Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is an old version of this entry, which may differ significantly from the current revision.
Subjects:
Environmental Sciences
There is growing interest in the creation of artificial microbial consortia, especially in the field of developing and applying various bioremediation processes. Heavy metals, dyes, synthetic polymers (microplastics), pesticides, polycyclic aromatic hydrocarbons and pharmaceutical agents are among the pollutants that have been mainly targeted by bioremediation based on various consortia containing fungi and yeasts. Such consortia can be designed both for the treatment of soil and water.
- consortium composition
- main trends
- genetically modified cells
- multicomponence effect
- pollutant removal efficacy
1. Genetically Modified Microorganisms in Artificial Consortia with Fungi
Despite the high efficiency of degradation of various pollutants by natural microbial consortia [1], there has recently been a growing demand for strains that are improved by using advanced methods of synthetic biology and metabolic engineering, and numerical methods in the field of genetic engineering [2]. To date, the use of genetically modified strains in mixed microbial consortia for the decomposition of various pollutants is recognized (Figure 1). The percentage of such consortia is comparable to variants composed of yeast and bacterial cells (Figure 1). One example of the use of such an artificial consortium, as described in Table 1, Table 2, Table 3, Table 4, Table 5, Table 6 and Table 7, is the heterologous expression of genes encoding MnP and LiP enzymes in non-ligninolytic fungi, which made it possible to complement the pathway of degradation of PAHs [3], without leaving toxic intermediates in the treated medium.
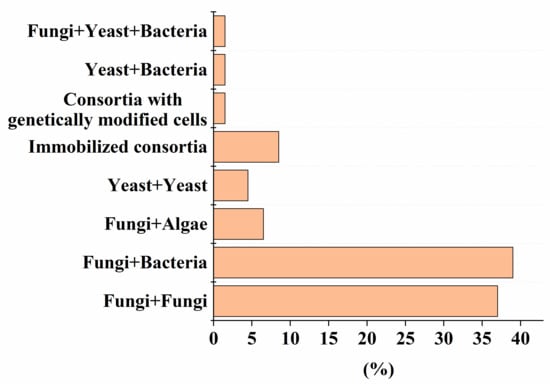
Figure 1. The percentage of artificial consortia of different composition of the total number (56) of reviewed studies, presented in Table 1, Table 2, Table 3, Table 4, Table 5, Table 6 and Table 7.
Table 1. Microbial consortia with fungal species for removal of heavy metals.
| Consortia [Reference] | Conditions | Pollutant/Process Efficiency |
|---|---|---|
| Aspergillus niveus, A. flavus, A. niger [4] |
1.4 × 106 spore/mL of each strain; pH 5.0, 110 rpm, 30 °C, 96 h |
Removal of Cr, Zn, Pb, Cd and Ni—70–90% |
| A. flavus, A. fumigatus [5] | Heavy metal concentration—100 mg/L, 1.2 × 106 spores/mL, pH 5.0, 30 °C, 144 h |
Removal of Cr(VI)—81%, Cd(II)—82%, mixture of metals—73% |
| Ascomycota and Basidiomycota fungi [6] | Initial metal concentration (23–2347 mg/kg), pH 7.9, soil moisture 60–65%, 28 °C, 100 days |
Removal of As—77%, Cr—60%, Cu—52%, Fe—52%, Mn—71% |
| Ascomycota and Basidiomycota fungi [7] | Initial metal concentration (400–800 mg/kg), pH 7.9, soil moisture 60–65%, 28 °C, 100 days | Removal efficiencies of Ni, Pb, Zn—52%, 44%, 32% respectively |
| A. fumigatus, A. terreus, Paenibacillus dendritiformis [8] |
Cd—100 mg/L, pH 5.0, 30 °C, 120 h | Removal of Cd(II)—95% |
| A. terreus, Talaromyces islandicus, Neurospora crassa, Aspergillus flavus [9] | Pb(II)—20.5–293.23 mg/L, Ni(II)—12.1–164.7 mg/L, inoculum 8%, pH 5.0, 30 °C 120 h |
Removal of Pb(II) and Ni(II)—95–97%
|
Table 2. Microbial consortia with fungal species for decolorization of dyes.
| Consortia [Reference] | Conditions | Pollutant/Process Efficiency |
|---|---|---|
| Yarrowia sp., Barnettozyma californica, Sterigmatomyces halophilus [10] | 100 mg/L of dye,30 °C, static conditions, 6–12 h |
Degradation of Scarlet GR, Red HE3B, Remazol Brilliant Blue R, Methyl Orange, Rubine GFL and Reactive Red 2—92–100% |
| Daldinia concentrica, Xylaria polymorpha [11] |
50 mg/L of dye, pH 4.5, 30 °C, 150 rpm, 48 h. |
Degradation of cibacron brilliant red 3B-A—99% |
| Rhodotorula sp., Raoultella planticola and Staphylococcus xylosus cells immobilized in Ca-alginate beads [12] | 200 mg/L of methylene blue in municipal wastewater and industrial effluent, 144 h | Degradation of methylene blue—100% and 78.5% in municipal wastewater and industrial effluent, respectively |
| A. niger, A. terrus, A. oryzae, A. fumigatus [13] |
20 mg/L of each dye, 150 rpm, 28 °C, 72 h |
Degradation of reactive blue 4, fast green, methyl red, crystal violet, alura red AC, tartrazine, naphthol blue black, janus green B, alizarin yellow R, evans blue, brilliant green, pararosaniline, ponceau S, cibacron brilliant red 3B-A, direct violet 51—57–100% |
| Aspergillus sp., Chlorella sorokiniana [14] |
Disperse Red—0.1 g/L, pH 6.0, 160 rpm, 25 °C, 4 days |
Degradation/adsorption of disperse red 3B—98.1% |
| Daedalea dickinsii, Pseudomonas aeruginosa [15] |
Methyl orange—100 mg/L, 30 °C, 7 days |
Degradation of methyl orange—98% |
| Sterigmatomyces halophilus, Meyerozyma guilliermondii [16] |
Reactive Black 5, Acid Orange 7; Reactive Green 19, Reactive Yellow, ABC, Atlantic Black C—50 mg/L, glucose as co-substrate, pH 7.0, 35 °C, 120 h |
Degradation—88–97% |
| Penicillium oxalicum, Aspergillus tubingensis [17] |
100 mg/L of congo red with dextrose (10 g/L), pH 5, 150 rpm, 28 °C, 12 h |
Congo red degradation—97.1% |
Table 3. Microbial consortia with fungal species for degradation of synthetic polymers.
| Consortia [Reference] | Conditions | Pollutant/Process Efficiency |
|---|---|---|
| Sterigmatomyces halophilus, Meyerozyma guilliermondii, M. caribbica [18] |
30 °C, 45 days | Low-density polyethylene (LDPE) mass reduction—33.2% |
| A. niger, P. aeruginosa [19] | 37 °C, 30 days | Polyurethane weight loss—20% |
| Curvularia lunata, Alternaria alternata, Penicillium simplicissimum, Fusarium sp. [20] |
90 days | Polyethylene weight loss—27% |
| A. niger, A. flavus, A. oryzae [21] | 55 days | Polyethylene weight loss—26.2% |
| Microorganisms isolated from activated sludge and river sediments (Lysinibacillus massiliensis, Bacillus licheniformis, B. indicus, B. megaterium, B. cereus, Pseudomonas alcaligenes, Aspergillus sp., Penicillium sp., Alternaria sp., Candida parapsilosis [22] |
160 rpm, 56 days at room temperature, 10 mL of bacterial and fungi suspension, and one film sample (1 cm2) of polymer materials |
Weight loss of sample (LDPE & thermoplastic starch & styrene-ethylene-styrene)—16% |
| Microorganisms isolated from compost (B. sonorensis, B. subtilis, Aspergillus sp. Trichoderma sp., Rhizopus sp.) [22] |
Weight loss—21.9% | |
| Microorganisms of enriched landfill soil (Achromobacter xylosoxidans, Trichosporon chiropterorum, Penicillium chalabudae) [23] | pH 7.2, 150 rpm, 30 °C, 90 days | LDPE weight loss—55.6% |
| Aspergillus sp., Penicillium sp. [24] | 29 °C, 85% humidity, 30 days |
Polypropylene/poly (butylene adipate-co-terephthalate)/thermoplastic starch weight loss—1.0–2.3% |
| Bacillus sp., Aspergillus sp. [25] | 30 °C, 150 rpm, 30 days | LDPE weight loss—12% |
Table 4. Microbial consortia with fungal species for degradation of pesticides.
| Consortia [Reference] | Conditions | Pollutant/Process Efficiency |
|---|---|---|
| Fomitopsis pinicola, B. subtilis [26] | 30 °C, 7 days | DDT (1,1,1-trichloro-2,2-bis(4-chlorophenyl) ethane) degradation—86% |
| Pleurotus ostreatus, P. aeruginosa [27] | 25 °C, 7 days | DDT degradation—86% |
| A. niger, Chlorella vulgaris [28] | 38 pesticides in mixture—total concentration—72.7µg/L, biomass—181.6 mg dry weight/L, pH 4.0, 100 rpm, 68 h | Degradation—23% |
| Verticilium sp., Metacordyceps sp. [29] | Concentration of each pesticide—50 mg/L, 100 rpm, pH 5.5, 27 °C, 21 days | Degradation of atrazine—81%, iprodione—96%; chlorpyrifos—99% |
| Verticilium sp., Metacordyceps sp. immobilized in Ca-alginate beads [29] | Concentration of each pesticide—50 mg/L, flow rate—90 mL/h, inoculum concentration—30 w/v, 100 rpm, 27 °C | Degradation of atrazine—64%, iprodione—96%; chlorpyrifos—85% (11–15 days) |
| Consortium of microorganisms present in coconut fiber, garden compost and agricultural soil and Trametes versicolor [30] | Mixture of pesticides—30–40 mg/kg, pH 6.4, 25 °C, 16 days | Degradation of atrazine—72.2%, carbendazim—96.7%, carbofuran—98.7%, metalaxyl—96.7% |
| Fomitopsis pinicola, Ralstonia pickettii [31] | DDT—5 mM, 30 °C, 7 days | DDT degradation—61% |
Table 5. Microbial consortia with fungal species for degradation of PAHs.
| Consortia [Reference] | Conditions | Pollutant/Process Efficiency |
|---|---|---|
| Acremonium sp, B. subtilis [32] | Concentration of each PAH in mixture—50 mg/L, 28 °C, 160 rpm, 10 days | Degradation of naphthalene—100%, fluorine—89%, phenanthrene—82%, anthracene—71%, fluoranthene—61% |
| Pleurotus ostreatus, Penicillium chrysogenum [33] | 30 °C, 30 days | Degradation of benzo[a]pyrene—86% |
| P. ostreatus, P. aeruginosa [33] | Degradation of benzo[a]pyrene—75% | |
| Consortium (Proteobacteria, Bacteroidota, Fusarium) immobilized on biochar [34] | Mixture of 50 mg/L of phenanthrene and 150 mg/L of Cd2+, 150 rpm, 30 °C, 7 days | Degradation of phenanthrene—92–98%, removing of Cd2+—94–99% |
| Consortium with two genetically modified strains of A. niger [3] | Mixture of pyrene and benzo(a)pyrene—1000 mg/kg soil, pH of 8.4, 30 °C, 14 days | Degradation efficiency of phenanthrene—92%, pyrene—64%, benzo(a)pyrene—65% |
| Kocuria rosea and A. sydowii immobilized in guargum-nanobentonite composite water dispersible granules [35] | Mixture of naphthalene, fluorene, phenanthrene, anthracene, and pyrene—100 µg of each PHA/g of soil, pH 8.3, 27 °C, 30 days | Degradation efficiency—85–100% |
| P. putida, yeast Basidioascus persicus [36] | 800 mg/L of pyrene, rhamnolipid biosurfactant 100 μL, 28 °C, 21 days | Degradation efficiency—78% |
| Ochrobactrum intermedium and white rot fungus Pleurotus ostreatus [37] | Concentrations of different PAH—138.2–268.0 mg/kg of soil, moisture—70%, 30 °C, 110 days | Degradation of fluoranthene, indene[1,2,3-cd]pyrene and benzo[g,h,i]perylene—100%; Anthracene, pyrene, chrysene and benzo[a]anthracene—96%, 86%, 98% and 98%, respectively |
| Pleurotus ostreatus, Azospirillum brasilense [38] | A mixture of anthracene, phenanthrene, fluorene, pyrene, and fluoranthene—50 mg/L, 130 rpm, 24 °C, 14 days | Degradation efficiency > 70% |
Table 6. Microbial consortia with fungal species for degradation of pharmaceutical pollutants.
| Consortia [Reference] | Conditions | Pollutant/Process Efficiency |
|---|---|---|
| Pycnoporus sanguineus, Phanerochaete chrysosporium [39] | Each antibiotic concentration—10 mg/L, biomass of each strain—0.15 g dry weight/L), pH 4.5, 30 °C, 4 days | Removal efficiency of ciprofloxacin, norfloxacin and sulfamethoxazole in their mixture—100% |
| Pycnoporus sanguineus, Alcaligenes faecalis [40] | Sulfamethoxazole (50 mg/L) and vitamins mixture (VB2, VB6, VB12 and VC), 28 °C, 120 rpm, 24 h | Sulfamethoxazole degradation—93% |
| Ganoderma applanatum, Laetiporus sulphureus [41] | Concentration of each of three pollutants—10 mg/L, pH 6.4, ambient temperature, 150 rpm, 72 h | Degradation (mixture of celecoxib, diclofenac and ibuprofen)—99.5% |
| A. niger, Mucor circinelloides, Trichoderma longibrachiatum, Trametes polyzona and Rhizopus microsporus [42] | Pollutants concentration—1 mg/L, pH 4.6, 30 °C, 7 days, consortium concentration—30% (v/v) | Degradation of carbamazepine—90%, diclofenac sodium—96% and ibuprofen—91% |
| A. niger, C. vulgaris [43] | Pharmaceutical substances—8–11 μg/L, microalgae-fungus biomass—75 mg dry weight/L, 72 h | Relative removal of initial ranitidine concentrations—50% |
| Penicillium rastrickii, P. oxalicum, Cladosporium cladosporoides, Micrococcus yunnanensis, Oligella ureolytica, Sphingobacterium jejuense [44] | Mixture of diclofenac, carbamazepine and ketoprofen with 100 μM of each compound, 28 °C, 10 days | Degradation of diclofenac—99%, ketoprofen—80% |
| Chlorella vulgaris, Aspergillus oryzae [45] | Simulated swine wastewater with addition of 0.1–0.5 mg/L Cu (II), 0.4 mg/L of mixture of antimicrobial agents, pH 7.2, 28 °C, 14 days | Removal efficiency of sulfamonomethoxine, sulfamethoxazole and sulfamethazine—58.8%, 63.5%, and 63.9%, respectively |
Table 7. Microbial consortia with fungal species for degradation of various pollutants not mentioned in Table 1, Table 2, Table 3, Table 4, Table 5 and Table 6.
| Consortia [Reference] | * Conditions | Pollutant/Process Efficiency |
|---|---|---|
| Acinetobacter baumannii, Talaromyces sp. [46] | The initial concentration of petroleum in soil—1220 mg/kg, pH 8.3, 30 °C, 28 days | Degradation of petroleum—65.6% |
| Paraburkholderia sp., Paraburkholderia tropica, Scedosporium boydii [47] | 1% v/v crude oil, 120 rpm, 30 °C, 7 days | Degradation of crude oil—81.5% |
| Scedosporium sp., Acinetobacter sp. [48] | Crude oil—200 mg/L, pH 7.0, 150 rpm, 30 °C, 7 days | Crude oil degradation—58.6% |
| Micrococcus luteus, Rhodococcus equi, A. niger [49] |
Greywater—COD—1165.6 mg/L, oil and grease—58 mg/L, sulphate—95.6 mg/L, pH 7, 35 °C, 96 h | Degradation of COD, oil and grease and sulphate were 78.7, 82.6 and 89.7%, respectively |
| Aspergillus versicolor and bacterial species (Pseudomonas, Klebsiella species, B. subtilis) [50] | Greywater with 100 μg/L of carbendazim and thiamethoxam, 80 rpm, 30 °C, 240 h |
Degradation of carbendazim and thiamethoxam 94.4 and 93.6%, respectively |
| A. flavus, Fusarium oxysporium [51] | Real textile effluent pH 8.7, COD—611 mg/L, pH 6.0–8.0, 28 °C, 7 days | Degradation—78.1%, COD removal—77.6% |
| Consortium of Brevibacillus laterosporus and Galactomyces geotrichum immobilized into Ca-alginate or polyvinyl alcohol-alginate beads [52] | Textile industry effluent pH 8.8, COD—2400 mg/L, 48–60 h | Degradation during 5 repeated cycles—76–95% |
| Ralstonia pickettii, Trichoderma viride [53] | Chlorobenzene—220 mg/L, 160 rpm, 28 °C, 60 h | Chlorobenzene degradation—100% |
| Chaetomium globosum, A. niger, Rhizopus oryzae [54] | Poly(vinyl acetate) processing wastewater pH 7.1, COD—23.48 g/L; pH 5.5, 150 rpm, 28 °C, 10 days | COD, poly(vinyl acetate) and color removal yields—97.8%, 98.5% and 99.8%, respectively. |
| Phanerochaete chrysosporium, Delftia lacustris [55] | Phenol (1000 mg/L) and selenite concentration—10 mg/L, 180 rpm, pH 6.5, 30 °C, 120 h | Phenol degradation—97.8% with the simultaneous reduction of selenite to Se(0) |
* COD—chemical oxygen demand.
However, the traditional method of random mutagenesis is a time-consuming approach, and searching for key amino acid mutations is extremely complex and requires operating across a large space. With the help of computer modeling, it is possible to “calculate” enzymes with improved binding affinity and greater specificity of action in relation to the substrates destruction, which are both necessary [56]; it is also necessary to successfully introduce producers of “improved” enzymes into consortia, which is another serious task that confronts researchers.
Another difficulty is that genetically modified strains often have to survive high concentrations of pollutants [2][57]. Several strategies have been used to improve cellular tolerance, including changing the composition of membrane lipids, phenotypic screening through adaptive laboratory evolution, modification of global gene expression, genome shuffling, directional evolution, and others.
Although genetically modified strains are effective in bioremediation processes [58], their use is limited to laboratory studies due to minimizing the risks of environmental impact. Since there are no globally accepted regulatory documents that concern the spread of genetically modified organisms, the regulation of the development and release of genetically modified organisms varies in different countries, depending on the purposes of their use, extending from a complete ban on their import, release or use to allowing their use, subject to varying degrees of regulation. However, despite this, methods of genetically engineering filamentous fungi continue to be actively developed and used in research around the world, making it possible to overcome many of the shortcomings of classical methods for improving strains [57][59][60]. To eliminate the negative effect of such cells, it is possible to use several genetic tools, including cell self-destruction systems [61]. Such approaches to realization of programmed cell death in a certain period of their functioning can be used and activated after the completion of bioremediation or after accumulation of certain concentrations of the cells.
3.2. Role of Composition in Artificial Consortia with Fungal Cells
Microbial consortia have a high self-organization, which allows them to carry out the catalytic conversion of substrates with high efficiency, providing an intensive exchange of metabolites. Moreover, in consortia, cells have higher adaptability and viability in relation to environmental factors (pH, temperature, concentration of pollutants, etc.) [62]. Despite this, most artificial microbial consortia face problems of non-sustainable functioning [63]. Even minor fluctuations in the composition and activities of consortia can have a negative impact on the effectiveness of the processes taking place with their participation. The difficulty of managing the bioremediation characteristics of artificial consortia is actually determined by the qualitative (Figure 1) and quantitative composition (Figure 2) of the participants in the artificial biosystems being formed.
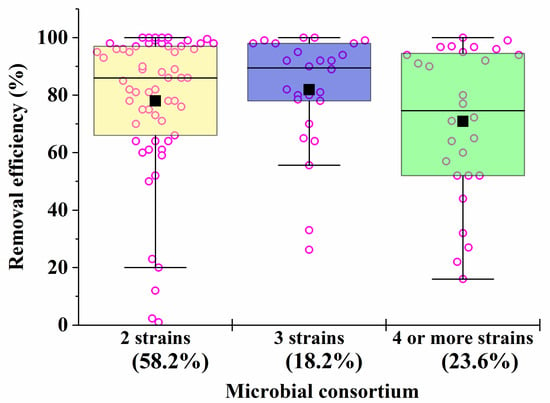
Figure 2. The average efficiency of degradation of pollutants by consortia with two, three, four and more strains in the composition. The percentage of the total number of all options presented in Table 1, Table 2, Table 3, Table 4, Table 5, Table 6 and Table 7 (56 studies reviewed) was calculated. Each point corresponds to the research results listed in Table 1, Table 2, Table 3, Table 4, Table 5, Table 6 and Table 7.
As the analysis of the data in Table 1, Table 2, Table 3, Table 4, Table 5, Table 6 and Table 7 shows that, despite the diversity of the consortia being developed (Figure 1), the ones most widely used to create effective bioremediation microbial systems, that have demonstrated their advantages over natural systems, consist of two or three microorganisms, of which one is a fungal culture (Figure 2). The maximum part of all artificial consortia currently being developed combines mycelial fungi with each other and with cells of different bacteria (Figure 1).
It is known that fungi and bacteria are characterized by different rates of synthesis of enzymes that are necessary for the catalysis of different processes in the bioremediation of pollutants. Consequently, the rates of different processes may be compared by varying the biomass of certain cells introduced into heterogeneous consortia in order to ensure their maximum effectiveness of action. This is the basis for the unification of certain microorganisms into artificial consortia (Figure 3).
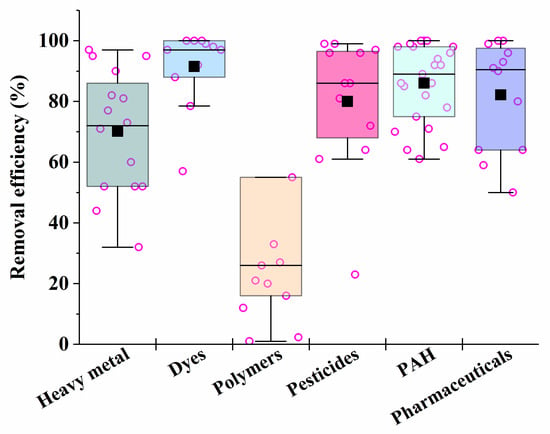
Figure 3. The average efficiency of the consortia for elimination of different types of pollutants, calculated on the basis of sources performed in Table 1, Table 2, Table 3, Table 4, Table 5, Table 6 and Table 7. Each point corresponds to the research results listed in Table 1, Table 2, Table 3, Table 4, Table 5, Table 6 and Table 7. The six differently colored squares correspond to the six pollutants shown on the abscissa axis.
After analyzing the data in Table 1, Table 2, Table 3, Table 4, Table 5, Table 6 and Table 7, it is possible to conclude that there are consortia consisting of two–three strains of various fungi, mainly mycelial types, that are most effective in removing 70–90% of heavy metals in mixtures from various media.
For the removal of dyes, the most effective are the consortia consisting of yeast cells, which provide bioremediation efficiency of up to 100%; the destruction of pesticides is the most successful under the action of consortia consisting of fungal cultures, such as Verticilium sp. and Metacordyceps sp.
Synthetic polymers undergo the most difficult microbial degradation, but the maximum weight loss of synthetic polymers (microplastics) is achieved under the action of consortia that combine bacteria, mycelial fungi or yeast.
In the case of the presence of mixtures of several PAHs in media subjected to bioremediation, the most effective (in terms of the degree of degradation of the pollutant (86–100%)) were consortia that included bacteria and white rot fungus Pleurotus ostreatus, as well as an immobilized consortium consisting of the bacteria Kocuria rosea and the fungi Aspergillus sydowii.
Among the most effective in the removal of pharmaceutical pollutants were selected consortia from cells of white rot saprobic fungus Pycnoporus sanguineus and Phanerochaete chrysosporium (the removal efficiency of ciprofloxacin-, norfloxacin and sulfamethoxazole in their mixture was 100%), as well as tinder fungi Ganoderma applanatum and Laetiporus sulphureus (efficiency of degradation of a mixture containing celecoxib, diclofenac and ibuprofen was 99.5%).
Bacterial and fungal consortia proved to be the most effective for the treatment of real wastewater from industrial enterprises and oil pollution. The fungal consortium consisting of Phanerochaete chrysosporium and Delftia lacustris can be used to remove phenol, with an efficiency of over 90%.
Of course, in order to achieve the effective functioning of synthetic microbial consortia, it is important to predict the possible types of interactions of all microorganisms involved in the functioning of artificially created biosystems [64][65].
Most often, these interactions are based on mutualism or competition [66][67]. Mutualism suggests that jointly cultivated microorganisms have a beneficial effect on each other, while the composition of artificial consortia is stabilized, but only at a certain cell density [55][68].
In case of competition for a substrate, consortium members can release metabolites into the environment that negatively affect other consortium members (organic acids, mycotoxins, antibiotics, antimicrobial peptides, enzymes) [69][70][71][72][73]. As a rule, these processes are regulated by cell Quorum Sensing (QS) [74]. QS molecules and the conditions of their formation can be used to control or regulate the expression of certain genes, control the composition of consortia, and ensure intercellular connections between certain consortium members [63][68][75]. In considering this, modern studies of artificial consortia should enable the creation of model systems for the accumulation of toxic metabolites of a number of microorganisms (fungi, microalgae, bacteria) and develop effective ways to detoxify them.
Cell immobilization can also solve the instability problems of microbial consortia, since cellular communication can be more active in a confined space and positively affects the speed of bioremediation processes. At the same time, the inclusion of cells in gel matrices [13][29][52] simulates the development of a stabilized state of cells and the additional formation of biofilms [37][47][55][58][68].
Immobilized fungal consortia can be used in non-sterile conditions at a separate stage, and then be integrated into conventional waste treatment systems before the action of aerobic sludge at water processing stations [76][77][78]. In some cases, the self-stabilization of the artificial fungi-containing consortia is sufficient for their use in non-immobilized form in laboratory conditions to treat various real contaminated industrial and environmental water and soil samples [4][9][10][12][51][52][54][79]. While such studies bring the introduction of fungal consortia closer to practice, there are not many known examples of pilot or industrial tests of artificial consortia [80][81][82]. Some of them follow:
- -
-
microbial consortium containing T. versicolor, P. ostreatus, Phanerochaete sp., Pseudomonas fluorescens and B. subtilis cells was applied for the treatment of non-domestic wastewater. This fungal/bacterial consortium was prepared by mixing fungal biomass pellets with suspensions of bacterial cells. The removal of colored substances (2700 Color Units550nm), COD (1.75 g/L) and nitrate (3 mg/L) was 91 ± 2%, 90 ± 4% and 17 ± 2%, respectively, after 15 days of water treatment at a pilot plant [80];
- -
-
consortium of A. niger, Mucor hiemalis and Galactomyces geotrichum, has been tested for the treatment of real wastewater from industry at a pilot scale station (110 L) and industrial wastewater treatment plant (1000 L). The efficiency of COD removal in the industrial reactor was 50% under the influence of this consortium [81];
- -
-
consortium containing Acinetobacter oleivorans, Corynebacterium sp., Pseudomonas sp, Rhodococcus sp., Micrococcus sp. and yeast Yarrowia sp. was tested by Ecophile Co., Ltd. (Korea) in the biodegradation of hydrocarbons in soil (2300 mg/kg) contaminated with diesel fuel. This large-scale experiment involved two samples of 100 metric tons of contaminated soil, both without (control) and with consortium treatment (109 cells/kg of soil). The introduction of consortium reduced pollution by 57.7% within 2 weeks, whereas in the control (without the consortium), degradation was only 10.1% [82].
Thus, such positive samples of scaling use of artificial fungal consortia not only demonstrate their real-world efficacy, but also addresses potential solutions encountered during practical applications.
This entry is adapted from the peer-reviewed paper 10.3390/microorganisms12030470
References
- Li, H.; Qiu, Y.; Yao, T.; Ma, Y.; Zhang, H.; Yang, X.; Li, C. Evaluation of seven chemical pesticides by mixed microbial culture (PCS-1): Degradation ability, microbial community, and Medicago sativa phytotoxicity. J. Hazard. Mater. 2020, 389, 121834.
- Tran, K.M.; Lee, H.M.; Thai, T.D.; Shen, J.; Eyun, S.I.; Na, D. Synthetically engineered microbial scavengers for enhanced bioremediation. J. Hazard. Mater. 2021, 419, 126516.
- Zafra, G.; Absalón, Á.E.; Anducho-Reyes, M.Á.; Fernandez, F.J.; Cortés-Espinosa, D.V. Construction of PAH-degrading mixed microbial consortia by induced selection in soil. Chemosphere 2017, 172, 120–126.
- Chaudhary, P.; Beniwal, V.; Sharma, P.; Goyal, S.; Kumar, R.; Alkhanjaf, A.A.M.; Umar, A. Unloading of hazardous Cr and tannic acid from real and synthetic waste water by novel fungal consortia. Environ. Technol. Innov. 2022, 26, 102230.
- Talukdar, D.; Jasrotia, T.; Sharma, R.; Jaglan, S.; Kumar, R.; Vats, R.; Umar, A. Evaluation of novel indigenous fungal consortium for enhanced bioremediation of heavy metals from contaminated sites. Environ. Technol. Innov. 2020, 20, 101050.
- Hassan, A.; Pariatamby, A.; Ossai, I.C.; Hamid, F.S. Bioaugmentation assisted mycoremediation of heavy metal and/metalloid landfill contaminated soil using consortia of filamentous fungi. Biochem. Eng. J. 2020, 157, 107550.
- Hassan, A.; Periathamby, A.; Ahmed, A.; Innocent, O.; Hamid, F.S. Effective bioremediation of heavy metal–contaminated landfill soil through bioaugmentation using consortia of fungi. J. Soils Sediments 2020, 20, 66–80.
- Zango, U.U.; Ahluwalia, S.S.; Sharma, A.K. Microbial consortium of Aspergillus fumigatus, Aspergillus terreus and Paenibacillus dendritiformis in the bioremoval of cadmium. Int. J. Pharm. Res. 2018, 10, 230–238.
- Sharma, R.; Jasrotia, T.; Kumar, R.; Kumar, R.; Alothman, A.A.; mana AL-Anazy, M.; Algahtani, K.N.; Umar, A. Multi-biological combined system: A mechanistic approach for removal of multiple heavy metals. Chemosphere 2021, 276, 130018.
- Ali, S.S.; Al-Tohamy, R.; Koutra, E.; El-Naggar, A.H.; Kornaros, M.; Sun, J. Valorizing lignin-like dyes and textile dyeing wastewater by a newly constructed lipid-producing and lignin modifying oleaginous yeast consortium valued for biodiesel and bioremediation. J. Hazard. Mater. 2021, 403, 123575.
- Bankole, P.O.; Adekunle, A.A.; Govindwar, S.P. Biodegradation of a monochlorotriazine dye, cibacron brilliant red 3B-A in solid state fermentation by wood-rot fungal consortium, Daldinia concentrica and Xylaria polymorpha: Co-biomass decolorization of cibacron brilliant red 3B-A dye. Int. J. Biol. Macromol. 2018, 120, 19–27.
- Eltarahony, M.; El-Fakharany, E.; Abu-Serie, M.; ElKady, M.; Ibrahim, A. Statistical modeling of methylene blue degradation by yeast-bacteria consortium; optimization via agro-industrial waste, immobilization and application in real effluents. Microb. Cell Fact. 2021, 20, 234.
- Abd El-Rahim, W.M.; Moawad, H.; Azeiz, A.Z.A.; Sadowsky, M.J. Biodegradation of azo dyes by bacterial or fungal consortium and identification of the biodegradation products. Egypt. J. Aquat. Res. 2021, 47, 269–276.
- Tang, W.; Xu, X.; Ye, B.C.; Cao, P.; Ali, A. Decolorization and degradation analysis of Disperse Red 3B by a consortium of the fungus Aspergillus sp. XJ-2 and the microalgae Chlorella sorokiniana XJK. RSC Adv. 2019, 9, 14558–14566.
- Purnomo, A.S.; Mawaddah, M.O. Biodecolorization of methyl orange by mixed cultures of brown-rot fungus Daedalea dickinsii and bacterium Pseudomonas aeruginosa. Biodiversitas 2020, 21, 2297–2302.
- Al-Tohamy, R.; Ali, S.S.; Xie, R.; Schagerl, M.; Khalil, M.A.; Sun, J. Decolorization of reactive azo dye using novel halotolerant yeast consortium HYC and proposed degradation pathway. Ecotoxicol. Environ. Saf. 2023, 263, 115258.
- Thakor, R.; Mistry, H.; Tapodhan, K.; Bariya, H. Efficient biodegradation of Congo red dye using fungal consortium incorporated with Penicillium oxalicum and Aspergillus tubingensis. Folia Microbiol. 2022, 67, 33–43.
- Elsamahy, T.; Sun, J.; Elsilk, S.E.; Ali, S.S. Biodegradation of low-density polyethylene plastic waste by a constructed tri-culture yeast consortium from wood-feeding termite: Degradation mechanism and pathway. J. Hazard. Mater. 2023, 448, 130944.
- Fernandes, I.P.; Barbosa, M.; Amaral, J.S.; Pinto, V.; Rodrigues, J.L.; Ferreira, M.J.; Barreiro, M.F. Biobased additives as biodegradability enhancers with application in TPU-based footwear components. J. Renew. Mater. 2016, 4, 47.
- Sowmya, H.V.; Ramalingappa, B.; Nayanashree, G.; Thippeswamy, B.; Krishnappa, M. Polyethylene degradation by fungal consortium. Int. J. Environ. Res. 2015, 9, 823–830.
- DSouza, G.C.; Sheriff, R.S.; Ullanat, V.; Shrikrishna, A.; Joshi, A.V.; Hiremath, L.; Entoori, K. Fungal biodegradation of low-density polyethylene using consortium of Aspergillus species under controlled conditions. Heliyon 2021, 7, 51–66.
- Kučić Grgić, D.; Miloloža, M.; Ocelić Bulatović, V.; Ukić, Š.; Slouf, M.; Gajdosova, V. Screening the efficacy of a microbial consortium of bacteria and fungi isolated from different environmental samples for the degradation of LDPE/TPS films. Separations 2023, 10, 79.
- Chigwada, A.D.; Ogola, H.J.O.; Tekere, M. Multivariate analysis of enriched landfill soil consortia provide insight on the community structural perturbation and functioning during low-density polyethylene degradation. Microbiol. Res. 2023, 274, 127425.
- de Oliveira, T.A.; Barbosa, R.; Mesquita, A.B.; Ferreira, J.H.; de Carvalho, L.H.; Alves, T.S. Fungal degradation of reprocessed PP/PBAT/thermoplastic starch blends. J. Mater. Res. Technol. 2020, 9, 2338–2349.
- Rajan, P.; Rangasamy, M.; Sundaram, S.K. Bioremediation of preprocessed plastic wastes through microbial consortium. J. Adv. Sci. Res. 2020, 11, 106–113.
- Sariwati, A.; Purnomo, A.S.; Kamei, I. Abilities of co-cultures of brown-rot fungus Fomitopsis pinicola and Bacillus subtilis on biodegradation of DDT. Curr. Microbiol. 2017, 74, 1068–1075.
- Purnomo, A.S.; Ashari, K.; Hermansyah, F.T. Evaluation of the synergistic effect of mixed cultures of white-rot fungus Pleurotus ostreatus and biosurfactant-producing bacteria on DDT biodegradation. J. Microbiol. Biotechnol. 2017, 27, 1306–1315.
- Hultberg, M.; Bodin, H. Effects of fungal-assisted algal harvesting through biopellet formation on pesticides in water. Biodegradation 2018, 29, 557–565.
- Levio-Raiman, M.; Briceño, G.; Leiva, B.; López, S.; Schalchli, H.; Lamilla, C.; Bornhardt, C.; Diez, M.C. Treatment of pesticide-contaminated water using a selected fungal consortium: Study in a batch and packed-bed bioreactor. Agronomy 2021, 11, 743.
- Castro-Gutiérrez, V.; Masís-Mora, M.; Carazo-Rojas, E.; Mora-López, M.; Rodríguez-Rodríguez, C.E. Fungal and bacterial co-bioaugmentation of a pesticide-degrading biomixture: Pesticide removal and community structure variations during different treatments. Water Air Soil Pollut. 2019, 230, 247.
- Purnomo, A.S.; Sariwati, A.; Kamei, I. Synergistic interaction of a consortium of the brown-rot fungus Fomitopsis pinicola and the bacterium Ralstonia pickettii for DDT biodegradation. Heliyon 2020, 6, e04027.
- Ma, X.K.; Li, T.T.; Fam, H.; Charles Peterson, E.; Zhao, W.W.; Guo, W.; Zhou, B. The influence of heavy metals on the bioremediation of polycyclic aromatic hydrocarbons in aquatic system by a bacterial-fungal consortium. Environ. Technol. 2018, 39, 2128–2137.
- Bhattacharya, S.; Das, A.; Palaniswamy, M.; Angayarkanni, J. Degradation of benzopyrene by Pleurotus ostreatus PO-3 in the presence of defined fungal and bacterial co-cultures. J. Basic Microbiol. 2017, 57, 95–103.
- Li, W.; Zhu, Y.; Li, K.; Wang, L.; Li, D.; Liu, N.; Huang, S. Synergistic remediation of phenanthrene–cadmium co-contaminants by an immobilized acclimated bacterial–fungal consortium and its community response. Chemosphere 2023, 336, 139234.
- Khandelwal, A.; Sugavanam, R.; Ramakrishnan, B.; Dutta, A.; Varghese, E.; Banerjee, T.; Nain, L.; Singh, S.B.; Singh, N. Bio-polysaccharide composites mediated degradation of polyaromatic hydrocarbons in a sandy soil using free and immobilized consortium of Kocuria rosea and Aspergillus sydowii. Environ. Sci. Pollut. Res. 2022, 29, 80005–80020.
- Kamyabi, A.; Nouri, H.; Moghimi, H. Characterization of pyrene degradation and metabolite identification by Basidioascus persicus and mineralization enhancement with bacterial-yeast co-culture. Ecotoxicol. Environ. Saf. 2018, 163, 471–477.
- Acevedo-Sandoval, O.; Gutierrez-Alcantara, E.J.; Perez-Balan, R.; Rodriguez-Vazquez, G.; Zamorategui-Molina, A.; Tirado-Torres, D. Degradation of polycyclic aromatic hydrocarbons using bacterial isolate from the contaminated soil and white rot fungus Pleurotus ostreatus. Appl. Ecol. Environ. Res. 2018, 16, 3815–3829.
- Pozdnyakova, N.; Muratova, A.; Turkovskaya, O. Degradation of polycyclic aromatic hydrocarbons by co-culture of Pleurotus ostreatus Florida and Azospirillum brasilense. Appl. Microbiol. 2022, 2, 735–748.
- Gao, N.; Liu, C.X.; Xu, Q.M.; Cheng, J.S.; Yuan, Y.J. Simultaneous removal of ciprofloxacin, norfloxacin, sulfamethoxazole by co-producing oxidative enzymes system of Phanerochaete chrysosporium and Pycnoporus sanguineus. Chemosphere 2018, 195, 146–155.
- Li, X.; Xu, Q.M.; Cheng, J.S.; Yuan, Y.J. Improving the bioremoval of sulfamethoxazole and alleviating cytotoxicity of its biotransformation by laccase producing system under coculture of Pycnoporus sanguineus and Alcaligenes faecalis. Bioresour. Technol. 2016, 220, 333–340.
- Bankole, P.O.; Adekunle, A.A.; Jeon, B.H.; Govindwar, S.P. Novel cobiomass degradation of NSAIDs by two wood rot fungi, Ganoderma applanatum and Laetiporus sulphureus: Ligninolytic enzymes induction, isotherm and kinetic studies. Ecotoxicol. Environ. Saf. 2020, 203, 110997.
- Kasonga, T.K.; Coetzee, M.A.; Van Zijl, C.; Momba, M.N.B. Removal of pharmaceutical’estrogenic activity of sequencing batch reactor effluents assessed in the T47D-KBluc reporter gene assay. J. Environ. Manag. 2019, 240, 209–218.
- Bodin, H.; Daneshvar, A.; Gros, M.; Hultberg, M. Effects of biopellets composed of microalgae and fungi on pharmaceuticals present at environmentally relevant levels in water. Ecol. Eng. 2016, 91, 169–172.
- Angeles-de Paz, G.; Ledezma-Villanueva, A.; Robledo-Mahón, T.; Pozo, C.; Calvo, C.; Aranda, E.; Purswani, J. Assembled mixed co-cultures for emerging pollutant removal using native microorganisms from sewage sludge. Chemosphere 2023, 313, 137472.
- Li, S.; Zhu, L. Copper regulates degradation of typical antibiotics by microalgal-fungal consortium in simulated swine wastewater: Insights into metabolic routes and dissolved organic matters. Water Res. 2023, 245, 120654.
- Liu, X.; He, L.; Zhang, X.; Kong, D.; Chen, Z.; Lin, J.; Wang, C. Bioremediation of petroleum-contaminated saline soil by Acinetobacter baumannii and Talaromyces sp. and functional potential analysis using metagenomic sequencing. Environ. Pollut. 2022, 311, 119970.
- Yuan, X.; Zhang, X.; Chen, X.; Kong, D.; Liu, X.; Shen, S. Synergistic degradation of crude oil by indigenous bacterial consortium and exogenous fungus Scedosporium boydii. Bioresour. Technol. 2018, 264, 190–197.
- Atakpa, E.O.; Zhou, H.; Jiang, L.; Ma, Y.; Liang, Y.; Li, Y.; Zhang, D.; Zhang, C. Improved degradation of petroleum hydrocarbons by co-culture of fungi and biosurfactant-producing bacteria. Chemosphere 2022, 290, 133337.
- Rajpal, N.; Ratan, J.K.; Divya, N.; Hebbani, A.V. Bioremediation of greywater using a novel bacterial–fungal consortium: Optimization and validation of the operating parameters in vitro. Environ. Technol. 2021, 43, 2430–2442.
- Rajpal, N.; Verma, S.; Kumar, N.; Lee, J.; Kim, K.H.; Ratan, J.K.; Divya, N. Bioremediation of carbendazim and thiamethoxam in domestic greywater using a bioaugmented microbial consortium. Environ. Technol. Innov. 2023, 30, 103087.
- Selim, M.T.; Salem, S.S.; Mohamed, A.A.; El-Gamal, M.S.; Awad, M.F.; Fouda, A. Biological treatment of real textile effluent using Aspergillus flavus and Fusarium oxysporium and their consortium along with the evaluation of their phytotoxicity. J. Fungi 2021, 7, 193.
- Kurade, M.B.; Waghmode, T.R.; Xiong, J.Q.; Govindwar, S.P.; Jeon, B.H. Decolorization of textile industry effluent using immobilized consortium cells in upflow fixed bed reactor. J. Clean. Prod. 2019, 213, 884–891.
- Cheng, Z.; Li, C.; Kennes, C.; Ye, J.; Chen, D.; Zhang, S.; Chen, S.; Yu, J. Improved biodegradation potential of chlorobenzene by a mixed fungal-bacterial consortium. Int. Biodeterior. Biodegrad. 2017, 123, 276–285.
- Ajmi, K.; Vismara, E.; Manai, I.; Haddad, M.; Hamdi, M.; Bouallagui, H. Polyvinyl acetate processing wastewater treatment using combined Fenton’s reagent and fungal consortium: Application of central composite design for conditions optimization. J. Hazard. Mater. 2018, 358, 243–255.
- Chakraborty, S.; Rene, E.R.; Lens, P.N.L. Reduction of selenite to elemental Se(0) with simultaneous degradation of phenol by co-cultures of Phanerochaete chrysosporium and Delftia lacustris. J. Microbiol. 2019, 57, 738–747.
- Lyagin, I.; Efremenko, E. Theoretical evaluation of suspected enzymatic hydrolysis of novichok agents. Catal. Commun. 2019, 120, 91–94.
- Liu, L.; Bilal, M.; Duan, X.; Iqbal, H.M. Mitigation of environmental pollution by genetically engineered bacteria—Current challenges and future perspectives. Sci. Total Environ. 2019, 667, 444–454.
- Lin, Y.; Zhang, H.; Li, P.; Jin, J.; Li, Z. The bacterial consortia promote plant growth and secondary metabolite accumulation in Astragalus mongholicus under drought stress. BMC Plant Biol. 2022, 22, 475.
- Meyer, V.; Basenko, E.Y.; Benz, J.P.; Braus, G.H.; Caddick, M.X.; Csukai, M.; de Vries, R.P.; Endy, D.; Frisvad, J.C.; Gunde-Cimerman, N.; et al. Growing a circular economy with fungal biotechnology: A white paper. Fungal Biol. Biotechnol. 2020, 7, 1–23.
- Salazar-Cerezo, S.; de Vries, R.P.; Garrigues, S. Strategies for the development of industrial fungal producing strains. J. Fungi 2023, 9, 834.
- Xue, Y.; Qiu, T.; Sun, Z.; Liu, F.; Yu, B. Mercury bioremediation by engineered Pseudomonas putida KT2440 with adaptationally optimized biosecurity circuit. Environ. Microbiol. 2022, 24, 3022–3036.
- Chaudhary, P.; Xu, M.; Ahamad, L.; Chaudhary, A.; Kumar, G.; Adeleke, B.S.; Verma, K.K.; Hu, D.-M.; Širić, I.; Kumar, P.; et al. Application of synthetic consortia for improvement of soil fertility, pollution remediation, and agricultural productivity: A Review. Agronomy 2023, 13, 643.
- Xu, C.; Yu, H. Insights into constructing a stable and efficient microbial consortium. Chin. J. Chem. Eng. 2021, 30, 112–120.
- Kong, W.; Meldgin, D.R.; Collins, J.J.; Lu, T. Designing microbial consortia with defined social interactions. Nat. Chem. Biol. 2018, 14, 821–829.
- Cao, Z.; Yan, W.; Ding, M.; Yuan, Y. Construction of microbial consortia for microbial degradation of complex compounds. Front. Bioeng. Biotechnol. 2022, 10, 1051233.
- Mittermeier, F.; Bäumler, M.; Arulrajah, P.; García Lima, J.J.; Hauke, S.; Stock, A.; Weuster-Botz, D. Artificial microbial consortia for bioproduction processes. Eng. Life Sci. 2022, 23, e2100152.
- Adamu, K.S.; Bichi, Y.H.; Nasiru, A.Y.; Babangida, A.M.; Umar, M.M.; Usman, G.; Muhammad, R. Synthetic microbial consortia in bioremediation and biodegradation. Int. J. Res. Sci. Innov. Appl. Sci. 2023, 8, 232–241.
- Hennig, S.; Wenzel, M.; Haas, C.; Hoffmann, A.; Weber, J.; Rödel, G.; Ostermann, K. New approaches in bioprocess-control: Consortium guidance by synthetic cell-cell communication based on fungal pheromones. Eng. Life Sci. 2018, 18, 387–400.
- Liu, G.L.; Chi, Z.; Wang, G.Y.; Wang, Z.P.; Li, Y.; Chi, Z.M. Yeast killer toxins, molecular mechanisms of their action and their applications. Crit. Rev. Biotechnol. 2015, 35, 222–234.
- Efremenko, E.; Aslanli, A.; Stepanov, N.; Senko, O.; Maslova, O. Various biomimetics, including peptides as antifungals. Biomimetics 2023, 8, 513.
- Costa, A.F.; Silva, L.D.C.; Amaral, A.C. Farnesol: An approach on biofilms and nanotechnology. Med. Mycol. 2021, 59, 958–969.
- Kischkel, B.; Souza, G.K.; Chiavelli, L.U.R.; Pomini, A.M.; Svidzinski, T.I.E.; Negri, M. The ability of farnesol to prevent adhesion and disrupt Fusarium keratoplasticum biofilm. Appl. Microbiol. Biotechnol. 2020, 104, 377–389.
- Greeff-Laubscher, M.R.; Beukes, I.; Marais, G.J.; Jacobs, K. Mycotoxin production by three different toxigenic fungi genera on formulated abalone feed and the effect of an aquatic environment on fumonisins. Mycology 2019, 11, 105–117.
- Efremenko, E.; Senko, O.; Stepanov, N.; Aslanli, A.; Maslova, O.; Lyagin, I. Quorum sensing as a trigger that improves characteristics of microbial biocatalysts. Microorganisms 2023, 11, 1395.
- Loncar, J.; Bellich, B.; Cescutti, P.; Motola, A.; Beccaccioli, M.; Zjalic, S.; Reverberi, M. The effect of mushroom culture filtrates on the inhibition of mycotoxins produced by Aspergillus flavus and Aspergillus carbonarius. Toxins 2023, 15, 177.
- Srinuanpan, S.; Chawpraknoi, A.; Chantarit, S.; Cheirsilp, B.; Prasertsan, P. A rapid method for harvesting and immobilization of oleaginous microalgae using pellet-forming filamentous fungi and the application in phytoremediation of secondary effluent. Int. J. Phytoremed. 2018, 20, 1017–1024.
- Gururani, P.; Bhatnagar, P.; Kumar, V.; Vlaskin, M.S.; Grigorenko, A.V. Algal consortiums: A novel and integrated approach for wastewater treatment. Water 2022, 14, 3784.
- Walls, L.E.; Velasquez-Orta, S.B.; Romero-Frasca, E.; Leary, P.; Noguez, I.Y.; Ledesma, M.T.O. Non-sterile heterotrophic cultivation of native wastewater yeast and microalgae for integrated municipal wastewater treatment and bioethanol production. Biochem. Eng. J. 2019, 151, 107319.
- Rios-Miguel, A.B.; van Bergen, T.J.; Zillien, C.; Ragas, A.M.; van Zelm, R.; Jetten, M.S.; Hedriks, A.J.; Welte, C.U. Predicting and improving the microbial removal of organic micropollutants during wastewater treatment: A review. Chemosphere 2023, 333, 138908.
- Céspedes-Bernal, D.N.; Mateus-Maldonado, J.F.; Rengel-Bustamante, J.A.; Quintero-Duque, M.C.; Rivera-Hoyos, C.M.; Poutou-Piñales, R.A.; Diaz-Ariza, L.A.; Castillo-Carvajal, L.C.; Paez-Moralez, A.; Pedroza-Rodríguez, A.M. Non-domestic wastewater treatment with fungal/bacterial consortium followed by Chlorella sp., and thermal conversion of the generated sludge. 3 Biotech 2021, 11, 1–18.
- Djelal, H.; Amrane, A. Biodegradation by bioaugmentation of dairy wastewater by fungal consortium on a bioreactor lab-scale and on a pilot-scale. J. Environ. Sci. 2013, 25, 1906–1912.
- Lee, Y.; Jeong, S.E.; Hur, M.; Ko, S.; Jeon, C.O. Construction and evaluation of a Korean native microbial consortium for the bioremediation of diesel fuel-contaminated soil in Korea. Front. Microbiol. 2018, 9, 2594.
This entry is offline, you can click here to edit this entry!
