Amino acids have been extensively studied in nutrition, mainly as key elements for maintaining optimal protein synthesis in the body as well as precursors of various nitrogen-containing compounds. However, it is now known that amino acid catabolism is an important element for the metabolic control of different biological processes, although it is still a developing field to have a deeper understanding of its biological implications. The mechanisms involved in the regulation of amino acid catabolism now include the contribution of the gut microbiota to amino acid oxidation and metabolite generation in the intestine, the molecular mechanisms of transcriptional control, and the participation of specific miRNAs involved in the regulation of amino acid degrading enzymes. In addition, molecules derived from amino acid catabolism play a role in metabolism as they are used in the epigenetic regulation of many genes.
- amino acid catabolism
- gut microbiota
- gene transcription
- epigenetics
- immune response
- obesity
- diabetes
- thermogenesis
1. Introduction
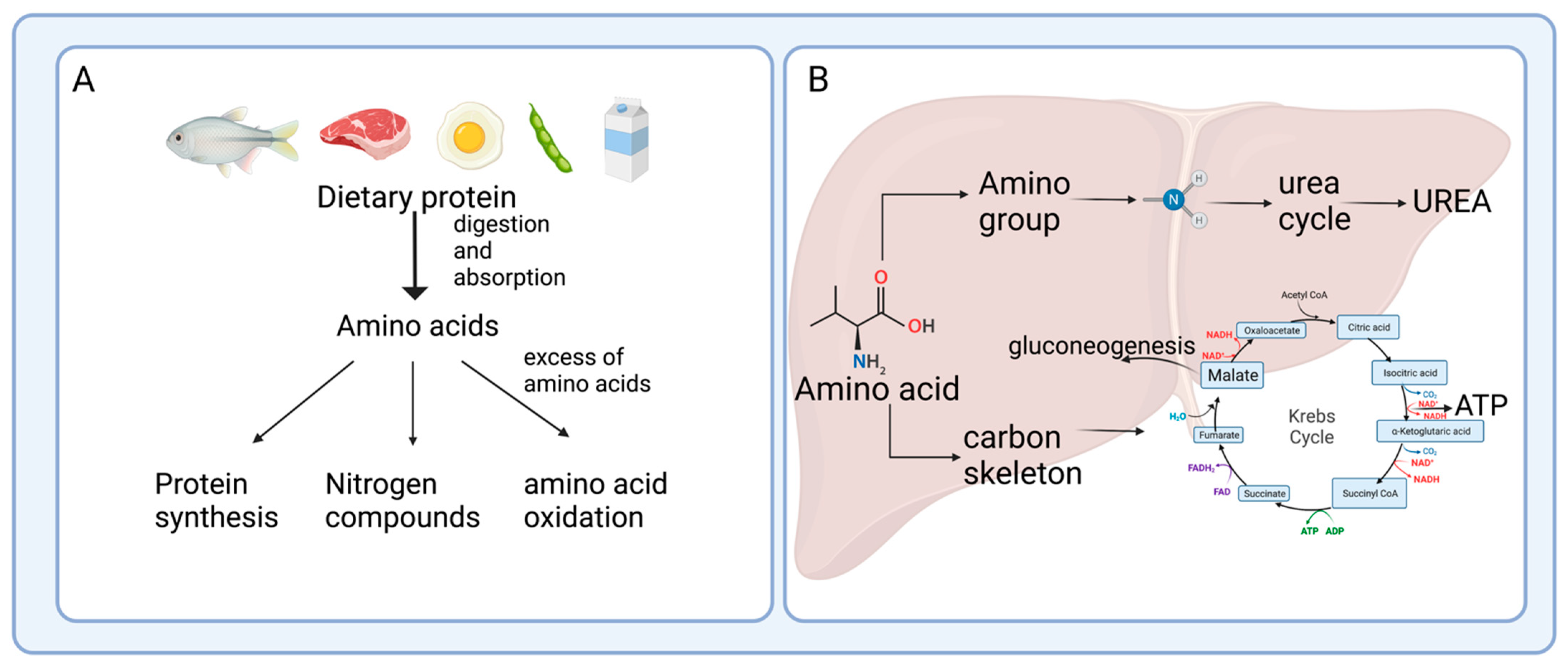
2. Enterohepatic Axis of Amino Acid Catabolism
2.1. Protein Hydrolysis and Amino Acid Absorption in the Gut
2.2. Amino Acid Catabolism in the Intestine
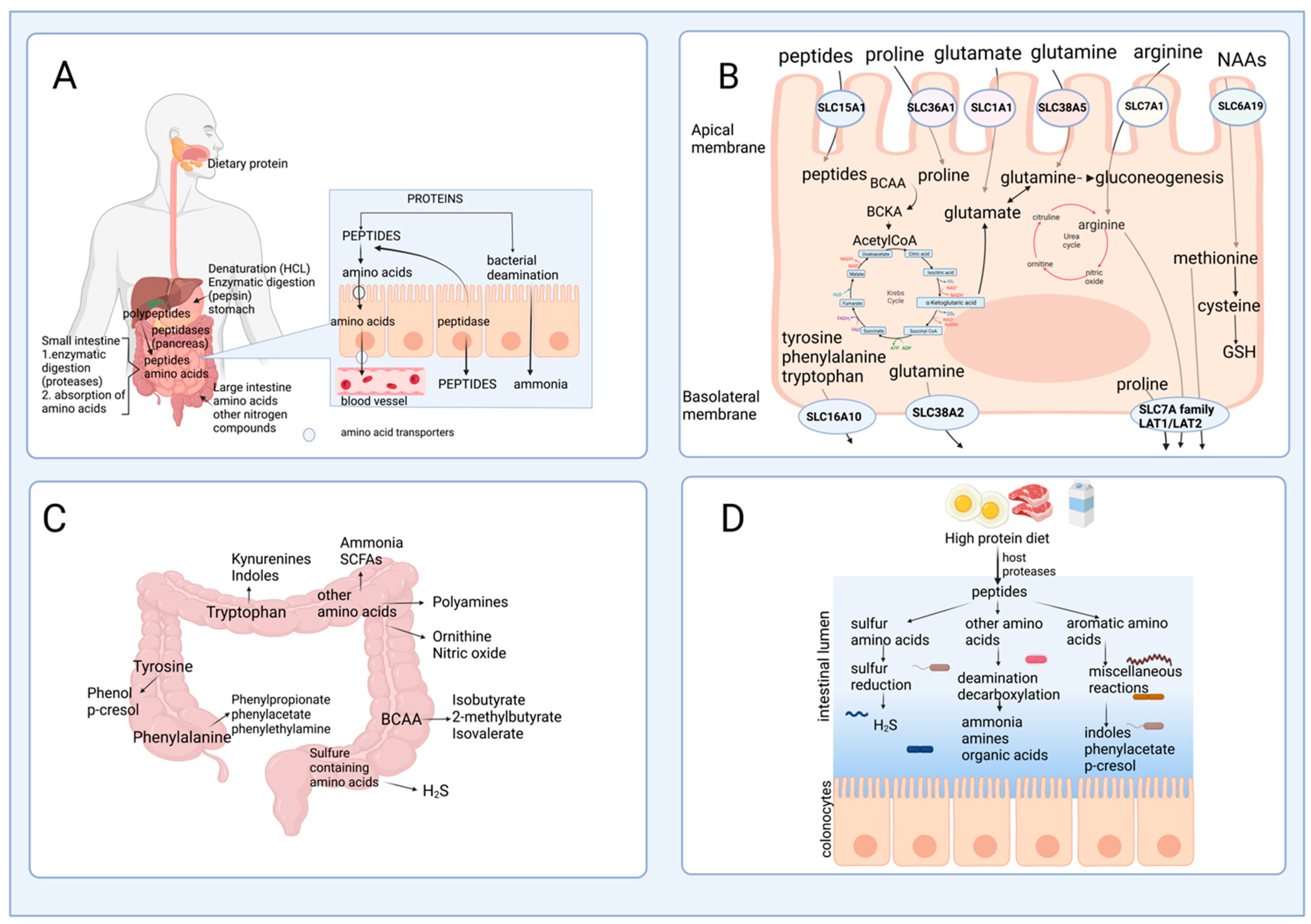
2.3. The Role of Gut Microbiota in Amino Acid Catabolism and Generation of End-Products
3. Mechanisms of Regulation of Amino Acid Oxidation
4. Amino Acid Catabolism in Health and Disease
4.1. Amino Acid Catabolism and Immunity
4.2. Amino Acid Catabolism in Obesity and Diabetes
4.3. Amino Acid Catabolism and Thermogenesis
This entry is adapted from the peer-reviewed paper 10.3390/nu15153378
References
- Chou, C.J.; Affolter, M.; Kussmann, M. A nutrigenomics view of protein intake: Macronutrient, bioactive peptides, and protein turnover. Prog. Mol. Biol. Transl. Sci. 2012, 108, 51–74.
- Harper, A.E.; Miller, R.H.; Block, K.P. Branched-chain amino acid metabolism. Annu. Rev. Nutr. 1984, 4, 409–454.
- Bender, D.A. The metabolism of “surplus” amino acids. Br. J. Nutr. 2012, 108 (Suppl. S2), S113–S121.
- Bartlett, A.; Kleiner, M. Dietary protein and the intestinal microbiota: An understudied relationship. iScience 2022, 25, 105313.
- Tome, D. Amino acid metabolism and signalling pathways: Potential targets in the control of infection and immunity. Nutr. Diabetes 2021, 11, 20.
- Broer, S. Amino acid transport across mammalian intestinal and renal epithelia. Physiol. Rev. 2008, 88, 249–286.
- Loveday, S.M. Protein digestion and absorption: The influence of food processing. Nutr. Res. Rev. 2022, 16, 1–16.
- Bertrand, J.; Goichon, A.; Dechelotte, P.; Coeffier, M. Regulation of intestinal protein metabolism by amino acids. Amino Acids 2013, 45, 443–450.
- Sleisenger, M.H.; Kim, Y.S. Protein digestion and absorption. N. Engl. J. Med. 1979, 300, 659–663.
- Davila, A.M.; Blachier, F.; Gotteland, M.; Andriamihaja, M.; Benetti, P.H.; Sanz, Y.; Tome, D. Intestinal luminal nitrogen metabolism: Role of the gut microbiota and consequences for the host. Pharmacol. Res. 2013, 68, 95–107.
- Chen, C.; Yin, Y.; Tu, Q.; Yang, H. Glucose and amino acid in enterocyte: Absorption, metabolism and maturation. Front. Biosci. 2018, 23, 1721–1739.
- Torres, N.; Lopez, G.; De Santiago, S.; Hutson, S.M.; Tovar, A.R. Dietary protein level regulates expression of the mitochondrial branched-chain aminotransferase in rats. J. Nutr. 1998, 128, 1368–1375.
- Ma, N.; Ma, X. Dietary Amino Acids and the Gut-Microbiome-Immune Axis: Physiological Metabolism and Therapeutic Prospects. Compr. Rev. Food Sci. Food Saf. 2019, 18, 221–242.
- Pink, D.B.; Gatrell, S.K.; Elango, R.; Turchinsky, J.; Kiess, A.S.; Blemings, K.P.; Dixon, W.T.; Ball, R.O. Lysine alpha-ketoglutarate reductase, but not saccharopine dehydrogenase, is subject to substrate inhibition in pig liver. Nutr. Res. 2011, 31, 544–554.
- Mitchell, A.D.; Benevenga, N.J. The role of transamination in methionine oxidation in the rat. J. Nutr. 1978, 108, 67–78.
- Cooper, A.J.; Meister, A. Comparative studies of glutamine transaminases from rat tissues. Comp. Biochem. Physiol. B Biochem. 1981, 69, 137–145.
- Schaart, M.W.; Schierbeek, H.; van der Schoor, S.R.; Stoll, B.; Burrin, D.G.; Reeds, P.J.; van Goudoever, J.B. Threonine utilization is high in the intestine of piglets. J. Nutr. 2005, 135, 765–770.
- Hoerr, R.A.; Matthews, D.E.; Bier, D.M.; Young, V.R. Effects of protein restriction and acute refeeding on leucine and lysine kinetics in young men. Am. J. Physiol. 1993, 264, E567–E575.
- Wu, G. Synthesis of citrulline and arginine from proline in enterocytes of postnatal pigs. Am. J. Physiol. 1997, 272, G1382–G1390.
- Bush, J.A.; Wu, G.; Suryawan, A.; Nguyen, H.V.; Davis, T.A. Somatotropin-induced amino acid conservation in pigs involves differential regulation of liver and gut urea cycle enzyme activity. J. Nutr. 2002, 132, 59–67.
- Morris, S.M., Jr. Regulation of enzymes of the urea cycle and arginine metabolism. Annu. Rev. Nutr. 2002, 22, 87–105.
- Wu, G.; Morris, S.M., Jr. Arginine metabolism: Nitric oxide and beyond. Biochem. J. 1998, 336 Pt 1, 1–17.
- Kan, C.C.; Chung, T.Y.; Juo, Y.A.; Hsieh, M.H. Glutamine rapidly induces the expression of key transcription factor genes involved in nitrogen and stress responses in rice roots. BMC Genom. 2015, 16, 731.
- Mouille, B.; Robert, V.; Blachier, F. Adaptative increase of ornithine production and decrease of ammonia metabolism in rat colonocytes after hyperproteic diet ingestion. Am. J. Physiol. Gastrointest. Liver Physiol. 2004, 287, G344–G351.
- Piper, D.W.; Fenton, B.H. pH stability and activity curves of pepsin with special reference to their clinical importance. Gut 1965, 6, 506–508.
- Sanchez-Tapia, M.; Moreno-Vicencio, D.; Ordaz-Nava, G.; Guevara-Cruz, M.; Granados-Portillo, O.; Vargas-Castillo, A.; Torres, N.; Tovar, A.R. Antibiotic Treatment Reduces the Health Benefits of Soy Protein. Mol. Nutr. Food Res. 2020, 64, e2000532.
- Dai, Z.L.; Li, X.L.; Xi, P.B.; Zhang, J.; Wu, G.; Zhu, W.Y. Metabolism of select amino acids in bacteria from the pig small intestine. Amino Acids 2012, 42, 1597–1608.
- Dai, Z.L.; Zhang, J.; Wu, G.; Zhu, W.Y. Utilization of amino acids by bacteria from the pig small intestine. Amino Acids 2010, 39, 1201–1215.
- Fan, P.; Liu, P.; Song, P.; Chen, X.; Ma, X. Moderate dietary protein restriction alters the composition of gut microbiota and improves ileal barrier function in adult pig model. Sci. Rep. 2017, 7, 43412.
- Windey, K.; De Preter, V.; Verbeke, K. Relevance of protein fermentation to gut health. Mol. Nutr. Food Res. 2012, 56, 184–196.
- Torrallardona, D.; Harris, C.I.; Coates, M.E.; Fuller, M.F. Microbial amino acid synthesis and utilization in rats: Incorporation of 15N from 15NH4Cl into lysine in the tissues of germ-free and conventional rats. Br. J. Nutr. 1996, 76, 689–700.
- Macfarlane, G.T.; Allison, C.; Gibson, S.A.; Cummings, J.H. Contribution of the microflora to proteolysis in the human large intestine. J. Appl. Bacteriol. 1988, 64, 37–46.
- Blachier, F.; Mariotti, F.; Huneau, J.F.; Tome, D. Effects of amino acid-derived luminal metabolites on the colonic epithelium and physiopathological consequences. Amino Acids 2007, 33, 547–562.
- Macfarlane, G.T.; Gibson, S.A.; Beatty, E.; Cummings, J.H. Estimation of short-chain fatty acid production from protein by human intestinal bacteria based on branched-chain fatty acid measurements. FEMS Microbiol. Lett. 1992, 101, 81–88.
- Desvergne, B.; Michalik, L.; Wahli, W. Transcriptional regulation of metabolism. Physiol. Rev. 2006, 86, 465–514.
- Vanweert, F.; Schrauwen, P.; Phielix, E. Role of branched-chain amino acid metabolism in the pathogenesis of obesity and type 2 diabetes-related metabolic disturbances BCAA metabolism in type 2 diabetes. Nutr. Diabetes 2022, 12, 35.
- Newsholme, P. Cellular and metabolic mechanisms of nutrient actions in immune function. Eur. J. Clin. Nutr. 2021, 75, 1328–1331.
- Kelly, B.; Pearce, E.L. Amino Assets: How Amino Acids Support Immunity. Cell Metab. 2020, 32, 154–175.
- Kedia-Mehta, N.; Finlay, D.K. Competition for nutrients and its role in controlling immune responses. Nat. Commun. 2019, 10, 2123.
- Loftus, R.M.; Finlay, D.K. Immunometabolism: Cellular Metabolism Turns Immune Regulator. J. Biol. Chem. 2016, 291, 1–10.
- Calder, P.C. Branched-chain amino acids and immunity. J. Nutr. 2006, 136, 288S–293S.
- McGaha, T.L.; Huang, L.; Lemos, H.; Metz, R.; Mautino, M.; Prendergast, G.C.; Mellor, A.L. Amino acid catabolism: A pivotal regulator of innate and adaptive immunity. Immunol. Rev. 2012, 249, 135–157.
- Guevara-Cruz, M.; Vargas-Morales, J.M.; Mendez-Garcia, A.L.; Lopez-Barradas, A.M.; Granados-Portillo, O.; Ordaz-Nava, G.; Rocha-Viggiano, A.K.; Gutierrez-Leyte, C.A.; Medina-Cerda, E.; Rosado, J.L.; et al. Amino acid profiles of young adults differ by sex, body mass index and insulin resistance. Nutr. Metab. Cardiovasc. Dis. 2018, 28, 393–401.
- Medina-Vera, I.; Sanchez-Tapia, M.; Noriega-Lopez, L.; Granados-Portillo, O.; Guevara-Cruz, M.; Flores-Lopez, A.; Avila-Nava, A.; Fernandez, M.L.; Tovar, A.R.; Torres, N. A dietary intervention with functional foods reduces metabolic endotoxaemia and attenuates biochemical abnormalities by modifying faecal microbiota in people with type 2 diabetes. Diabetes Metab. 2019, 45, 122–131.
- Wang, T.J.; Larson, M.G.; Vasan, R.S.; Cheng, S.; Rhee, E.P.; McCabe, E.; Lewis, G.D.; Fox, C.S.; Jacques, P.F.; Fernandez, C.; et al. Metabolite profiles and the risk of developing diabetes. Nat. Med. 2011, 17, 448–453.
- Neinast, M.; Murashige, D.; Arany, Z. Branched Chain Amino Acids. Annu. Rev. Physiol. 2019, 81, 139–164.
- Estrada-Alcalde, I.; Tenorio-Guzman, M.R.; Tovar, A.R.; Salinas-Rubio, D.; Torre-Villalvazo, I.; Torres, N.; Noriega, L.G. Metabolic Fate of Branched-Chain Amino Acids During Adipogenesis, in Adipocytes From Obese Mice and C2C12 Myotubes. J. Cell Biochem. 2017, 118, 808–818.
- Lackey, D.E.; Lynch, C.J.; Olson, K.C.; Mostaedi, R.; Ali, M.; Smith, W.H.; Karpe, F.; Humphreys, S.; Bedinger, D.H.; Dunn, T.N.; et al. Regulation of adipose branched-chain amino acid catabolism enzyme expression and cross-adipose amino acid flux in human obesity. Am. J. Physiol. Endocrinol. Metab. 2013, 304, E1175–E1187.
- Salinas-Rubio, D.; Tovar, A.R.; Torre-Villalvazo, I.; Granados-Portillo, O.; Torres, N.; Pedraza-Chaverri, J.; Noriega, L.G. Interaction between leucine and palmitate catabolism in 3T3-L1 adipocytes and primary adipocytes from control and obese rats. J. Nutr. Biochem. 2018, 59, 29–36.
- She, P.; Van Horn, C.; Reid, T.; Hutson, S.M.; Cooney, R.N.; Lynch, C.J. Obesity-related elevations in plasma leucine are associated with alterations in enzymes involved in branched-chain amino acid metabolism. Am. J. Physiol. Endocrinol. Metab. 2007, 293, E1552–E1563.
- Serralde-Zuniga, A.E.; Guevara-Cruz, M.; Tovar, A.R.; Herrera-Hernandez, M.F.; Noriega, L.G.; Granados, O.; Torres, N. Omental adipose tissue gene expression, gene variants, branched-chain amino acids, and their relationship with metabolic syndrome and insulin resistance in humans. Genes Nutr. 2014, 9, 431.
- Zoncu, R.; Bar-Peled, L.; Efeyan, A.; Wang, S.; Sancak, Y.; Sabatini, D.M. mTORC1 senses lysosomal amino acids through an inside-out mechanism that requires the vacuolar H(+)-ATPase. Science 2011, 334, 678–683.
- Zoncu, R.; Efeyan, A.; Sabatini, D.M. mTOR: From growth signal integration to cancer, diabetes and ageing. Nat. Rev. Mol. Cell Biol. 2011, 12, 21–35.
- Newgard, C.B.; An, J.; Bain, J.R.; Muehlbauer, M.J.; Stevens, R.D.; Lien, L.F.; Haqq, A.M.; Shah, S.H.; Arlotto, M.; Slentz, C.A.; et al. A branched-chain amino acid-related metabolic signature that differentiates obese and lean humans and contributes to insulin resistance. Cell Metab. 2009, 9, 311–326.
- Green, C.R.; Wallace, M.; Divakaruni, A.S.; Phillips, S.A.; Murphy, A.N.; Ciaraldi, T.P.; Metallo, C.M. Branched-chain amino acid catabolism fuels adipocyte differentiation and lipogenesis. Nat. Chem. Biol. 2016, 12, 15–21.
- Halama, A.; Horsch, M.; Kastenmuller, G.; Moller, G.; Kumar, P.; Prehn, C.; Laumen, H.; Hauner, H.; Hrabe de Angelis, M.; Beckers, J.; et al. Metabolic switch during adipogenesis: From branched chain amino acid catabolism to lipid synthesis. Arch. Biochem. Biophys. 2016, 589, 93–107.
- Hong, S.; Zhou, W.; Fang, B.; Lu, W.; Loro, E.; Damle, M.; Ding, G.; Jager, J.; Zhang, S.; Zhang, Y.; et al. Dissociation of muscle insulin sensitivity from exercise endurance in mice by HDAC3 depletion. Nat. Med. 2017, 23, 223–234.
- Lerin, C.; Goldfine, A.B.; Boes, T.; Liu, M.; Kasif, S.; Dreyfuss, J.M.; De Sousa-Coelho, A.L.; Daher, G.; Manoli, I.; Sysol, J.R.; et al. Defects in muscle branched-chain amino acid oxidation contribute to impaired lipid metabolism. Mol. Metab. 2016, 5, 926–936.
- Muoio, D.M. Metabolic inflexibility: When mitochondrial indecision leads to metabolic gridlock. Cell 2014, 159, 1253–1262.
- Chandel, N.S. Amino Acid Metabolism. Cold Spring Harb. Perspect. Biol. 2021, 13, a040584.
- Oxenkrug, G.F. Genetic and hormonal regulation of tryptophan kynurenine metabolism: Implications for vascular cognitive impairment, major depressive disorder, and aging. Ann. N. Y. Acad. Sci. 2007, 1122, 35–49.
- Schwarcz, R. The kynurenine pathway of tryptophan degradation as a drug target. Curr. Opin. Pharmacol. 2004, 4, 12–17.
- Chen, T.; Zheng, X.; Ma, X.; Bao, Y.; Ni, Y.; Hu, C.; Rajani, C.; Huang, F.; Zhao, A.; Jia, W.; et al. Tryptophan Predicts the Risk for Future Type 2 Diabetes. PLoS ONE 2016, 11, e0162192.
- Breum, L.; Rasmussen, M.H.; Hilsted, J.; Fernstrom, J.D. Twenty-four-hour plasma tryptophan concentrations and ratios are below normal in obese subjects and are not normalized by substantial weight reduction. Am. J. Clin. Nutr. 2003, 77, 1112–1118.
- Muzik, O.; Burghardt, P.; Yi, Z.; Kumar, A.; Seyoum, B. Successful metformin treatment of insulin resistance is associated with down-regulation of the kynurenine pathway. Biochem. Biophys. Res. Commun. 2017, 488, 29–32.
- Oxenkrug, G.; van der Hart, M.; Summergrad, P. Elevated anthranilic acid plasma concentrations in type 1 but not type 2 diabetes mellitus. Integr. Mol. Med. 2015, 2, 365–368.
- Kotake, Y.; Ueda, T.; Mori, T.; Igaki, S.; Hattori, M. Abnormal tryptophan metabolism and experimental diabetes by xanthurenic acid (XA). Acta Vitaminol. Enzymol. 1975, 29, 236–239.
- Lin, H.V.; Efanov, A.M.; Fang, X.; Beavers, L.S.; Wang, X.; Wang, J.; Gonzalez Valcarcel, I.C.; Ma, T. GPR142 Controls Tryptophan-Induced Insulin and Incretin Hormone Secretion to Improve Glucose Metabolism. PLoS ONE 2016, 11, e0157298.
- Chouchani, E.T.; Kazak, L.; Spiegelman, B.M. New Advances in Adaptive Thermogenesis: UCP1 and Beyond. Cell Metab. 2019, 29, 27–37.
- Nicholls, D.G. The hunt for the molecular mechanism of brown fat thermogenesis. Biochimie 2017, 134, 9–18.
- Wang, Z.; Wang, Q.A.; Liu, Y.; Jiang, L. Energy metabolism in brown adipose tissue. FEBS J. 2021, 288, 3647–3662.
- Yoneshiro, T.; Wang, Q.; Tajima, K.; Matsushita, M.; Maki, H.; Igarashi, K.; Dai, Z.; White, P.J.; McGarrah, R.W.; Ilkayeva, O.R.; et al. BCAA catabolism in brown fat controls energy homeostasis through SLC25A44. Nature 2019, 572, 614–619.
