Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is an old version of this entry, which may differ significantly from the current revision.
The compositional asymmetry of biological membranes has attracted significant attention over the last decade. Harboring more differences from symmetric membranes than previously appreciated, asymmetric bilayers have proven quite challenging to study with familiar concepts and techniques, leaving many unanswered questions about the reach of the asymmetry effects. One particular area of active research is the computational investigation of composition- and number-asymmetric lipid bilayers with molecular dynamics (MD) simulations.
- membrane asymmetry
- differential stress
- interleaflet coupling
- molecular dynamics simulations
- lipid bilayers
- computational model
1. Introduction
Lipid bilayers are ubiquitous in biology. Constituting the core of cellular membranes, they have long fascinated scientists trying to uncover their multifaceted roles in biological processes. Of particular interest are membrane biophysical properties and their relation to phenomena such as protein-lipid interactions and selective solute permeability [1][2]. The effects of lipid composition—one of the major determinants of the bilayer’s structural and mechanical properties—have been extensively studied in symmetric model membranes (defined as having two leaflets with identical lipid composition and number density) that can be easily prepared and manipulated [3].
A major discovery in the early 1970s found that the plasma membrane of erythrocytes is compositionally asymmetric, with sphingomyelin and phosphocholine lipids enriched in the outer leaflet, and the aminophospholipids PS and PE largely confined to the inner leaflet [4][5]. These observations were later confirmed for the plasma membranes of many eukaryotic cells [6], which necessitated the development of novel experimental and theoretical paradigms for the investigation of membrane asymmetry [7][8][9][10]. Central questions in early studies concerned the effects of asymmetry on protein-membrane interactions and interleaflet coupling of phase behavior (Figure 1) [11][12][13][14][15][16][17].
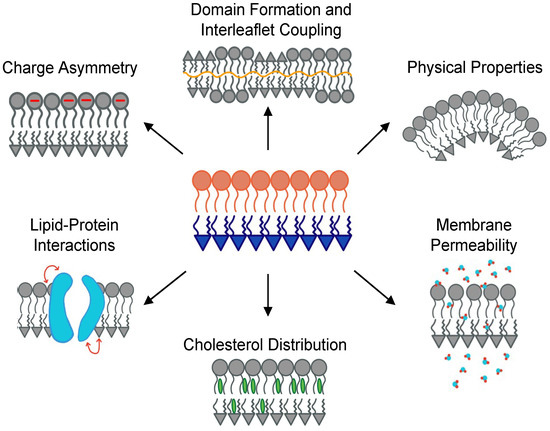
Figure 1. Various properties of asymmetric membranes studied with experiments and simulations. Clockwise from top: Effects of asymmetry on phase separation and interleaflet communication and adjustments; physical properties such as curvature, rigidity, and differential stress; permeability of water and small molecules; distribution of cholesterol between the two leaflets; interaction of membrane proteins with the bilayer; and charge asymmetry.
Offering an atomic resolution that is difficult to achieve with experiments, molecular dynamics (MD) simulations have often provided invaluable insights into the origins of membrane behavior and properties (e.g., [18][19][20][21][22][23]). Therefore, it is not surprising that the computational investigation of membrane asymmetry quickly gained speed alongside the rather challenging experimental characterization of asymmetric model membranes. Inspired mainly by observations in cells, some of the first MD simulation studies focused on the effects of asymmetry on membrane electrostatics and permeability [24][25][26][27], the structural properties of asymmetrically distributed gangliosides [27], and the peculiar confinement of raft-forming mixtures to the outer plasma membrane leaflet [28]. Later, advances in related experimental techniques [7] stimulated the investigation of interleaflet coupling [10][29][30][31][32][33][34][35][36][37][38], cholesterol interleaflet distribution [36][39][40][41][42][43], and protein interactions with asymmetric bilayers [44][45][46][47][48][49][50] (Figure 1).
In an MD simulation, a molecular model is first constructed, and its dynamics are simulated by evaluating inter-atomic forces from sets of parameters (often experimentally calibrated) and propagating the system in time following Newton’s laws of motion. Freely available software packages have been developed to assist with both steps, making MD simulations and their analysis readily accessible [51][52][53][54][55][56][57][58][59][60]. Although an asymmetric bilayer with well-defined leaflet lipid compositions can be built in minutes and generally simulated without major hurdles, it has become clear that one decision in the initial bilayer design—namely, the relative numbers of slowly-flipping lipids to place in the two leaflets—can be more consequential even than the choice of the specific lipid compositions of the leaflets [10].
2. Protocols for Bilayer Construction
To date, there are four main approaches for the construction of compositionally asymmetric bilayers for MD simulations:
-
Ensure equal numbers of lipids in the two leaflets (EqN).
-
Match the surface areas (or lipid packing densities) of the two leaflets to those from cognate symmetric bilayers (SA).
-
Eliminate differential stress, i.e., ensure zero leaflet tension (0-DS).
-
Emulate biological asymmetry (EmBioAsym).
Most of these methods require performing at least one simulation to arrive at the desired asymmetric membrane model. Once the relative leaflet compositions and abundances are determined, the bilayer can be simulated with any conventional software to investigate its properties in detail. Below we briefly describe each of these approaches and discuss a few studies in which they have been applied.
2.1. Same Number of Lipids (EqN)
Since in symmetric bilayer simulations, the two leaflets generally have equal numbers of lipids (Figure 2A), one approach to building asymmetric bilayers is to ensure that same condition (Figure 2B). This can be accomplished by building a symmetric bilayer and replacing individual lipids with different ones, or by specifying the same total numbers of lipids in the two leaflets and letting software generate the asymmetric lipid compositions. Here, all lipids are treated identically, thus disregarding any structural differences or packing preferences between species. This approach has been used to study the physical properties of both simpler lipid mixtures [27][30] and asymmetric plasma membrane models of increasing complexity [61][62].
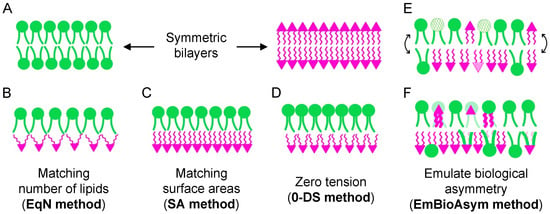
Figure 2. Protocols for the construction of asymmetric bilayers for MD simulations. (A) Symmetric bilayers have the same number and type of lipids in their two leaflets. An asymmetric bilayer can be built (B) with equal numbers of lipids in the two leaflets (EqN method) or (C) by ensuring that the relative packing densities (or surface areas) of the two leaflets are initially the same as in cognate symmetric bilayers (the SA method). Alternatively, (D) iterative simulations and analysis can be used to identify the leaflet number asymmetry that eliminates differential stress (the 0-DS method) or (E,F) advanced simulations can be performed to optimize the leaflet lipid compositions. The latter is achieved by letting a subpopulation of the lipids equilibrate their leaflet concentrations by either (E) freely exchanging between leaflets or (F) changing their identity, in the presence of constraints keeping other lipids in place (e.g., lipids with patterned headgroups in (E) can diffuse only within their respective leaflets).
2.2. Match Surface Areas (SA)
An alternative to the EqN method is choosing the relative numbers of lipids in the two leaflets so that at the initial stage of bilayer construction, their average areas per lipid (or surface areas) match those of cognate symmetric bilayers (Figure 2C). This can be accomplished by first simulating two symmetric bilayers with the lipid compositions of the asymmetric membrane leaflets, followed by either stitching their leaflets together or using the obtained equilibrium areas per lipid to calculate the respective numbers of lipids and building the asymmetric bilayer from scratch. The latter approach is more general and allows for the construction of asymmetric bilayers of different sizes. This method ensures that the relative packing densities of the two leaflets remain fixed even as the two individual leaflet areas may simultaneously increase or decrease in comparison to their symmetric counterparts. This protocol has been applied to the analysis of the membrane potential [24][25][26], interleaflet coupling [29][32][34], the permeability of plasma membrane models [63], and cholesterol interleaflet distribution [39][42].
To avoid simulating two symmetric bilayers, one can alternatively use reported areas per lipid (APL) for the individual lipids and, if the leaflets contain more than one component, assume ideal mixing and calculate the respective mole-fraction-weighted packing densities. For instance, CHARMM-GUI [54] uses individual lipid APLs to estimate the number of lipids in each leaflet when only lipid composition and lateral box dimensions are provided. This approach was recently used in a study comparing different construction methods for asymmetric bilayers [8].
2.3. Eliminate Differential Stress (0-DS)
Recent experiments have found that preferred lipid packing densities in symmetric bilayers can be altered in asymmetric bilayers, presumably due to the effects of the interleaflet coupling [64]. A corollary is that lipid areas in asymmetric bilayers cannot be estimated a priori from their values in symmetric bilayers. It follows that the EqN and SA methods described above may produce bilayers with differentially stressed leaflets [65][66]. To ensure zero leaflet tension (Figure 2C), one can follow the protocol outlined in [65], which is based on the following principle. An asymmetric bilayer is first constructed (for example, using the EqN or SA methods) and simulated until the APL is converged over the last ~200 ns. Then, the lateral pressure profile is calculated from the converged portion and used to obtain the leaflet tension. If the tension is non-zero, the number of lipids is adjusted by either removing lipids from the leaflet with negative tension or adding lipids to the leaflet with positive tension. The exact number of lipids to add or remove can be chosen on a trial-and-error basis or estimated from the relationship between the bilayer area compressibility modulus and leaflet tension (Equation 3 in [65]). A new asymmetric bilayer is then built from scratch with the updated leaflet lipid numbers and the same steps are repeated (i.e., simulation, calculation of leaflet tension, adjustment, and rebuilding if necessary) until zero leaflet tension is reached. This approach has been used to examine the effects of asymmetry on the mechanical properties of anionic asymmetric bilayers [67] and the ability of gramicidin to scramble lipids [46], as well as the effects of differential stress on the interaction of small molecules with asymmetric membranes [66].
2.4. Emulate Biological Asymmetry (EmBioAsym)
The asymmetry in cell plasma membranes (PM) is maintained via the activity of flippase and floppase enzymes which, when active, move lipids between leaflets against their concentration gradients [68]. Since the lipid specificity of these enzymes is arguably restricted to certain lipid types [69], one hypothesis is that cells regulate their PM lipid organization by restricting the asymmetry of some lipids while letting others equilibrate between leaflets according to their chemical potential. One notable example is cholesterol, a major component of mammalian cell plasma membranes that can rapidly flip between leaflets [6][70][71][72]. Interestingly, while cholesterol is capable of alleviating stresses in the membrane [73], its strong preference for interaction with saturated lipids may dominate over elastic and entropic forces and drive its distribution in a way that increases the differential stress [10][36][74]. This illustrates both the natural tendency of a bilayer constituent to equilibrate its distribution based on its chemical potential and the fact that realizing this tendency may produce rather than eliminate stresses in the membrane. In that respect, two methods have emerged to examine the equilibration of lipid redistribution in a simulated bilayer in the presence of imposed asymmetry [8][9].
The first approach involves simulations in the NPT ensemble and utilizes P21 boundary conditions (Figure 2E) [8]. These PBCs allow lipids to sample both leaflet environments by freely exchanging between leaflets during the simulation [75]. To mimic the activity of flippases and simulate asymmetric membranes, the method involves constraining some lipids to stay in one leaflet while allowing others to equilibrate their leaflet concentrations via an interleaflet redistribution [8]. Thus, it is possible to start with a bilayer constructed with one of the methods described above, then restrict some lipids to their respective leaflets and simulate the system with P21 PBCs to examine the preferred lipid distribution of the unconstrained membrane components in the presence of the imposed asymmetry. Consequently, since the relative numbers of lipids in the two leaflets are not constrained, they can dynamically change during the simulation. As noted by the authors, while opening transient pores in the membrane can also accelerate the exchange of lipids between leaflets, the advantage of P21 PBCs is that the chemical equilibrium reached by the freely diffusing lipids is a property of the asymmetric leaflets in the absence of any mechanical perturbations such as those imposed by a pore [8].
The second approach starts with a compositionally symmetric bilayer and replaces some of the lipids with new ones to generate the initial asymmetry (if the bilayer contains the same lipid numbers across leaflets, this is equivalent to the EqN method). It then proceeds with a simulation not in the NPT ensemble (as discussed above), but instead in a semi-grand canonical ensemble that allows dynamic changes in lipid identity (specifically their saturation or headgroup type) during the simulation (Figure 2F) [9]. This approach emulates the action of lipid-translocating enzymes by imposing a chemical potential difference between some molecules in one leaflet while letting the leaflet lipid compositions adjust in accordance with the chemical potential of all lipid species subject to the imposed constraints. In these simulations, the lipid number asymmetry is fixed, but the changes in the degree of lipid saturation or type across species dynamically alter the relative packing densities of the leaflets by virtue of their changing lipid compositions. This method is well suited for investigating how some asymmetries might naturally arise from others and has thus helped explain certain experimental observations of the leaflet lipid compositions in erythrocyte membranes [9].
This entry is adapted from the peer-reviewed paper 10.3390/membranes13070629
References
- Levental, K.R.; Malmberg, E.; Symons, J.L.; Fan, Y.Y.; Chapkin, R.S.; Ernst, R.; Levental, I. Lipidomic and biophysical homeostasis of mammalian membranes counteracts dietary lipid perturbations to maintain cellular fitness. Nat. Commun. 2020, 11, 1339.
- Epand, R.M.; Ruysschaert, J.M. The Biophysics of Cell Membranes; Springer: Berlin/Heidelberg, Germany, 2018.
- Marsh, D. Handbook of Lipid Bilayers, 2nd ed.; CRC Press: Boca Raton, FL, USA, 2013.
- Verkleij, A.J.; Zwaal, R.F.; Roelofsen, B.; Comfurius, P.; Kastelijn, D.; van Deenen, L.L. The asymmetric distribution of phospholipids in the human red cell membrane. A combined study using phospholipases and freeze-etch electron microscopy. Biochim. Biophys. Acta 1973, 323, 178–193.
- Bretscher, M.S. Asymmetrical lipid bilayer structure for biological membranes. Nat. New Biol. 1972, 236, 11–12.
- Doktorova, M.; Symons, J.L.; Levental, I. Structural and functional consequences of reversible lipid asymmetry in living membranes. Nat. Chem. Biol. 2020, 16, 1321–1330.
- Scott, H.L.; Kennison, K.B.; Enoki, T.A.; Doktorova, M.; Kinnun, J.J.; Heberle, F.A.; Katsaras, J. Model Membrane Systems Used to Study Plasma Membrane Lipid Asymmetry. Symmetry 2021, 13, 1356.
- Park, S.; Im, W.; Pastor, R.W. Developing initial conditions for simulations of asymmetric membranes: A practical recommendation. Biophys. J. 2021, 120, 5041–5059.
- Girard, M.; Bereau, T. Induced asymmetries in membranes. Biophys. J. 2023, 122, 2092–2098.
- Hossein, A.; Deserno, M. Spontaneous Curvature, Differential Stress, and Bending Modulus of Asymmetric Lipid Membranes. Biophys. J. 2020, 118, 624–642.
- Chiantia, S.; London, E. Acyl chain length and saturation modulate interleaflet coupling in asymmetric bilayers: Effects on dynamics and structural order. Biophys. J. 2012, 103, 2311–2319.
- Cheng, H.T.; Megha; London, E. Preparation and properties of asymmetric vesicles that mimic cell membranes: Effect upon lipid raft formation and transmembrane helix orientation. J. Biol. Chem. 2009, 284, 6079–6092.
- May, S. Trans-monolayer coupling of fluid domains in lipid bilayers. Soft Matter 2009, 5, 3148–3156.
- Garg, S.; Ruhe, J.; Ludtke, K.; Jordan, R.; Naumann, C.A. Domain registration in raft-mimicking lipid mixtures studied using polymer-tethered lipid bilayers. Biophys. J. 2007, 92, 1263–1270.
- Kiessling, V.; Crane, J.M.; Tamm, L.K. Transbilayer effects of raft-like lipid domains in asymmetric planar bilayers measured by single molecule tracking. Biophys. J. 2006, 91, 3313–3326.
- Collins, M.D.; Keller, S.L. Tuning lipid mixtures to induce or suppress domain formation across leaflets of unsupported asymmetric bilayers. Proc. Natl. Acad. Sci. USA 2008, 105, 124–128.
- Wagner, A.J.; Loew, S.; May, S. Influence of monolayer-monolayer coupling on the phase behavior of a fluid lipid bilayer. Biophys. J. 2007, 93, 4268–4277.
- Moradi, S.; Nowroozi, A.; Shahlaei, M. Shedding light on the structural properties of lipid bilayers using molecular dynamics simulation: A review study. RSC Adv. 2019, 9, 4644–4658.
- Ballweg, S.; Sezgin, E.; Doktorova, M.; Covino, R.; Reinhard, J.; Wunnicke, D.; Hanelt, I.; Levental, I.; Hummer, G.; Ernst, R. Regulation of lipid saturation without sensing membrane fluidity. Nat. Commun. 2020, 11, 756.
- Doktorova, M.; Kucerka, N.; Kinnun, J.J.; Pan, J.; Marquardt, D.; Scott, H.L.; Venable, R.M.; Pastor, R.W.; Wassall, S.R.; Katsaras, J.; et al. Molecular Structure of Sphingomyelin in Fluid Phase Bilayers Determined by the Joint Analysis of Small-Angle Neutron and X-ray Scattering Data. J. Phys. Chem. B 2020, 124, 5186–5200.
- Heberle, F.A.; Doktorova, M.; Scott, H.L.; Skinkle, A.D.; Waxham, M.N.; Levental, I. Direct label-free imaging of nanodomains in biomimetic and biological membranes by cryogenic electron microscopy. Proc. Natl. Acad. Sci. USA 2020, 117, 19943–19952.
- Doktorova, M.; LeVine, M.V.; Khelashvili, G.; Weinstein, H. A new computational method for membrane compressibility: Bilayer mechanical thickness revisited. Biophys. J. 2019, 116, 487–502.
- Doktorova, M.; Heberle, F.A.; Kingston, R.L.; Khelashvili, G.; Cuendet, M.A.; Wen, Y.; Katsaras, J.; Feigenson, G.W.; Vogt, V.M.; Dick, R.A. Cholesterol Promotes Protein Binding by Affecting Membrane Electrostatics and Solvation Properties. Biophys. J. 2017, 113, 2004–2015.
- Gurtovenko, A.A.; Vattulainen, I. Membrane potential and electrostatics of phospholipid bilayers with asymmetric transmembrane distribution of anionic lipids. J. Phys. Chem. B 2008, 112, 4629–4634.
- Gurtovenko, A.A.; Vattulainen, I. Lipid transmembrane asymmetry and intrinsic membrane potential: Two sides of the same coin. J. Am. Chem. Soc. 2007, 129, 5358–5359.
- Gurtovenko, A.A.; Vattulainen, I. Calculation of the electrostatic potential of lipid bilayers from molecular dynamics simulations: Methodological issues. J. Chem. Phys. 2009, 130, 215107.
- Patel, R.Y.; Balaji, P.V. Characterization of symmetric and asymmetric lipid bilayers composed of varying concentrations of ganglioside GM1 and DPPC. J. Phys. Chem. B 2008, 112, 3346–3356.
- Bhide, S.Y.; Zhang, Z.; Berkowitz, M.L. Molecular dynamics simulations of SOPS and sphingomyelin bilayers containing cholesterol. Biophys. J. 2007, 92, 1284–1295.
- Perlmutter, J.D.; Sachs, J.N. Interleaflet interaction and asymmetry in phase separated lipid bilayers: Molecular dynamics simulations. J. Am. Chem. Soc. 2011, 133, 6563–6577.
- Polley, A.; Vemparala, S.; Rao, M. Atomistic simulations of a multicomponent asymmetric lipid bilayer. J. Phys. Chem. B 2012, 116, 13403–13410.
- Tian, J.H.; Nickels, J.; Katsaras, J.; Cheng, X.L. Behavior of Bilayer Leaflets in Asymmetric Model Membranes: Atomistic Simulation Studies. J. Phys. Chem. B 2016, 120, 8438–8448.
- Rog, T.; Orlowski, A.; Llorente, A.; Skotland, T.; Sylvanne, T.; Kauhanen, D.; Ekroos, K.; Sandvig, K.; Vattulainen, I. Interdigitation of long-chain sphingomyelin induces coupling of membrane leaflets in a cholesterol dependent manner. Biochim. Biophys. Acta 2016, 1858, 281–288.
- Mohideen, N.; Weiner, M.D.; Feigenson, G.W. Bilayer compositional asymmetry influences the nanoscopic to macroscopic phase domain size transition. Chem. Phys. Lipids 2020, 232, 104972.
- Weiner, M.D.; Feigenson, G.W. Molecular Dynamics Simulations Reveal Leaflet Coupling in Compositionally Asymmetric Phase-Separated Lipid Membranes. J. Phys. Chem. B 2019, 123, 3968–3975.
- Blumer, M.; Harris, S.; Li, M.; Martinez, L.; Untereiner, M.; Saeta, P.N.; Carpenter, T.S.; Ingolfsson, H.I.; Bennett, W.F.D. Simulations of Asymmetric Membranes Illustrate Cooperative Leaflet Coupling and Lipid Adaptability. Front. Cell Dev. Biol. 2020, 8, 575.
- Varma, M.; Deserno, M. Distribution of cholesterol in asymmetric membranes driven by composition and differential stress. Biophys. J. 2022, 121, 4001–4018.
- Hossein, A.; Deserno, M. Stiffening transition in asymmetric lipid bilayers: The role of highly ordered domains and the effect of temperature and size. J. Chem. Phys. 2021, 154, 014704.
- Foley, S.; Deserno, M. Stabilizing Leaflet Asymmetry under Differential Stress in a Highly Coarse-Grained Lipid Membrane Model. J. Chem. Theory Comput. 2020, 16, 7195–7206.
- Yesylevskyy, S.O.; Demchenko, A.P. How cholesterol is distributed between monolayers in asymmetric lipid membranes. Eur. Biophys. J. 2012, 41, 1043–1054.
- Yesylevskyy, S.O.; Demchenko, A.P.; Kraszewski, S.; Ramseyer, C. Cholesterol induces uneven curvature of asymmetric lipid bilayers. Sci. World J. 2013, 2013, 965230.
- Karlsen, M.L.; Bruhn, D.S.; Pezeshkian, W.; Khandelia, H. Long chain sphingomyelin depletes cholesterol from the cytoplasmic leaflet in asymmetric lipid membranes. RSC Adv. 2021, 11, 22677–22682.
- Courtney, K.C.; Pezeshkian, W.; Raghupathy, R.; Zhang, C.; Darbyson, A.; Ipsen, J.H.; Ford, D.A.; Khandelia, H.; Presley, J.F.; Zha, X. C24 Sphingolipids Govern the Transbilayer Asymmetry of Cholesterol and Lateral Organization of Model and Live-Cell Plasma Membranes. Cell Rep. 2018, 24, 1037–1049.
- Aghaaminiha, M.; Farnoud, A.M.; Sharma, S. Quantitative relationship between cholesterol distribution and ordering of lipids in asymmetric lipid bilayers. Soft Matter 2021, 17, 2742–2752.
- Wu, E.L.; Fleming, P.J.; Yeom, M.S.; Widmalm, G.; Klauda, J.B.; Fleming, K.G.; Im, W.E. coli outer membrane and interactions with OmpLA. Biophys. J. 2014, 106, 2493–2502.
- Liu, H.; Yang, L.; Zhang, Q.; Mao, L.; Jiang, H.; Yang, H. Probing the structure and dynamics of caveolin-1 in a caveolae-mimicking asymmetric lipid bilayer model. Eur. Biophys. J. 2016, 45, 511–521.
- Doktorova, M.; Heberle, F.A.; Marquardt, D.; Rusinova, R.; Sanford, R.L.; Peyear, T.A.; Katsaras, J.; Feigenson, G.W.; Weinstein, H.; Andersen, O.S. Gramicidin Increases Lipid Flip-Flop in Symmetric and Asymmetric Lipid Vesicles. Biophys. J. 2019, 116, 860–873.
- Piggot, T.J.; Holdbrook, D.A.; Khalid, S. Conformational dynamics and membrane interactions of the E. coli outer membrane protein FecA: A molecular dynamics simulation study. Biochim. Biophys. Acta 2013, 1828, 284–293.
- Sharma, S.; Kim, B.N.; Stansfeld, P.J.; Sansom, M.S.P.; Lindau, M. A Coarse Grained Model for a Lipid Membrane with Physiological Composition and Leaflet Asymmetry. PLoS ONE 2015, 10, e0144814.
- Park, S.; Beaven, A.H.; Klauda, J.B.; Im, W. How Tolerant are Membrane Simulations with Mismatch in Area per Lipid between Leaflets? J. Chem. Theory Comput. 2015, 11, 3466–3477.
- Araya, M.K.; Gorfe, A.A. Phosphatidylserine and Phosphatidylethanolamine Asymmetry Have a Negligible Effect on the Global Structure, Dynamics, and Interactions of the KRAS Lipid Anchor. J. Phys. Chem. B 2022, 126, 4491–4500.
- Lee, J.; Patel, D.S.; Stahle, J.; Park, S.J.; Kern, N.R.; Kim, S.; Lee, J.; Cheng, X.; Valvano, M.A.; Holst, O.; et al. CHARMM-GUI Membrane Builder for Complex Biological Membrane Simulations with Glycolipids and Lipoglycans. J. Chem. Theory Comput. 2019, 15, 775–786.
- Wu, E.L.; Cheng, X.; Jo, S.; Rui, H.; Song, K.C.; Davila-Contreras, E.M.; Qi, Y.; Lee, J.; Monje-Galvan, V.; Venable, R.M.; et al. CHARMM-GUI Membrane Builder toward realistic biological membrane simulations. J. Comput. Chem. 2014, 35, 1997–2004.
- Jo, S.; Lim, J.B.; Klauda, J.B.; Im, W. CHARMM-GUI Membrane Builder for mixed bilayers and its application to yeast membranes. Biophys. J. 2009, 97, 50–58.
- Jo, S.; Kim, T.; Iyer, V.G.; Im, W. CHARMM-GUI: A web-based graphical user interface for CHARMM. J. Comput. Chem. 2008, 29, 1859–1865.
- Phillips, J.C.; Hardy, D.J.; Maia, J.D.C.; Stone, J.E.; Ribeiro, J.V.; Bernardi, R.C.; Buch, R.; Fiorin, G.; Henin, J.; Jiang, W.; et al. Scalable molecular dynamics on CPU and GPU architectures with NAMD. J. Chem. Phys. 2020, 153, 044130.
- Phillips, J.C.; Braun, R.; Wang, W.; Gumbart, J.; Tajkhorshid, E.; Villa, E.; Chipot, C.; Skeel, R.D.; Kale, L.; Schulten, K. Scalable molecular dynamics with NAMD. J. Comput. Chem. 2005, 26, 1781–1802.
- Abraham, M.J.; Murtola, T.; Schulz, R.; Páll, S.; Smith, J.C.; Hess, B.; Lindahl, E. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 2015, 1, 19–25.
- Pall, S.; Zhmurov, A.; Bauer, P.; Abraham, M.; Lundborg, M.; Gray, A.; Hess, B.; Lindahl, E. Heterogeneous parallelization and acceleration of molecular dynamics simulations in GROMACS. J. Chem. Phys. 2020, 153, 134110.
- Martinez, L.; Andrade, R.; Birgin, E.G.; Martinez, J.M. PACKMOL: A package for building initial configurations for molecular dynamics simulations. J. Comput. Chem. 2009, 30, 2157–2164.
- Eastman, P.; Swails, J.; Chodera, J.D.; McGibbon, R.T.; Zhao, Y.; Beauchamp, K.A.; Wang, L.P.; Simmonett, A.C.; Harrigan, M.P.; Stern, C.D.; et al. OpenMM 7: Rapid development of high performance algorithms for molecular dynamics. PLoS Comput. Biol. 2017, 13, e1005659.
- Koldso, H.; Shorthouse, D.; Helie, J.; Sansom, M.S.P. Lipid Clustering Correlates with Membrane Curvature as Revealed by Molecular Simulations of Complex Lipid Bilayers. PLoS Comput. Biol. 2014, 10, e1003911.
- Shahane, G.; Ding, W.; Palaiokostas, M.; Orsi, M. Physical properties of model biological lipid bilayers: Insights from all-atom molecular dynamics simulations. J. Mol. Model. 2019, 25, 76.
- Vacha, R.; Berkowitz, M.L.; Jungwirth, P. Molecular model of a cell plasma membrane with an asymmetric multicomponent composition: Water permeation and ion effects. Biophys. J. 2009, 96, 4493–4501.
- Heberle, F.A.; Marquardt, D.; Doktorova, M.; Geier, B.; Standaert, R.F.; Heftberger, P.; Kollmitzer, B.; Nickels, J.D.; Dick, R.A.; Feigenson, G.W.; et al. Subnanometer Structure of an Asymmetric Model Membrane: Interleaflet Coupling Influences Domain Properties. Langmuir 2016, 32, 5195–5200.
- Doktorova, M.; Weinstein, H. Accurate in silico modeling of asymmetric bilayers based on biophysical principles. Biophys. J. 2018, 115, 1638–1643.
- Pirhadi, E.; Vanegas, J.M.; Farin, M.; Schertzer, J.W.; Yong, X. Effect of Local Stress on Accurate Modeling of Bacterial Outer Membranes Using All-Atom Molecular Dynamics. J. Chem. Theory Comput. 2023, 19, 363–372.
- Jiang, W.; Lin, Y.C.; Luo, Y.L. Mechanical properties of anionic asymmetric bilayers from atomistic simulations. J. Chem. Phys. 2021, 154, 224701.
- Sakuragi, T.; Nagata, S. Regulation of phospholipid distribution in the lipid bilayer by flippases and scramblases. Nat. Rev. Mol. Cell Biol. 2023, 1–21.
- Montigny, C.; Lyons, J.; Champeil, P.; Nissen, P.; Lenoir, G. On the molecular mechanism of flippase- and scramblase-mediated phospholipid transport. Biochim. Biophys. Acta 2016, 1861 Pt B, 767–783.
- Ayuyan, A.G.; Cohen, F.S. The Chemical Potential of Plasma Membrane Cholesterol: Implications for Cell Biology. Biophys. J. 2018, 114, 904–918.
- Steck, T.L.; Lange, Y. Transverse distribution of plasma membrane bilayer cholesterol: Picking sides. Traffic 2018, 19, 750–760.
- Shaw, T.R.; Wisser, K.C.; Schaffner, T.A.; Gaffney, A.D.; Machta, B.B.; Veatch, S.L. Chemical potential measurements constrain models of cholesterol-phosphatidylcholine interactions. Biophys. J. 2023, 122, 1105–1117.
- Miettinen, M.S.; Lipowsky, R. Bilayer Membranes with Frequent Flip-Flops Have Tensionless Leaflets. Nano Lett. 2019, 19, 5011–5016.
- Doktorova, M.; Levental, I. Cholesterol’s balancing act: Defying the status quo. Biophys. J. 2022, 121, 3771–3773.
- Dolan, E.A.; Venable, R.M.; Pastor, R.W.; Brooks, B.R. Simulations of membranes and other interfacial systems using P2(1) and Pc periodic boundary conditions. Biophys. J. 2002, 82, 2317–2325.
This entry is offline, you can click here to edit this entry!
