SARS-CoV2 has caused the current pandemic of new coronavirus disease 2019 (COVID-19) worldwide. Clinical outcomes of COVID-19 illness range broadly from asymptotic and mild to a life-threatening situation. This casts uncertainties for defining host determinants underlying the disease severity. Recent genetic analyses based on extensive clinical sample cohorts using genome-wide association studies (GWAS) and high throughput sequencing curation revealed genetic errors and gene loci associated with about 20% of life-threatening COVID-19 cases. Significantly, most of these critical genetic loci are enriched in two immune signaling pathways, i.e., interferon-mediated antiviral signaling and chemokine-mediated/inflammatory signaling. In line with these genetic profiling studies, the broad spectrum of COVID-19 illness could be explained by immuno-pathological regulation of these critical immunogenetic pathways through various epigenetic mechanisms, which further interconnect to other vital components such as those in the renin–angiotensin–aldosterone system (RAAS) because of its direct interaction with the virus causing COVID-19. Together, key genes unraveled by genetic profiling may provide targets for precisely early risk diagnosis and prophylactic design to relieve severe COVID-19. The confounding epigenetic mechanisms may be key to understanding the clinical broadness of COVID-19 illness.
- COVID-19
- interferon signaling
- chemokine signaling, genome-wide association
- epigenetic regulation
1. The Broad Spectrum and Critical Illness in COVID-19 Progression
The coronavirus disease 2019 (COVID-19), which has been declared a worldwide pandemic by the WHO since March of 2020, is caused by the novel coronavirus severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) [1,2,3,4]. The virus evolves at a highly contagious rate in human beings, with a basic reproduction number (R0) ranging at 1.4–5.7. The clinical outcome of COVID-19 varies broadly among infected people, ranging from asymptotic infection and common cold-like sickness to a severe pneumonia leading to acute respiratory distress syndrome (ARS) and multi-organ complications that potentially have fatal prognosis [5,6,7,8]. Complications of severe COVID-19 include vasculitis, coagulopathy, thrombosis, septic shock, and even multi-organ failure [5,6,7,8]. The epidemiology of COVID-19 shows a diverse pattern across people who are different in age, sex, ethnicity, and particularly among those with pre-existing medical conditions [6,7,8,9,10,11]. For example, the US statistics showed that older patients (aged ≥65 years) accounted for 31% of all cases, 45% of hospitalizations, 53% admissions of intensive care unit (ICU), and 80% of deaths, with the highest incidence of severe outcomes in patients aged ≥85 years [1,4,8]. Similarly, increased risk of critical and life-threatening illnesses was reported to associate with males and particularly pre-existing comorbidities, including cardiovascular, renal, liver, diabetes, and other autoimmune diseases as well as obesity condition [4,5,6,7,8,9,10,11]. In contrast, evidence indicates that children (median age 4–7 years) have a lower susceptibility and risk for critical illness. However, under the circumstance of comorbidity and genetic risks, the disparity of the risk for severe COVID-19 becomes vague concerning the factors of age, sex, and ethnicity [4,5,6,7,8,9,10,11]. With a critical viral disease like COVID-19, illness comes from both the virus infection and interacting with immune responses, especially a consequential imbalance of harmful immunopathies over proper immune responses. Upon exposure to the same virus, whereas individuals show asymptotic or mild illness plausibly mounting effective immune reactions, severe COVID-19 patients, however, may reflect dysfunctional immune reactions that further leads to pathological exacerbation accompanying uncontrolled virus spreading and immune overwhelming [9,10,11,12,13,14,15,16,17]. As the virological branch focuses on diminishing viral spreading and virulence to cause disease, deciphering the genetic and especially epigenetic associations underlie severe COVID-19 will grasp the immunogenetic theme for severity prognosis in the host, thus providing manageable targets for early risk diagnosis and development of prophylactic and therapeutic remedies to face current pandemic [18,19,20].
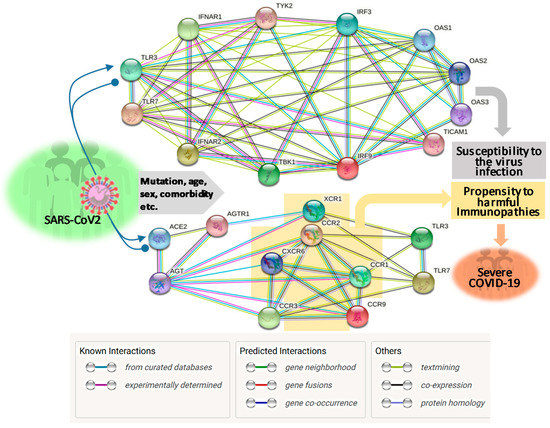
2. Genetic Association: Interferon and Chemokine Response Representing the Centric Immune Determinants Underlying Severe COVID-19
| Chr. Location (Key Genes Covered, or Epigenetic Effect) |
Association (Appr./OR: Freq.) |
Major Immune Pathway Involved |
References & Notes |
|---|---|---|---|
| 3p21.31 (SLC6A20, LZTFL1, FYCO1, CXCR6, XCR1, and CCR9; Neanderthal-originated allelic region) |
GWAS 95% CI/ (1.95–2.79: 1610 vs. 2205) [22] (2.14: 2244 vs. ~5X 2244) [23] |
ACE2 mediated amino acid transport (SLC6A20); Chemokine and Inflammation signaling, chemotaxis, immunopathies for lung injury (others) |
Associated at [22,23,24] |
| 6p22.1–33 (HLA-G, CCHCR1, NOTCH4) |
GWAS 95% CI/ (1.30–1.85: 2244 vs. ~5X 2244) |
Antigen processing and presentation (HLA); P-body component for RNA metabolism, associated with psoriasis (CCHCR1); lymphocyte development (NOTCH4) | Associated by [23] |
| 9q34.2 (ABO blood type locus) |
GWAS 95% CI/ (1.37–1.45: 1610 patient vs. 2205 control) |
Blood type-dependent pathological reaction, such as coagulation and thrombolysis | Associated by [22] |
| 12q24.13 (OAS1, OAS2, OAS3) |
GWAS 95% CI/ (1.29: 2244 vs. ~5X 2244) [23] |
IFN-mediated antiviral signaling | Associated by [23] |
| 19p13.3 (DPP9, TYK2) |
GWAS 95% CI/ (1.36–1.59: 2244 vs. ~5X 2244) [23] |
Innate antiviral defense (TYK2), and antigen presentation, CXCL10 signaling, and associated to obesity, diabetes, and cancer (DPP9) | Associated by [23] |
| 21q22.1 (IFNAR2) |
GWAS 95% CI/ (1.28: 2244 vs. ~5X 2244) [23] |
IFN-mediated immune signaling | Associated by [23] |
| Several Chr. (TLR3, UNC93B1, TICAM1, TRAF3, TBK1, IRF3/7/9, IFNAR1/2, STAT1/2) |
NGS and variant calling, wet-bench validation (3.5% of 659 severe COVID-19 vs few in 534 control) |
IFN mediated immune signaling |
Detected by [31] |
| Epigenetic obtaining (Autoantibody against IFNs, 94% in male) |
Wet-bench detection (13.7% of 987 severe COVID-19 vs. 0.33% in 1227 control) |
IFN mediated immune signaling | Detected by [39] |
| Epigenetic obtaining (Higher incidence of severe COVID-19 in aged, male, and comorbid patients) |
Inclusive studies and evidence (Higher incidence of severe COVID-19 in aged, male, and comorbid patients) |
Dysregulated IFN and chemokine responses, chronic/systemic inflammation, impaired other immune responses | Exemplified by [18,19,20,39,44,45] |
This entry is adapted from the peer-reviewed paper 10.3390/vaccines8040700
