Recent publications on gall formation induced on the leaves of dicotyledonous flowering plants by eriophyoid mites (Eriophyoidea) and representatives of four insect orders (Diptera, Hemiptera, Hymenoptera, Lepidoptera) are analyzed. Cellular and molecular level data on the stimuli that induce and sustain the development of both mite and insect galls, the expression of host plant genes during gallogenesis, and the effects of these galling arthropods on photosynthesis are considered. A hypothesis is proposed for the relationship between the size of galls and the volume of secretions injected by a parasite. Multistep, varying patterns of plant gene expression and accompanying histo-morphological changes in the transformed gall tissues are apparent. The main obstacle to better elucidating the nature of the induction of gallogenesis is the impossibility of collecting a sufficient amount of saliva for analysis, which is especially important in the case of microscopic eriophyoids. The use of modern omics technologies at the organismal level has revealed a spectrum of genetic mechanisms of gall formation at the molecular level but has not yet answered the questions regarding the nature of gall-inducing agents and the features of events occurring in plant cells at the very beginning of gall growth.
- :eriophyoid mites
- tenuipalpids
- galling arthropods
- gene expression
1. Introduction
2. Functional Diversity of Leaf Galls Induced by Arthropods
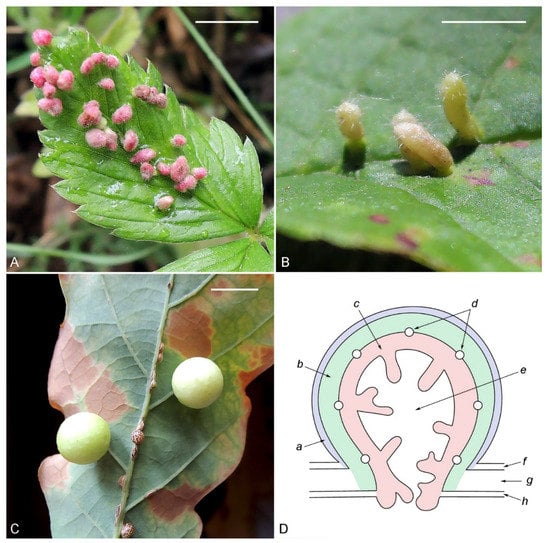
3. Stimuli That Induce the Development of Galls
4. Gall Formation Is a Multistep Process
5. Expression of Plant Genes during Gall Formation
The eriophyoid mite Fragariocoptes setiger induces galls on the leaves of the green strawberry, Fragaria viridis (Weston, 1771) (Rosaceae). During the initiation and growth of young galls, the increase in the expression of the CYCD3 and CYCB1 cell cycle genes in their tissues is associated with active cell proliferation [15]. By the time of gall maturation, a sharp decrease in the expression of cell cycle genes was found. A similar dynamic of changes in gene expression during mite galling on strawberry leaves was found for the homeobox genes, KNOX and WOX. These two genes are the universal regulators of normal plant development [61]. Finally, during the development of galls on strawberry leaves, there was an abrupt change in the expression pattern of the genes responsible for the adaxial–abaxial polarity of the leaf.
The development of leaf galls induced by several wasp species of the family Cynipidae (Amphibolips michoacaensis Nieves-Aldrey et Maldonado, 2012, B. pallida, Dryocosmus kuriphilus Yasumatsu, 1951) when they deposit eggs in the meristematic tissues of the leaves of oaks (genus Quercus) and some other trees of the beech family (Fagaceae) has been studied [16,55,56,57]. These studies revealed major differences in gene expression between gall cells and the cells of normal (control) leaves, as well as changes in the expression pattern of hundreds or possibly thousands of plant genes during gall development.
Particularly noteworthy is a study in which gene expression was analyzed during the development of both participants in a “parasite–host dialogue” involving larval B. pallida and the leaves of the common oak, Q. robur. The study demonstrated that “gall development involves expression of oak and gall wasp genes in repeatable, growth stage-specific patterns” [16]: (p. 19). In particular, in the tissues of young galls, the enhanced expression of ENOD genes occurred. These genes had been discovered in nitrogen-fixing nodules of a legume family (Fabaceae) and were later found in many other plants [62]. Nodulin-like proteins encoded by these genes belong to the large family of arabinogalactan proteins, which are glycoproteins involved in plant growth and development processes, including somatic embryogenesis [63,64,65].
In the young larvae of B. pallida, the expression of PCWDE genes, which code for plant cell wall degrading enzymes, including six pectin/pectate lyases, four cellulases and four rhamnogalacturonan lyases, occurs [16]. Enzymes encoded by these genes disrupt the wall structure of plant cells in this wasp’s feeding area. Then, numerous secreted peptides, including wasp chitinases, move into the gall tissues surrounding the larva, although it is not yet clear which larval tissue produces chitinases. The same authors inferred that their data supported a hypothesis, although it is not yet generally accepted, that galls induced by wasps of the family Cynipidae can be considered “modified somatic embryos”, with their development being similar to the somatic embryogenesis of plants. They stated that “host arabinogalactan proteins and gall wasp chitinases interact in young galls to generate a somatic embryogenesis-like process in oak tissues surrounding the gall wasp larvae” [16] (p. 1). During somatic embryogenesis, a fully developed fertile plant organism develops from a single somatic cell [67,68]; the molecular aspects of this process have been intensively studied [69,70,71].
A very recent study [53,54] revealed for the first time the tissue-specific gene expression in the active growth phase of young galls induced by the wasp Dryocosmus quercuspalustris (Osten-Sacken, 1861) on the leaves of the red oak, Quercus rubra (L., 1753). The analysis was carried out during a single stage of gall formation, specifically a young actively growing gall with a feeding larva in the internal gall chamber (approximately 5–6 days after oviposition). For the first time, not only were significant differences (28%) revealed between the transcriptomes of the whole gall and the adjacent leaf tissue, but also between the outer gall tissue, which performs a predominantly protective function, and the internal tissue of the gall, on which the parasite feeds. In general, the transcriptome of the outer tissue of the gall was more similar to the transcriptomes of the tissues of leaf buds, twigs and reproductive structures of oak than to the transcriptome of normal leaf tissue.
Leaf galling is also induced by some hemipterans. In the case of a parasite–plant host pair, the aphid, Schlechtendalia chinensis (Bell, 1851) (Aphididae), and sumac, Rhus javanica (L., 1753) (Anacardiaceae), in the early stages of gall formation, increased expression of the KNOX genes occurs [58].
An attempt to identify the genes involved in leaf gallogenesis involving the simultaneous use of several parasite–host pairs was recently undertaken by Takeda et al. [3,72]. Transcriptomes from galls formed by three pairs were studied: Rhopalomyia yomogicola (Matsumura, 1931) (Diptera, Cecidomyiidae)—Artemisia montana (Pampanini, 1930) (Asteraceae), Caloptilia cecidophora (Lepidoptera, Gracillariidae)—Glochidion obovatum (Phyllanthaceae) and Borboryctis euryae (Kumata et Kuroko, 1988) (Lepidoptera, Gracillariidae)—Eurya japonica (Thunberg, 1783) (Pentaphylacaceae). Their molecular data were discussed in combination with the molecular data obtained by the same authors for the pair Schlechtendalia chinensis—Rhus javanica [58].
6. Gallogenesis and Photosynthesis
This entry is adapted from the peer-reviewed paper 10.3390/life13061347
