Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is a comparison between Version 2 by Conner Chen and Version 1 by Lucia Krasničanová.
Telomere dynamics play a crucial role in the maintenance of chromosome integrity; changes in telomere length may thus contribute to the development of various diseases including cancer. Understanding the role of telomeric DNA in carcinogenesis and detecting the presence of cell-free telomeric DNA (cf-telDNA) in body fluids offer a potential biomarker for novel cancer screening and diagnostic strategies. Telomeres are protected by specialized nucleoprotein capping structures consisting of DNA and shelterin protein complexes.
- telomere length
- telomerase
- telomeric cfDNA
1. Introduction
Telomeres have been studied for decades, as they are related to important biological processes such as aging and the development of various diseases. As they play a crucial role in maintaining chromosome integrity, its deregulation and telomere length (TL) changes may lead to various age-related disorders, cardiovascular disease, or cancer development. Recent findings suggest cell-free telomeric sequences in circulation may also play regulatory roles in fine-tuning of the immune system [1]. Despite many recent advances in the study of telomeres and telomerase, numerous challenges and questions remain unexplored [2]. Understanding the role of telomeric DNA in carcinogenesis gives hope for new screening/diagnostic utility and cancer treatment strategies [3]. Several potential applications in the molecular assessment of human diseases have been suggested. However, the organization and function of telomeres in the extracellular milieu remain understudied.
Liquid biopsies involving the analysis of various biomolecules contained in the patient’s bodily fluids has become increasingly popular recently because of its undeniable benefits over conventional invasive methods [4]. This approach is gaining significant attention in clinical settings due to its potential to detect and monitor cancers at early disease stages [5]. The analysis of telomeres could enhance the existing assays, providing valuable information regarding cancer diagnosis, prognosis, and treatment response. Furthermore, it may contribute to a better comprehension of the mechanisms essential for the development and progression of various cancers.
2. Telomere Structure and Homeostasis
Telomeres are protected by specialized nucleoprotein capping structures consisting of DNA and shelterin protein complexes (Figure 1). Human telomeric DNA is composed of tandem repeats (10–15 kb at birth) of double-stranded DNA nucleotide sequence 5′-TTAGGG-3′, and a final 3′ G-rich single-stranded overhang (150–200-nucleotide-long), linked by telomere-binding proteins [6]. The 3′ G-rich single-stranded overhang folds back and invades the homologous double-stranded TTAGGG region, forming a telomeric loop (T-loop). This higher-order structure provides protection to the 3′-end by sequestering it from recognition by the DNA damage response (DDR) machinery [7]. Although the length of telomeres differs among various chromosomes and species, their sequence is similar across all eukaryotes. This would indicate that telomeres are to a great extent a long-preserved and archaic structure with a major evolutionary role in the protection of genome integrity [8].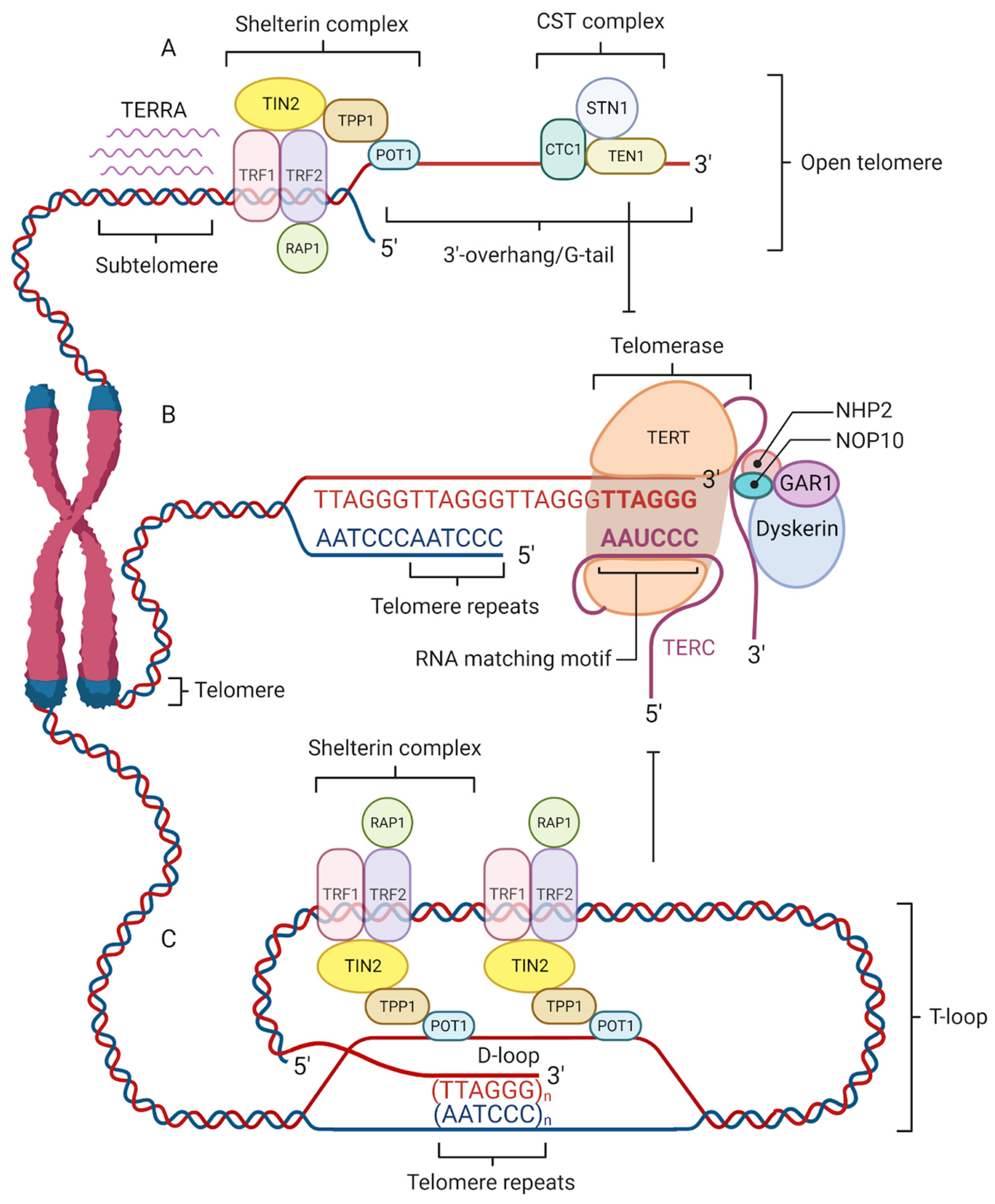
Figure 1. Four factors contribute to telomere maintenance: the shelterin complex, telomerase, telomeric repeat-containing RNA (TERRA), and the CST complex. The shelterin complex protects telomeres and regulates telomere elongation. TRF1/2, RAP1, and TIN2 are associated with double-stranded DNA; POT1 and TPP1 bind to the single-stranded G-tail DNA and are responsible for recruiting telomerase to telomeres (A). The shelterin complex coordinates the T-loop (Telomere loop) formation into which a 3′-overhang extends to form a small D-loop (Displacement-loop) and protects the end of the chromosome from damage (C). The absence of a shelterin complex causes telomere uncapping and thereby activates damage-signaling pathways. Telomerase is a reverse transcriptase enzyme that carries its own RNA molecule, which is used as a template in telomere elongation. This ribonucleoprotein complex consists of TERT (telomerase reverse transcriptase), TERC (telomerase RNA component), dyskerin, NOP10, NHP2, and GAR1 (B). TERRA is transcribed from telomere DNA and together with the shelterin complex inhibits telomere lengthening by telomerase (A). The CST complex localized on the single-stranded 3′ overhang prevents telomerase from binding to the 3′-overhang and interacts with DNA Polα-primase during telomere replication (A).
References
- Zinkova, A.; Brynychova, I.; Svacina, A.; Jirkovska, M.; Korabecna, M. Cell-Free DNA from Human Plasma and Serum Differs in Content of Telomeric Sequences and Its Ability to Promote Immune Response. Sci. Rep. 2017, 7, 2591.
- Shay, J.W.; Wright, W.E. Telomeres and Telomerase: Three Decades of Progress. Nat. Rev. Genet. 2019, 20, 299–309.
- Trybek, T.; Kowalik, A.; Góźdź, S.; Kowalska, A. Telomeres and Telomerase in Oncogenesis (Review). Oncol. Lett. 2020, 20, 1015–1027.
- Pös, O.; Biró, O.; Szemes, T.; Nagy, B. Circulating Cell-Free Nucleic Acids: Characteristics and Applications. Eur. J. Hum. Genet. 2018, 26, 937–945.
- Szilágyi, M.; Pös, O.; Márton, É.; Buglyó, G.; Soltész, B.; Keserű, J.; Penyige, A.; Szemes, T.; Nagy, B. Circulating Cell-Free Nucleic Acids: Main Characteristics and Clinical Application. Int. J. Mol. Sci. 2020, 21, 6827.
- Jafri, M.A.; Ansari, S.A.; Alqahtani, M.H.; Shay, J.W. Roles of Telomeres and Telomerase in Cancer, and Advances in Telomerase-Targeted Therapies. Genome Med. 2016, 8, 69.
- Doksani, Y.; Wu, J.Y.; de Lange, T.; Zhuang, X. Super-Resolution Fluorescence Imaging of Telomeres Reveals TRF2-Dependent T-Loop Formation. Cell 2013, 155, 345–356.
- Tommerup, H.; Dousmanis, A.; de Lange, T. Unusual Chromatin in Human Telomeres. Mol. Cell. Biol. 1994, 14, 5777–5785.
- Hemann, M.T.; Strong, M.A.; Hao, L.Y.; Greider, C.W. The Shortest Telomere, not Average Telomere Length, Is Critical for Cell Viability and Chromosome Stability. Cell 2001, 107, 67–77.
- Harley, C.B.; Futcher, A.B.; Greider, C.W. Telomeres Shorten during Ageing of Human Fibroblasts. Nature 1990, 345, 458–460.
- Reichert, S.; Stier, A. Does Oxidative Stress Shorten Telomeres in vivo? A Review. Biol. Lett. 2017, 13, 20170463.
- von Zglinicki, T. Oxidative Stress Shortens Telomeres. Trends Biochem. Sci. 2002, 27, 339–344.
- Fouquerel, E.; Lormand, J.; Bose, A.; Lee, H.-T.; Kim, G.S.; Li, J.; Sobol, R.W.; Freudenthal, B.D.; Myong, S.; Opresko, P.L. Oxidative Guanine Base Damage Regulates Human Telomerase Activity. Nat. Struct. Mol. Biol. 2016, 23, 1092–1100.
- Barnes, R.P.; Fouquerel, E.; Opresko, P.L. The Impact of Oxidative DNA Damage and Stress on Telomere Homeostasis. Mech. Ageing Dev. 2019, 177, 37–45.
- Victorelli, S.; Passos, J.F. Telomeres and Cell Senescence—Size Matters Not. EBioMedicine 2017, 21, 14–20.
- di Fagagna, F.D.; Reaper, P.M.; Clay-Farrace, L.; Fiegler, H.; Carr, P.; von Zglinicki, T.; Saretzki, G.; Carter, N.P.; Jackson, S.P. A DNA Damage Checkpoint Response in Telomere-Initiated Senescence. Nature 2003, 426, 194–198.
- Okamoto, K.; Seimiya, H. Revisiting Telomere Shortening in Cancer. Cells 2019, 8, 107.
- Blackburn, E.H.; Epel, E.S.; Lin, J. Human Telomere Biology: A Contributory and Interactive Factor in Aging, Disease Risks, and Protection. Science 2015, 350, 1193–1198.
- Muraki, K.; Nyhan, K.; Han, L.; Murnane, J.P. Mechanisms of Telomere Loss and Their Consequences for Chromosome Instability. Front. Oncol. 2012, 2, 135.
- Hanahan, D.; Weinberg, R.A. Hallmarks of Cancer: The next Generation. Cell 2011, 144, 646–674.
- de Lange, T. How Telomeres Solve the End-Protection Problem. Science 2009, 326, 948–952.
- Bekaert, S.; Derradji, H.; Baatout, S. Telomere Biology in Mammalian Germ Cells and during Development. Dev. Biol. 2004, 274, 15–30.
- Wright, W.E.; Piatyszek, M.A.; Rainey, W.E.; Byrd, W.; Shay, J.W. Telomerase Activity in Human Germline and Embryonic Tissues and Cells. Dev. Genet. 1996, 18, 173–179.
- Hiyama, E.; Hiyama, K. Telomere and Telomerase in Stem Cells. Br. J. Cancer 2007, 96, 1020–1024.
- De Vitis, M.; Berardinelli, F.; Sgura, A. Telomere Length Maintenance in Cancer: At the Crossroad between Telomerase and Alternative Lengthening of Telomeres (ALT). Int. J. Mol. Sci. 2018, 19, 606.
- Devereux, T.R.; Horikawa, I.; Anna, C.H.; Annab, L.A.; Afshari, C.A.; Barrett, J.C. DNA Methylation Analysis of the Promoter Region of the Human Telomerase Reverse Transcriptase (hTERT) Gene. Cancer Res. 1999, 59, 6087–6090.
- Shin, K.-H.; Kang, M.K.; Dicterow, E.; Park, N.-H. Hypermethylation of the hTERT Promoter Inhibits the Expression of Telomerase Activity in Normal Oral Fibroblasts and Senescent Normal Oral Keratinocytes. Br. J. Cancer 2003, 89, 1473–1478.
- Zinn, R.L.; Pruitt, K.; Eguchi, S.; Baylin, S.B.; Herman, J.G. hTERT Is Expressed in Cancer Cell Lines despite Promoter DNA Methylation by Preservation of Unmethylated DNA and Active Chromatin around the Transcription Start Site. Cancer Res. 2007, 67, 194–201.
- Barthel, F.P.; Wei, W.; Tang, M.; Martinez-Ledesma, E.; Hu, X.; Amin, S.B.; Akdemir, K.C.; Seth, S.; Song, X.; Wang, Q.; et al. Systematic Analysis of Telomere Length and Somatic Alterations in 31 Cancer Types. Nat. Genet. 2017, 49, 349–357.
- Novakovic, B.; Napier, C.E.; Vryer, R.; Dimitriadis, E.; Manuelpillai, U.; Sharkey, A.; Craig, J.M.; Reddel, R.R.; Saffery, R. DNA Methylation Mediated up-Regulation of TERRA Non-Coding RNA Is Coincident with Elongated Telomeres in the Human Placenta. Mol. Hum. Reprod. 2016, 22, 791–799.
- Slatter, T.L.; Tan, X.; Yuen, Y.C.; Gunningham, S.; Ma, S.S.; Daly, E.; Packer, S.; Devenish, C.; Royds, J.A.; Hung, N.A. The Alternative Lengthening of Telomeres Pathway May Operate in Non-Neoplastic Human Cells. J. Pathol. 2012, 226, 509–518.
- Pedram, M.; Sprung, C.N.; Gao, Q.; Lo, A.W.I.; Reynolds, G.E.; Murnane, J.P. Telomere Position Effect and Silencing of Transgenes near Telomeres in the Mouse. Mol. Cell. Biol. 2006, 26, 1865–1878.
- Robin, J.D.; Ludlow, A.T.; Batten, K.; Magdinier, F.; Stadler, G.; Wagner, K.R.; Shay, J.W.; Wright, W.E. Telomere Position Effect: Regulation of Gene Expression with Progressive Telomere Shortening over Long Distances. Genes Dev. 2014, 28, 2464–2476.
- Rippe, K.; Luke, B. TERRA and the State of the Telomere. Nat. Struct. Mol. Biol. 2015, 22, 853–858.
- Wang, C.; Zhao, L.; Lu, S. Role of TERRA in the Regulation of Telomere Length. Int. J. Biol. Sci. 2015, 11, 316–323.
More
