Alzheimer’s disease (AD) is the most common form of dementia that affects millions of individuals worldwide. Although the research over the last decades has provided new insight into AD pathophysiology, there is currently no cure for the disease. AD is often only diagnosed once the symptoms have become prominent, particularly in the late-onset (sporadic) form of AD. Consequently, it is essential to further new avenues for early diagnosis. With recent advances in genomic analysis and a lower cost of use, the exploration of genetic markers alongside RNA molecules can offer a key avenue for early diagnosis.
- Alzheimer’s disease
- genes
- genetics
- Late Onset Alzheimer's Disease
- Sporadic Alzheimer's Disease
- RNA
1. Introduction
With recent advances in technology leading to lower costs, genetic screening has become increasingly utilised in the diagnosis of a range of diseases, including for the diagnosis of progressive neurological disorders such as Spino–Cerebellar Ataxia (SCA) [15]. Although the familial form of AD has strong genetic origins with well-characterised pathogenic variants (Amyloid Protein Precursor (APP), Presenilin-1 (PSEN1) and Presenilin-2 (PSEN2) [16,17]), the genetic landscape of sporadic AD is more complex and remains less well understood. Taking into account the multifactorial aetiology of AD (including environmental factors) [1], it is essential to consider not only the multiple genes involved in AD but also the regulation of the transcriptome. Small non-coding molecules, such as microRNAs, circular-RNAs, and long non-coding RNAs, can regulate gene expression at the transcriptional and post-transcriptional level. Consequently, it is essential to further explore the interplay between the genome and its regulation via non-coding RNAs, which may be influenced by and respond to environmental factors.
2. AD Genetic Susceptibility
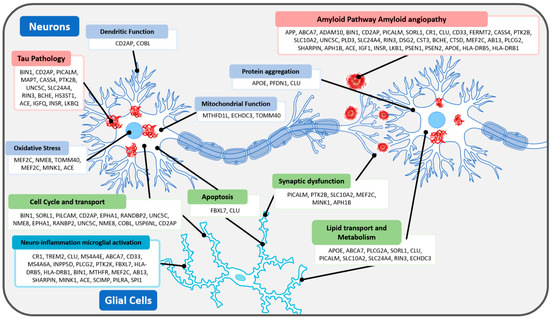
| Mechanism of AD Pathophysiology |
Genes |
|---|---|
| Amyloid Pathway Amyloid angiopathy |
APP, ABCA7, ADAM10, BIN1, CD2AP, PICALM, SORL1, CR1, CLU, CD33, FERMT2, CASS4, PTK2B, SLC10A2, UNC5C, PLD3, SLC24A4, RIN3, DSG2, CST3, BCHE, CTSD, MEF2C, AB13, PLCG2, SHARPIN, APH1B, ACE, IGF1, INSR, LKB1, PSEN1, PSEN2, APOE, HLA-DRB5, HLA-DRB1 |
| Tau Pathology | BIN1, CD2AP, PICALM, MAPT, CASS4, PTK2B, UNC5C, SLC24A4, RIN3, BCHE, HS3ST1, ACE, IGFQ, INSR, LKBQ |
| Lipid transport Lipid Metabolism |
APOE, ABCA7, PLCG2A, SORL1, CLU, PICALM, SLC10A2, SLC24A4, RIN3, ECHDC3 |
| Dendrites | CD2AP, COBL |
| Neuroinflammation microglial activation |
CR1, TREM2, CLU, MS4A4E, ABCA7, CD33, MS4A6A, INPP5D, PLCG2, PTK2K, FBXL7, HLA-DRB5, HLA-DRB1, BIN1, MTHFR, MEF2C, AB13, SHARPIN, MINK1, ACE, SCIMP, PILRA, SPI1 |
| Protein aggregation | APOE, PFDN1, CLU |
| Mitochondrial Function | MTHFD1L, ECHDC3, TOMM40 |
| Synaptic dysfunction | PICALM, PTK2B, SLC10A2, MEF2C, MINK1, APH1B |
| Blood Brain Barrier disruption, vascular damage | CD2AP, EPHA1, MTHFR |
| Apoptotic genes | FBXL7, CLU |
| Oxidative Stress | MEF2C, NME8, TOMM40, MEF2C, MINK1, ACE |
| Cell Cycle and transport | BIN1, SORL1, PILCAM, CD2AP, EPHA1, RANDBP2, UNC5C, NME8, EPHA1, RANBP2, UNC5C, NME8, COBL, USP6NL, CD2AP |
2.1. Genes Related to AD Hallmarks of Pathophysiology
2.2. Candidate Genes Involved in the Regulation of AD Pathways
| Ranking | Gene | p-Value | Source |
|---|---|---|---|
| 1 | BIN1 | 2.06 × 10−30 | [18] |
| 1.10 × 10−54 | [53] | ||
| 7.30 × 10−49 | [54] | ||
| 2 | PICALM | 4.29 × 10−14 | [18] |
| 5.21 × 10−26 | [53] | ||
| 5.10 × 10−36 | [54] | ||
| 3 | CLU | 6.60 × 10−13 | [18] |
| 7.71 × 10−26 | [53] | ||
| 1.10 × 10−28 | [54] | ||
| 4 | CR1 | 1.51 × 10−14 | [18] |
| 1.40 × 10−23 | [53] | ||
| 1.60 × 10−28 | [54] | ||
| 5 | MS4A | 2.87 × 10−20 | [18] |
| 9.33 × 10−20 | [53] | ||
| 1.10 × 10−18 | [54] | ||
| 6 | TREM2 | 2.34 × 10−11 | [18] |
| 1.83 × 10−23 | [53] | ||
| 7 | PILRA | 7.41 × 10−10 | [18] |
| 3.28 × 10−18 | [53] | ||
| 1.10 × 10−18 | [54] | ||
| 8 | SORL1 | 1.76 × 10−8 | [18] |
| 5.59 × 10−14 | [53] | ||
| 4.80 × 10−17 | [54] | ||
| 9 | HLA | 2.88 × 10−15 | [53] |
| 1.20 × 10−14 | [54] | ||
| 10 | CD2AP | 2.95 × 10−12 | [18] |
| 1.11 × 10−11 | [53] | ||
| 5.80 × 10−14 | [54] | ||
| 11 | ABCA7 | 2.36 × 10−9 | [18] |
| 2.41 × 10−13 | [53] | ||
| 3.70 × 10−10 | [54] | ||
| 12 | SLC24A4 | 3.55 × 10−7 | [18] |
| 7.45 × 10−14 | [53] | ||
| 1.10 × 10−10 | [54] | ||
| 14 | CLNK/EPHA1/ECHDC3/HS3ST1 | 5.02 × 10−8 | [18] |
| 1.08 × 10−11 | [53] | ||
| 3.40 × 10−7 | [54] | ||
| 15 | ADAM10 | 2.67 × 10−11 | [53] |
| 5.50 × 10−11 | [54] | ||
| 16 | CASS4 | 1.56 × 10−6 | [18] |
| 1.07 × 10−10 | [53] |
3. RNA Regulation
3.1. Micro-RNAs
3.2. Emerging Regulators of the Transcriptome: circRNAs and lncRNAs
4. Summary
This entry is adapted from the peer-reviewed paper 10.3390/ijms241713480
