Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is an old version of this entry, which may differ significantly from the current revision.
Subjects:
Infectious Diseases
NRF2 belongs to the cap “n” collar (CNC) family of transcription factors and is found in the cytoplasm of non-stressed cells in a combined form with KEAP1. Oxidative stress activates the transcription factor NRF2, which plays a key role in alleviating redox-induced cellular injury.
- NRF2 pathway
- respiratory viruses
- viral replication
- inflammation
1. Introduction
Respiratory viruses target the human respiratory system and cause various clinical symptoms in humans, ranging from mild upper respiratory infections to organ failure and life-threatening respiratory diseases [1,2]. The most common respiratory viruses are rhinoviruses, coronaviruses (CoVs), influenza virus, respiratory syncytial virus (RSV), parainfluenza viruses, enteroviruses, adenoviruses, and human metapneumovirus (hMPV) [3]. Each year, nearly 4 million deaths are attributed to lower respiratory tract infections, with Influenza contributing to approximately half a million of these fatalities [4]. Moreover, morbidity and mortality caused by respiratory viruses increased drastically with the emergence of the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) and the COVID-19 pandemic. Although several host factors have been found to play crucial role in the pathogenesis of respiratory viral infections, the interaction between respiratory viruses and the host cellular response remains poorly understood. Understanding antiviral host pathways and defining their role in the pathogenesis of respiratory viruses may set the foundation for novel antiviral therapies for viral respiratory diseases.
Viral respiratory infections are commonly associated with increased production of reactive oxygen and nitrogen species (ROS and RNS), leading to oxidative stress [5,6]. Subsequently, increased oxidative stress contributes to reduced host antiviral response, enhanced replication and virus-induced cell and tissue injury apoptosis, ferroptosis, inflammation, causing organ damage [6,7] and the occurrence of clinical symptoms [5,8,9]. During oxidative stress induced by respiratory viruses, the host deploys antioxidant mechanisms to control signaling pathways and reestablish cellular redox balance. Many respiratory viruses, including influenza [10,11,12,13,14,15,16,17], CoVs [18,19,20,21,22,23,24,25,26], RSV [27,28,29,30,31,32,33,34,35,36,37,38,39,40], rhinoviruses [41,42,43,44,45], enteroviruses such as Coxsackievirus B3 (CVB3) [46], EV71 [47,48,49,50], metapneumoviruses [34], and parainfluenza viruses [51,52], have been demonstrated to disrupt the cell redox homeostasis and induce the production of ROS.
However, in response to virus-induced oxidative stress, host cells deploy a strong antioxidant response characterized by the production of proteins (enzymes) and/or small molecules (vitamins C and E) that are mainly mediated by the nuclear factor erythroid 2-related factor (NRF2) to counteract the redox-induced toxicity and restore cellular redox homeostasis [53,54].
2. The NRF2 Pathway Regulates Cellular Responses to Stress
The production of ROS and activation of an antioxidant response is known to be controlled by the Kelch-like ECH-associated protein 1 (KEAP1)–NRF2 axis. This regulation occurs through intrinsic mechanisms within different cell types of the airway epithelium (e.g., nasal versus bronchial cells). NRF2 belongs to the cap “n” collar (CNC) family of transcription factors and is found in the cytoplasm of non-stressed cells in a combined form with KEAP1. In quiescent cells, an adapter protein, KEAP1, interacts with NRF2 and recruits cullin-3 (CUL3)-containing E3 ubiquitin ligase to form a complex that regulates the ubiquitination of NRF2. Consequently, polyubiquitination of NRF2 leads to NRF2 degradation via the 26S proteasome machinery, which ensures that the NRF2 level and its activity remain low during redox homeostasis [55]. Contrarily, during a viral respiratory infection or other induced oxidative stress, NRF2 escapes repression by KEAP1. The CUL3/KEAP1 complex that targets NRF2 for ubiquitination undergoes a change to a nonfunctional conformation [56,57,58,59]. Thus, upon activation, newly synthesized NRF2 is no longer ubiquitinated/degraded, rapidly accumulates, and translocates to the nucleus where it binds the small maf protein (sMaf) [56,57,58,59]. The NRF2–Maf heterodimer binds to the antioxidant response element (ARE) (or multiple Maf recognition elements (MAREs)). This interaction induces the transcription of a wide variety of antioxidant genes, including HO-1 and genes that are involved in the synthesis and recycling of glutathione (Figure 1). A heme sensor known as BTB and CNC homology 1 (BACH-1) can also bind to the ARE in a KEAP1-independent manner and directly competes with NRF2 for binding to AREs. The interaction of BACH-1 with ARE prevents NRF2 from binding to the ARE, thus repressing HO-1 [60,61,62,63,64,65] (Figure 1). HO-1 catalyzes the degradation of heme into carbon monoxide (CO), Fe2+, and biliverdin, and has antiviral properties through multiple pathways (Figure 2). Importantly, NRF2 activation appears to have an inhibitory effect on the interferon response, which is an important component of the innate immune system’s antiviral defense. The balance between NRF2 activation and the interferon response is regulated by intrinsic cell factors, for which the cell composition varies from nasal to bronchial cells, and potentially influences the susceptibility to viral infections. Notably, activation of the NRF2 pathway appears to mediate protection against viral respiratory infections, including SARS-CoV-2, influenza viruses, and several RNA and DNA viruses that induce oxidative stress (Supplementary Table S1) [5,8,9,10,11,12,13,14,18,19,20,21,22,23,24,27,28,29,30,31,32,33,34,35,36,41,42,43,44,46,47,48,49,51,52,53,54,65,66,67,68,69,70,71,72,73,74,75,76,77,78,79,80,81,82,83,84,85,86,87,88,89,90,91,92,93]. Overall, the crosstalk between NRF2 and viruses is bidirectional and complex. With regards to the impact of viruses on the NRF2 pathway, respiratory viruses such as influenza [12,94], SARS-CoV-2 [19,20,22,24,95,96], RSV [27,31,32], rhinovirus [43,44], and enteroviruses [47] directly alter NRF2 levels and signaling [5,8,9,10,18,33] (Table 1). With regards to the impact of the NRF2 pathway on viruses, host NRF2 pathways also regulate the replication of several respiratory viruses such as influenza [11,12,13], coronaviruses [19,20,21,22,23], RSV [34,35], rhinovirus [44], enterovirus [47], metapneumovirus [34], and parainfluenza [51,52] (Table 2). Notably, the host NRF2 pathways also regulate viral replication, apoptosis, ferroptosis, and inflammation (Table 2, Table 3, Table 4, Table 5 and Table 6). Besides playing an essential role in cell defense against redox stresses by trans-activating cytoprotective genes encoding antioxidant and detoxifying enzymes, NRF2 contributes to the regulation of the anti-inflammatory response and metabolic reprogramming [97].
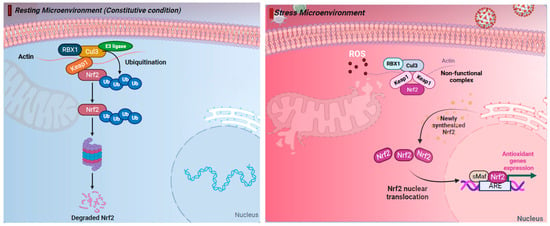
Figure 1. The nuclear factor erythroid 2-related factor 2 (NRF2) pathway regulates cellular responses to stress. (Left) Under resting (constitutive) condition, in the cytoplasm NRF2 is anchored with Kelch-like ECH-associated protein 1 (KEAP1). NRF2 binds KEAP1 and becomes ubiquinated, leading to degradation by the 26S proteasome. (Right) Under oxidative stress response, NRF2 escapes repression by KEAP1. The CUL3/KEAP1 complex that targets NRF2 for ubiquitination undergoes a change to a nonfunctional conformation. Thus, newly synthesized NRF2 is no longer ubiquitinated/degraded, rapidly accumulates, and translocates to the nucleus where it binds the small maf protein (sMaf) and antioxidant response element (ARE). Activation of ARE increases the expression of the antioxidant genes heme oxygenase 1 (HO-1), quinone oxidoreductase (NQO1), and glutathione (GSH), which blocks the progression of oxidative stress (OS). Thus, activation of the NRF2 pathway has cytoprotective effects and plays a key role in maintaining redox balance. Figure generated with Biorender (https://biorender.com/, accessed on 10 November 2023).
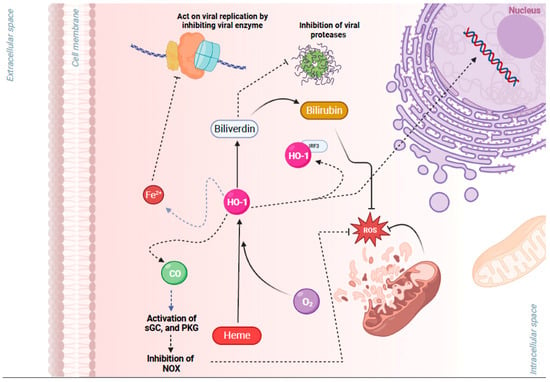
Figure 2. Cytoprotective effects of heme oxygenase 1 (HO-1), a key gene of the nuclear factor erythroid 2-related factor 2 (NRF2) pathway, in viral infection. HO-1 is a metabolic enzyme that utilizes oxygen (O2), heme, and NADPH to catalyze the degradation of heme into carbon monoxide (CO), Fe2+, and biliverdin. HO-1 has antiviral properties through multiple pathways: (1) Free Fe2+ may act on viral replication by binding to the highly conserved divalent metal-binding pocket of the viral RNA and inhibiting enzymes that mediate viral replication. (2) Biliverdin may inhibit viral proteases. (3) Heterodimerization of HO-1 with interferon regulatory factor 3 (IRF3) facilitates the phosphorylation and nuclear translocation of IRF3 and the induction of type I interferon (IFN) gene expression that has antiviral properties. (4) CO activates protein kinase G (PKG), which inhibits NAPDH oxidases (NOX), preventing an increase in reactive oxygen species (ROS) and associated damage. (5) Biliverdin also has antioxidant properties, and it is converted by NRF2/ARE-regulated gene biliverdin reductase to the potent antioxidant bilirubin. Figure generated with Biorender (https://biorender.com/, accessed on 10 November 2023).
Table 1. Impact of respiratory viruses on the NRF2 pathway.
| Viruses | Mechanism of NRF2 Activation | Reference |
|---|---|---|
| Several (respiratory) viruses | ∙ ↑ ROS and mito-ROS ↑ ARE elements and phosphorylation of p62. | [5,8,9] |
| Influenza | ∙ IV strains are thought to activate the NRF2/ARE defense pathway in vitro and in mice by inducing oxidative stress and nuclear translocation and transcriptional activity of NRF2 because transcription of the NRF2 target gene HO-1 was shown to be augmented. | [12,94] |
| SARS-CoV-2 | ∙ SARS-CoV-2 infection ↓ levels of NRF2 in epithelial cells in vitro [19,20,22]. ∙ NRF2 was ↓ in RNA Seq analysis of lung biopsies of COVID-19 patients [22]. ∙ NRF2 deficiency ↑ ACE2, enhancing viral entry and as a result viral replication [19,20]. ∙ The nonstructural SARS-CoV-2 NSP14 viral protein inhibits NRF2 through ↓ of SIRT1 [95]. ∙ The SARS-CoV-2 ORF3a viral protein recruits KEAP-1 which inhibits NRF2 activity, thereby facilitating ferroptosis through the built-up ROS and the downregulation of genes like HO-1 and NQO1 [96]. ∙ The SARS-CoV-2 ORF3a viral protein binds to human HO-1 protein in vitro [24]. |
[19,20,22,24,95,96] |
| RSV | ∙ RSV deregulates the NRF2 expression and its activity along with the upregulation of downstream ARE-responsive genes [98]. ∙ RSV ↓ mRNA levels of NRF2 in airway epithelial cells [27]. ∙ RSV ↑ NRF2 deacetylation, ubiquitination, and degradation through a proteasome-dependent pathway in a SUMO-specific E3 ubiquitin ligase RNF4-dependent manner. ∙ Another possible mechanism of RSV-associated NRF2 activation is the activation of [31], which activates the NRF2 pathway through direct alkylation of the NRF2 partner—KEAP1 [32]. |
[27,31,32,98] |
| Rhinovirus | ∙ Rhinovirus RNA stimulates the innate immune sensor RIG-I within airway epithelial cells and activates the antiviral interferon response (greater activation in nasal cells than in bronchial cells) and the NRF2-mediated response to oxidative stress (greater activation in bronchial cells compared to nasal cells) [43]. ∙ However, the inhibitory effects were reversed in cells pretreated with the antioxidant, N-acetyl cysteine. Moreover, the secretion of anti-viral interferons ↑ in cells treated with the NRF2 agonist sulforaphane but ↓ in cells where NRF2 was silenced [44]. |
[43,44] |
| Enterovirus 71 (EV71) | ∙ EV71 ↑ KEAP1 and ↓ NRF2 [47]. | [47] |
| RSV, influenza, coronaviruses, HCV | ∙ ↑ phosphorylation of the redox-sensitive PKC ↑ NRF2 dissociation from KEAP1. | [10,18,33] |
Abbreviations: ACE2: angiotensin-converting enzyme, ARE: antioxidant response element, COX-2: cyclooxygenase 2, EBV: Epstein Barr Virus, ER: endoplasmic reticulum, EV71: Enterovirus 71, GSK3: glycogen synthase kinase 3, HBV: Hepatitis B virus, HCV: Hepatitis C virus, HO-1: Heme Oxygenase 1, IRG1: immune-responsive gene 1, IV: influenza virus, mito-ROS: mitochondrial ROS, KEAP1: Kelch-like ECH-associated protein 1, KSHV: Kaposi sarcoma-associated herpesvirus, NQO1: NAD(P)H quinone oxidoreductase, NRF2: nuclear factor erythroid 2-related factor, NSP14: nonstructural protein 14, ORF3a: 3a open reading frame 3a, PKC: protein kinase C, PI3K: phosphatidylinositol 3-kinase (PI3K), PGE2: Prostaglandin E2. RNF4: RING finger protein 4, ROS: reactive oxygen species, RSV: respiratory syncytial virus, SIRT1: sirtuin 1, SFTS: severe fever with thrombocytopenia syndrome, sMafs: small Maf proteins, SUMO: small ubiquitin-like modifiers.
Table 2. Impact of NRF2 on the replication of respiratory viruses.
| Virus | Role of NRF2 in Viral Replication | Reference |
|---|---|---|
| Influenza virus | ∙ Activation of NRF2 leads to ↓ viral replication through interferon host responses ∙ Downregulation of NRF2 results in major ↑ of viral entry and, subsequently, viral replication. ∙ Inhibition of viral replication, growth, and protein expression after activation of NRF2. Influenza also regulates autophagy, which interacts with NRF2 and is involved in influenza replication [99]. |
[11,12,13] |
| Coronaviruses | ∙ NRF2 deficiency upregulates ACE2, ↑ viral entry, and, as a result, viral replication. ∙ NRF2 induced production of HO-1 and generated Fe+2, which binds to the RNA- dependent RNA polymerase of SARS-CoV-2, inhibiting its activity and thus viral replication. ∙ NRF2 agonists like 4-OI and DMF inhibit SARS-CoV-2 replication. ∙ Vero cells infected with SARS-CoV-2 and transfected with siRNA to silence KEAP-1, thereby activating NRF2, had a decreased viral load. ∙ Absence of NRF2 in knockout mice ↑ the severity of SARS-CoV-2 infection and viral replication. |
[19,20,21,22,23] |
| RSV | ∙ NRF2 knockout mice showed significantly ↑ viral titers in the lungs. ∙ Treatment of the NRF2 agonist sulforaphane on NRF2−/− and NRF2+/+ mice before RSV infection ↓ virus replication, but this significant effect was not observed in NRF2−/− mice [37]. ∙ Compared to wild-type mice, RSV-infected NRF2 KO had ↓ antioxidant enzymes and enzymes in the airway, which modulated the endogenous hydrogen sulfide (H2S) pathway that has a significant antiviral function [34]. ∙ Inducers of the NRF2-ARE pathway, such as BHA treatment, ↑ the viral clearance in murine lungs [35]. |
[34,35,37] |
| Rhinovirus | ∙ Silence of NRF2 in cells led to a ↓ in the secretion of antiviral interferons and higher viral titers. | [44] |
| Enterovirus 71 (EV71) | ∙ Silencing of NRF2 is beneficial for viral replication. ∙ Activating NRF2 through downregulation of KEAP1 led to ↓ viral replication in RD cells. |
[47] |
| Metapneumovirus | ∙ ↓ of NRF2-dependent genes ↑ viral replication and clinical disease upon hMPV infection. | [34] |
| Parainfluenza viruses | ∙ Cotreatment or post infection treatment with curcumin, ↓ the expression of HN viral protein, indicating that curcumin may ↓ viral entry affecting viral replication and subsequently different steps in viral replication ∙ Curcumin, an NRF2 activator, led to ↓ of F-actin, ↓ the formation of viral IBs, and ↓ viral replication. ∙ Curcumin ↓ HPIV3 replication by ↓ the endogenous PI4KB level in the cells, and ↓ the colocalization of PI4KB and IBs, affecting IB formation. |
[51,52] |
Abbreviations: ACE2: angiotensin-converting enzyme, ARE: antioxidant response element, BHA: butylated hydroxyanisole, DMF: Dimethyl fumarate, EV-71: Enterovirus 71, hMPV: Human Metapneumovirus, HO-1: Heme Oxygenase 1, HPIV3: Human Parainfluenza virus 3, IBs: inclusion bodies, KEAP1: Kelch-like ECH-associated protein 1, NRF2: nuclear factor erythroid 2-related factor, 4-OI: 4-octyl itaconate, PI4KB: 1-phosphatidylinositol 4-kinase beta, RD: Rhabdomyosarcoma, RSV: respiratory syncytial virus, SARS-CoV-2: severe acute respiratory syndrome-coronavirus 2, siRNA: Small interfering RNA.
Table 3. Antiviral activity of Heme Oxygenase 1 (HO-1), a key downstream gene of the NRF2 pathway.
| Viruses/NRF2 | HO-1 Antiviral Activity | Reference |
|---|---|---|
| Influenza | ∙ ↑ expression of HO-1 leads to ↓ viral replication during infection from Influenza A, through the upregulation of IFN- α/β and ISGs. | [14] |
| RSV | ∙ Harmacological activation of HO-1 by CoPP ↓ viral replication of RSV in lung cells of infected mice via induction of IFN-α/β expression. ∙ In vitro data suggest that ↑ of HO-1 can moderate the susceptibility of cells to hRSV infection [36]. |
[36] |
| HCV | ∙ Type 1 IFN-dependent anti-HCV activity due to ↑ levels of HO-1 resulting from the usage of HO-1 agonists/inducers. ∙ Iron stopped viral replication of HCV by direct binding to the Mg+2 binding pocket of the RNA polymerase of the virus. |
[88,89,90,91,92] |
| Coronaviruses | ∙ Fe+2 binds to the RNA-dependent RNA polymerase of SARS-CoV-2 inhibiting its activity and ↓ viral replication. ORF3a protein binds to human HMOX1 protein in vitro [24]. |
[19] |
| EV71 | ∙ The overexpression of HO-1 ↓ NADPH oxidase/ROS production that is induced by enterovirus 71 and hence ↓ viral replication. This effect was abolished if cells were pretreated with zinc–protoporphyrin IX, an HO-1 activity inhibitor. ∙ Bilirubin has also been found to exert antiviral activity against EV71 reducing its replication and as a result infectivity in vitro. |
[48,49] |
| HIV | ∙ BV and BR have been identified to act as inhibitors for the protease of HIV, interfering with the life cycle of the virus. | [93] |
Abbreviations: CoPP: cobalt protoporphyrin, EV-71: Enterovirus 71, HCV: Hepatitis C virus, HO-1: Heme Oxygenase 1, ISGs: interferon stimulated genes. NRF2: nuclear factor erythroid 2-related factor, SARS-CoV-2: severe acute respiratory syndrome coronavirus 2, RSV: respiratory syncytial virus.
Table 4. Respiratory viruses and the role of NRF2 role in inflammation.
| Virus | Role of NRF2 in Inflammation | References |
|---|---|---|
| Influenza virus | ∙ Inactivation of NF-κΒ transcription factor. ∙ ↓ of NF-κΒ-mediated inflammation and associated lung permeability damage, mucus hypersecretion, lung permeability damage, as well as mucus hypersecretion, through reduced NF-κΒ-mediated inflammation and associated proinflammatory cytokines [15]. ∙ Induces anti-inflammatory effects in vivo through the HO-1 pathway [100,101]. ∙ NLRP3 activation form PB1-F2 influenza A protein. ∙ K+ efflux and ROS dependent activation of NLRP3 inflammasome ∙ Impacts function of alveolar macrophages (AMϕ) that are important essential for preventing respiratory failure and mortality after infection from influenza virus in mice [17]. ∙ Attenuates virus-induced inflammation through increased GSH levels and IL-8 secretion in ATI-like cells (alveolar epithelial cells) in vitro [12]. |
[15,16,17] |
| Coronavirus | ∙ NRF2 is directly able to inhibit IL6, IL-1B, a key hallmark of the cytokine storm in SARS-CoV-2 infection. ∙ Absence of NRF2 in knockout mice ↑ the severity of SARS-CoV-2 infection, pulmonary inflammation. ∙ In humans, SNPs in the Nrf2 gene promoter region can determine susceptibility to respiratory failure with COPD, indicating the importance of NRF2 in pulmonary inflammation. ∙ Cytokine storm due to T cell depletion and widespread pulmonary inflammation. ∙ Contradictory effect on proinflammatory nature of factors like NF-kB. |
[23,25,26] |
| RSV | ∙ Severe inflammation in NRF2−/− mice compared to NRF2+/+ mice. ∙ RSV-infected NRF2 KO mice are reported to have a significant ↑ in airway neutrophilia and inflammatory cytokines. ∙ ↓ lung inflammation when pretreated with sulforaphane. ∙ ↓ ROS- and K+ efflux-dependent activation of NLRP3 inflammasome. ∙ SH viroporin activates NLRP3 inflammasome. ∙ Impacts function of alveolar macrophages (AMϕ), which are important to attenuate virus-induced inflammation. |
[37,38,39,40,102] |
| Metapneumovirus | ∙ NRF2 KO mice infected with hMPV had ↓ expression of antioxidant enzymes (AOE) and ↑ viral-mediated oxidative stress and airway damage compared to NRF2+/+ mice. | [34] |
| Enterovirus 71 | ∙ By silencing KEAP1, the induced ROS, apoptosis, and inflammation was ↓ in the EV71 infected cells. However, when both KEAP1 and NRF2 were silenced in Vero and RD cells, these effects were restored. ∙ Inflammation-promoting cytokines and chemokines influence the severity of the EV71 infection. |
[47,50] |
| Rhinovirus | ∙ 2B viroporin activates NLRP3 inflammasome | [16,45] |
Abbreviations: EV-71: Enterovirus 71, KEAP1: Kelch-like ECH-associated protein 1, hMPV: human metapneumovirus, NRF2: nuclear factor erythroid 2-related factor, NRF2-KO: NRF2 knocked out; NF-κB: nuclear factor kappa B, ROS: reactive oxygen species, RSV: respiratory syncytial virus.
Table 5. Impact of viruses and NRF2 on the apoptosis pathway.
| Viruses/NRF2 | Impact on Apoptosis | Reference |
|---|---|---|
| Adenoviruses | ∙ Complex effects. ∙ ↑ apoptosis: ↑ sensitivity to TNFa that induces apoptosis, ↑ PP2A, and ↑ p53. ∙ ↓ apoptosis through several mechanisms: interacts with FADD, ↓ CD95-mediated apoptosis, ↓ phospholipase A2, ↓ Fas, ↓ p53, and ↓ pro-apoptotic proteins of the Bcl-2 family, such as Bax, Bak, BNIP3, and Bnip3L. ∙ ↓ apoptosis of the host cell in order to ↑ efficiently and the capacity of the virus to ‘hijack’ host cell apoptotic machinery. |
[103,104,105,106] |
| RSV | ∙ ↑ interferons and caspase 1. ∙ Experimental studies have shown that autophagy plays a very crucial role in RSV replication [107]. |
[105] |
| Influenza | ∙ ↑ Fas expression. ∙ ↓ PKR and apoptosis. ∙ Apoptosis plays a role in viral release. |
[105,106] |
| Rhinovirus, enteroviruses | ∙ ↑ apoptosis through unknown mechanism. | [105] |
| Coronaviruses | ∙ ↑ apoptosis through ORF proteins and unknown mechanisms. | [105] |
Abbreviations: FADD: Fas-associated death domain protein, GPX4: Glutathione Peroxidase 4, HO-1: Heme Oxygenase 1, NRF2: nuclear factor erythroid 2-related factor, ORF: Open Reading Frame, PKR: protein kinase R, p53: tumor protein 53, PP2A: Protein Phosphatase 2A, ROS: reactive oxygen species, RSV: respiratory syncytial virus, TNFa: tumor necrosis factor a.
Table 6. Impact of viruses and NRF2 on the ferroptosis pathway.
| Viruses/NRF2 | Impact on Ferroptosis | Reference |
|---|---|---|
| NRF2 | ∙ NRF2 ↓ ROS and ↑ antioxidant responses, and ↑ GPX4-induced ↓ of ferroptosis. ∙ NRF2 ↑ Heme Oxygenase 1 (HO-1) that ↓ ferroptosis. ∙ NRF2 ↑ antioxidant enzymes. |
[9] |
| Influenza | ∙ Iron ↓ viral genome amplification and viral replication. ∙ Influenza ↓ cellular GSH and/or affects GPX4 activity. ∙ Neuraminidase of Influenza A binds lysosome-associated membrane proteins and ↑ lysosome rupture. |
[9,108,109,110,111,112] |
| SARS-CoV-2, SARS-CoV, other coronaviruses | ∙ SARS-CoV-2 Potentially causes cellular iron overload and iron scavenging. ∙ SARS-CoV-2 ↑serum ferritin concentration. ∙ CoVs↓ cellular GSH and/or affect GPX4 activity. ∙ SARS-CoV ORF-3a viral protein ↑ lysosomal damage and dysfunction. |
[113,114,115,116,117] |
| RSV | ∙ ↑ the expression of 12/15-LOX and mitochondrial iron content. | [118] |
| EV-71, CB3 | ∙ Iron ↓ viral genome amplification and viral replication of EV-71. ∙ CB3 ↑the expression NRAMP (DMT) and ↑cellular iron uptake. |
[108,119,120] |
| Non-respiratory viruses: HBV, HCV, WNV, Dengue virus, HSV, KSHV |
∙ HBV, HCV: ↑ serum and cellular iron uptake and ↓ hepcidin expression, ↑ serum ferritin concentration, and uses TfR1 as a cellular receptor. ∙ HIV ↓ serum iron, ↑ the expression of hepcidin, ↑cellular iron via hepcidin mediated degradation of ferroportin, ↓ cellular GSH and/or affects GPX4 activity, and upregulates the expression of system xc-. ∙ WNV ↑ the expression NRAMP (DMT) and ↑cellular iron uptake. ∙ Dengue virus ↓ cellular GSH and/or affects GPX4 activity. ∙ HSV ↓ cellular GSH and/or affects GPX4 activity. ∙ JEV ↓ cellular GSH and/or affects GPX4 activity, produces lipid peroxide free radicals, and ↑ the expression of system xc-. ∙ KSHV ↓ cellular GSH and/or affects GPX4 activity. ∙ Zika virus ↓ cellular GSH and/or affects GPX4 activity. ∙ Other viruses (e.g., hemorrhagic viruses) use NRAMP or TfR1 as a cellular receptor. |
[121] |
Abbreviations: CB3: Coxsackievirus B3, EV-71: Enterovirus 71, GSH: Glutathione, GPX4: Glutathione Peroxidase 4, HBV: Hepatitis B virus, HCV: Hepatitis C virus, HIV: human immunodeficiency virus, HSV: Herpes simplex virus, JEV: Japanese encephalitis virus, KSHV: Kaposi sarcoma-associated herpesvirus, 12/15-LOX:12/15-lipoxygenase, NRAMP: Natural Resistance-Associated Macrophage Proteins, ORF-3a: Open Reading Frame-3a, RSV: respiratory syncytial virus, SARS-CoV-2: severe acute respiratory syndrome-coronavirus-2, SARS-CoV: severe acute respiratory syndrome coronavirus, TfR1: transferrin receptor 1 protein, WNV: West Nile virus.
This entry is adapted from the peer-reviewed paper 10.3390/pathogens13010039
This entry is offline, you can click here to edit this entry!
