts (e.g., attachment sites, replicons, selection markers, promoters, terminators) [79–84] were optimized for introduction and maintenance of additional or new genetic material in actinomycetes strains. Methods for transfer of the generated construct into the cell (e.g., protoplast transformation, conjugation) were established [74,85].
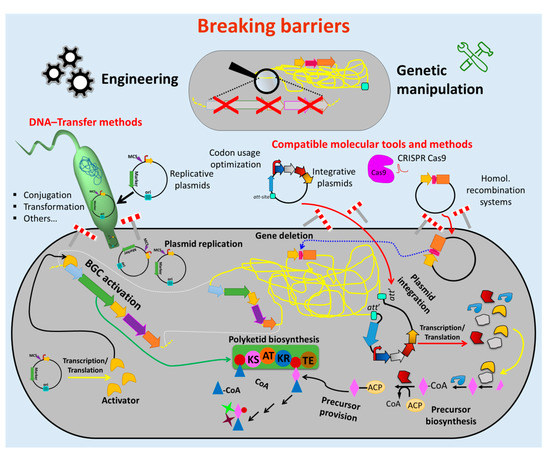
Figure 2. Breaking down the barriers to genetic manipulation and engineering of actinomycetes. Numerous methods and tools were developed to facilitate the genetic manipulation of actinomycetes. Suitable vectors and other vehicles (e.g., integrative and replicative systems) as well as fast and easy cloning strategies were developed to assemble genetic constructs, which are introduced into the host by using different transfer methods (e.g., conjugation and protoplast transformation). Protocols for avoiding DNA-degradation (e.g., treatment of the DNA with NaOH) and solutions for ensuring correct transcription/translation (e.g., codon usage optimization) were established. These innovations led to the generation of many gene deletions and (over)expression mutants (e.g., deletion of competitive biosynthetic pathways, overexpression of activators for activation of silent biosynthetic gene cluster (BGC)). This enormous progress facilitated the metabolic engineering of actinomycetes chassis for manufacturing bioactive natural products, including antibiotics. (Single elements of the figure (e.g., schematic representation of plasmid backbone) were re-used from a previous publication [74] with permission from the Royal Society of Chemistry.).
During the past five years (2015–2020) a variety of technologies and protocols for engineering of actinomycetes genomes have been established. Therefore, we believe that a summary of the recent developments (Antibiotics 2020, 9(8), 494; https://doi.org/10.3390/antibiotics9080494)[1] will be a valuable reference for the community focusing on actinomycetes and their natural products. To this end, we present an update on the molecular tools and methods for genetic engineering of actinomycetes. In the first part, details on new cloning strategies were described. The second part contains an overview on recent advances in “genetic bricks” and molecular tools for genetic manipulation of actinomycetes. As there is tremendous progress in the development of cloning strategies and CRISPR/Cas (clustered regularly interspaced short palindromic repeats/CRISPR associated)-based tools, these two sections have a “lexicon-like” structure. This should ensure an easy and time-saving search for the diverse tools and their explanation.
This entry is adapted from the peer-reviewed paper 10.3390/antibiotics9080494
References
- Mitousis, Lena; Thoma, Yvonne; and Ewa M. Musiol-Kroll, Ewa M.; An Update on Molecular Tools for Genetic Engineering of Actinomycetes—The Source of Important Antibiotics and Other Valuable Compounds . Antibiotics 2020, 9, article number: 494, https://doi.org/10.3390/antibiotics9080494.
