Fungal plant pathogens use proteinaceous effectors as well as newly identified secondary metabolites (SMs) and small non-coding RNA (sRNA) effectors to manipulate the host plant’s defense system via diverse plant cell compartments, distinct organelles, and many host genes. However, most molecular studies of plant–fungal interactions have focused on secreted effector proteins without exploring the possibly equivalent functions performed by fungal (SMs) and sRNAs, which are collectively known as “non-proteinaceous effectors”. Fungal SMs have been shown to be generated throughout the plant colonization process, particularly in the early biotrophic stages of infection. The fungal repertoire of non-proteinaceous effectors has been broadened by the discovery of fungal sRNAs that specifically target plant genes involved in resistance and defense responses. Many RNAs, particularly sRNAs involved in gene silencing, have been shown to transmit bidirectionally between fungal pathogens and their hosts.
- SMs
- sRNAs
- interactions
1. Introduction
Plants have developed a broad spectrum of responses to counter pathogen invasion. Likewise, plant pathogens orchestrate a highly calibrated array of pathogenicity strategies in their quest to cause diseases [1]. The recent increased availability of fungal and plant genomes in the public domain has facilitated considerable progress in molecular plant–fungal interaction studies. During interaction with their hosts, fungal plant pathogens secrete many proteins known as effectors which manipulate the physiology of the host or suppress the host’s immunity to promote infection [2][3]. Most studies on effectors have focused almost exclusively on secreted proteins, without exploring the possibly equivalent functions performed by fungal secondary metabolites (SMs) (chemical effectors) and sRNAs (sRNA effectors) which are collectively referred to as non-proteinaceous effectors [2][4]. Accumulating evidence has indicated that, pathogens use sRNAs (such as siRNAs and microRNAs) and SMs to manipulate host cell functions [5][6]. Fungal SMs and sRNAs have been shown to manipulate host defense-related genes in the same was as proteinaceous effectors [7][8]. In general, SM and sRNA effectors are increasingly becoming important targets for studying the pathogenesis mechanisms of fungal pathogens [9].
2. Fungal Secondary Metabolites Enhance Pathogenicity during Plant-Fungal Pathogen Interactions
Fungal SMs are not required for the growth and development of the fungus, but they have the potential to improve the pathogen’s fitness under certain conditions. Fungal SMs are often divided into polyketides, terpenes, non-ribosomal peptides and alkaloids on the basis of the primary enzymes and precursors that are involved in their biosynthesis [10][11]. The existence of fungal SMs, which have no discernible effect on the viability of the producer, raises issues about their potential influence on the environment [12]. SMs production by fungal pathogens and the presence of a host protein that is specifically susceptible to the corresponding toxin determines the ability of the pathogen to infect the host plant. Because host-specific toxin targets are encoded by plant genes, such genes can be referred to as dominant susceptibility genes [13]. Accumulating evidence indicates that fungal SMs serve as avirulence factors, host defense suppressors, and fungal cell wall hardening factors [8][14]. Fungal SMs are most effective during the early stage of infection (biotrophic phase), enhancing the fungus’ ability to penetrate and colonize its host without killing its host [2]. Fungal SMs can be host specific or non-host specific and generate necrosis in plant tissue. The majority of the fungal SMs have not been defined chemically, and the plants that they are intended to affect are still a mystery. The biological actions that have been reported to be caused by fungal SMs generated in-planta suggest that they have a broad range of plant cellular targets. The production and transport of proteins are targets of a wide range of fungal SMs [15][16].
3. Small non-coding sRNAs—The Secret agents in Plant-Fungal Pathogen Interactions
Plant immune responses are tightly regulated by an array of immunity-associated regulators such as sRNAs and some transcription factors [17]. Based on their biogenesis and structural features, sRNAs can be classified into three categories: short-interfering RNAs (siRNAs), dicer-independent microRNAs (miRNAs) and dicer-independent piwi interacting RNAs (piRNAs) [18][19]. The fundamental sRNA pathway components and other various sRNAs function as critical gene expression regulators to fine-tune the immunity of some cereal plants such as wheat and rice against pathogen invasion [17]. Normally, when a pathogen attacks its host, these sRNAs are either upregulated or downregulated in order to inhibit expression or to release suppression of their targets [5][20]. Thus, plant endogenous sRNAs and sRNA pathway components play key roles in regulating and fine-tuning host immune responses to pathogens such as fungi, bacteria, and oomycetes [21]. Accumulating evidence indicates that sRNAs produced by fungal pathogens can function as effector molecules, modulating host gene expression as a counter-defense mechanism (Table 1).| sRNA | sRNA Origin | Target Origin | Target Genes | Function | Reference |
|---|---|---|---|---|---|
| miR408 | Puccinia striiformis f. sp. tritici (Pst) | T. aestivum | CLP1 | Negatively regulates host immune response by suppressing the expression of CLP1. | [22] |
| Pst-milR1 | Pst | T. aestivum | PR2 | Represses plant innate immune response by suppressing the expression of PR2. | [7] |
| Pst-milR1 | Pst | T. aestivum | SM638 | Innate immunity. | [7] |
| pt-mil-RNA1 | Pt | T. aestivum | TCP14, CYB5R, and EF2 | Suppresses wheat defense response to Pt by targeting wheat TCP14, CYB5R and EF2. | [18] |
| pt-mil-RNA2 | Pt | T. aestivum | TCP14, CYB5R and EF2 | Suppresses wheat defense response to Pt by targeting wheat TCP14, CYB5R and EF2. | [18] |
| miR398 | Bgh | Barley | HvSOD1 | Negatively regulates host immunity by repressing HvSOD1 accumulation. | [23] |
| miR9836 | Bgh | Barley | MLA1 | Dampens immune response signaling triggered by host MLA immune receptors. | [24] |
| Fg-sRNA1 | F. graminearum | Chinese spring wheat | TaCEBiP | Suppresses wheat defense response by targeting and silencing TaCEBiP. | [25] |
| Fol-milR1 | Fusarium oxysporum | Tomato | SlyFRG4 | Suppresses host immunity by silencing SylFRG4. | [26] |
| Osa-miR167d | M. oryzae | Rice | ARF12, WRKY45 | Negatively regulates host immunity by downregulating AR12 expression. | [27] |
| miR156 | M. oryzae | Rice | SPL14 | Enhances host susceptibility by suppressing the expression of SPL14 and WRKY45. | [28] |
| Osa-miR164a | M. oryzae | Rice | OsNAC60 | Negatively regulates host immunity by suppressing OsNAC60 expression. | [29] |
| miR168 | M. oryzae | Rice | AGO1 | Negatively regulates host immunity by suppressing AGO1 expression. | [30] |
| Osa-miR169 | M. oryzae | Rice | NF-YAs | Enhances host susceptibility by suppressing the expression of nuclear factor N-Y (NF-YA) genes. | [31] |
| miR319 | M. oryzae | Rice | TCP21 | Negatively regulates host immunity by suppressing TCP21 expression. | [32] |
| miR396 | M. oryzae | Rice | OsGRFs | Negatively regulates host immunity by suppressing the expression of OsGRFs. | [33] |
| Osa-miR439 | M. oryzae | Rice | Predicted target genes LOC_Os01g23940, LOC_Os01g36270, LOC_Os01g26340 and LOC_Os06g19250 | Enhances host susceptibility by suppressing the expression of predicted target genes LOC_Os01g23940, LOC_Os01g36270, LOC_Os01g26340 and LOC_Os06g19250. | [34][35] |
| miR444b.2 | M. oryzae | Rice | MADS-box family genes | Negatively regulates host immunity by suppressing the expression of MADS-box family genes. | [36] |
| siR109944 | Rhizoctonia solani | Rice | FBL55 | Suppresses host immunity to sheath blight. | [37] |
| Bc-siR3.2 | Botrytis cinerea (B. cinerea) | A. thaliana | MPK1, MPK2 | Suppresses MPK1, MPK2 function in plant immunity. | [38] |
| Bc-siR3.1 | B. cinerea | A. thaliana | PRXIIF | Suppresses PRXIIF genes. | [38] |
| Bc-siR3.2 | B. cinerea | Solanum lycopersicum | MAPKKK4 | Suppresses MAPKKK4 function. | [38] |
| Bc-siR5 | B. cinerea | A. thaliana | WAK | Suppression the function WAK genes. | [38] |
| Bc-siR37 | B. cinerea | A. thaliana | WRKY7, PMR6 and FEI2 | Suppresses plant immunity by repressing the expression of WRKY7, PMR6 and FEI2. | [39] |
4. Cross-Kingdom RNA Interferencei during Plant-Fungal Pathogen Interactions
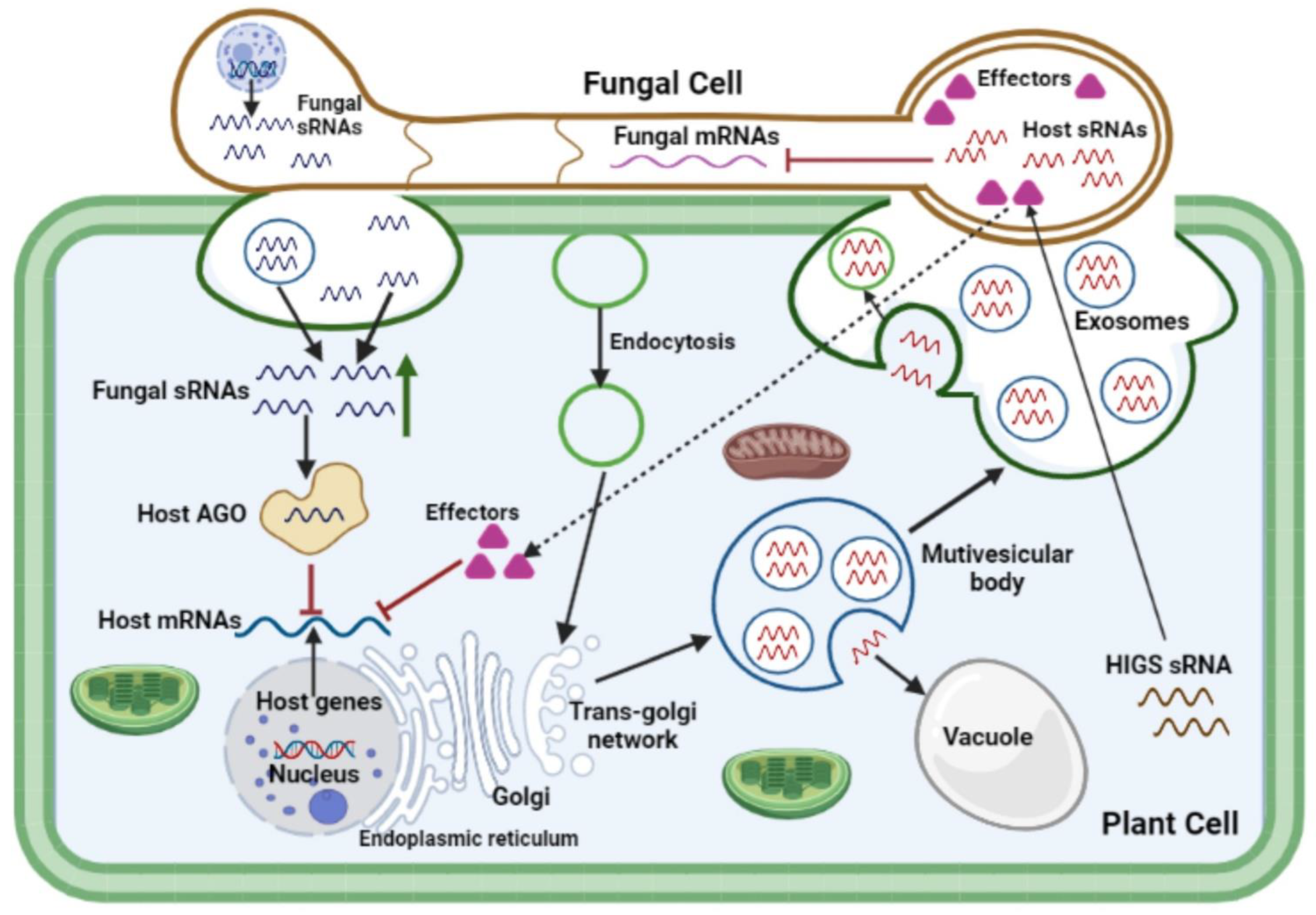
References
- Rangel, L.I.; Bolton, M.D. The unsung roles of microbial secondary metabolite effectors in the plant disease cacophony. Curr. Opin. Plant Biol. 2022, 68, 102233. https://doi.org/10.1016/j.pbi.2022.102233.
- Jaswal, R.; Kiran, K.; Rajarammohan, S.; Dubey, H.; Singh, P.K.; Sharma, Y.; Deshmukh, R.; Sonah, H.; Gupta, N.; Sharma, T. Effector Biology of Biotrophic Plant Fungal Pathogens: Current Advances and Future Prospects. Microbiol. Res. 2020, 241, 126567. https://doi.org/10.1016/j.micres.2020.126567.
- Mapuranga, J.; Zhang, N.; Zhang, L.; Chang, J.; Yang, W. Infection Strategies and Pathogenicity of Biotrophic Plant Fungal Pathogens. Front. Microbiol. 2022, 13, 799396. https://doi.org/10.3389/fmicb.2022.799396.
- Varden, F.A.; De la Concepcion, J.C.; Maidment, J.H.; Banfield, M.J. Taking the stage: Effectors in the spotlight. Curr. Opin. Plant Biol. 2017, 38, 25–33. https://doi.org/10.1016/j.pbi.2017.04.013.
- Weiberg, A.; Jin, H. Small RNAs—The secret agents in the plant-pathogen interactions. Curr. Opin. Plant Biol. 2015, 26, 87–94. https://doi.org/10.1016/j.pbi.2015.05.033.
- Mueth, N.A.; Ramachandran, S.R.; Hulbert, S.H. Small RNAs from the wheat stripe rust fungus (Puccinia striiformis f.sp. tritici). BMC Genom. 2015, 16, 1–16. https://doi.org/10.1186/s12864-015-1895-4.
- Wang, B.; Sun, Y.; Song, N.; Zhao, M.; Liu, R.; Feng, H.; Wang, X.; Kang, Z. Puccinia striiformis f. sp. tritici mi croRNA ‐like RNA 1 (Pst ‐milR1), an important pathogenicity factor of Pst, impairs wheat resistance to Pst by suppressing the wheat pathogenesis‐related 2 gene. N. Phytol. 2017, 215, 338–350. https://doi.org/10.1111/nph.14577
- Pusztahelyi, T.; Holb, I.J.; Pã³Csi, I. Secondary metabolites in fungus-plant interactions. Front. Plant Sci. 2015, 6, 573. https://doi.org/10.3389/fpls.2015.00573.
- Castro-Moretti, F.R.; Gentzel, I.N.; Mackey, D.; Alonso, A.P. Metabolomics as an Emerging Tool for the Study of Plant–Pathogen Interactions. Metabolites 2020, 10, 52. https://doi.org/10.3390/metabo10020052.
- Keller, N.P.; Turner, G.; Bennett, J.W. Fungal secondary metabolism—From biochemistry to genomics. Nat. Rev. Genet. 2005, 3, 937–947. https://doi.org/10.1038/nrmicro1286.
- Brakhage, A.A. Regulation of fungal secondary metabolism. Nat. Rev. Genet. 2012, 11, 21–32. https://doi.org/10.1038/nrmicro2916.
- Rangel, L.I.; Hamilton, O.; Jonge, R.; Bolton, M.D. Fungal social influencers: Secondary metabolites as a platform for shaping the plant‐associated community. Plant J. 2021, 108, 632–645. https://doi.org/10.1111/tpj.15490.
- Collemare, J.; Billard, A.; Böhnert, H.; Lebrun, M.-H. Biosynthesis of secondary metabolites in the rice blast fungus Magnaporthe grisea: The role of hybrid PKS-NRPS in pathogenicity. Mycol. Res. 2008, 112, 207–215. https://doi.org/10.1016/j.mycres.2007.08.003.
- Lo Presti, L.; Lanver, D.; Schweizer, G.; Tanaka, S.; Liang, L.; Tollot, M.; Zuccaro, A.; Reissmann, S.; Kahmann, R. Fungal Effectors and Plant Susceptibility. Annu. Rev. Plant Biol. 2015, 66, 513–545. https://doi.org/10.1146/annurev-arplant-043014-114623.
- Kwon, C.; Bednarek, P.; Schulze-Lefert, P. Secretory Pathways in Plant Immune Responses. Plant Physiol. 2008, 147, 1575–1583. https://doi.org/10.1104/pp.108.121566.
- Nielsen, M.E.; Feechan, A.; Böhlenius, H.; Ueda, T.; Thordal-Christensen, H. Arabidopsis ARF-GTP exchange factor, GNOM, mediates transport required for innate immunity and focal accumulation of syntaxin PEN1. Proc. Natl. Acad. Sci. USA 2012, 109, 11443–11448. https://doi.org/10.1073/pnas.1117596109.
- Mapuranga, J.; Zhang, N.; Zhang, L.; Liu, W.; Chang, J.; Yang, W. Harnessing genetic resistance to rusts in wheat and inte-grated rust management methods to develop more durable resistant cultivars. Front. Plant Sci. 2022, 13, 951095. https://doi.org/10.3389/fpls.2022.951095.
- Dubey, H.; Kiran, K.; Jaswal, R.; Jain, P.; Kayastha, A.M.; Bhardwaj, S.C.; Mondal, T.K.; Sharma, T.R. Discovery and profiling of small RNAs from Puccinia triticina by deep sequencing and identification of their potential targets in wheat. Funct. Integr. Genom. 2019, 19, 391–407. https://doi.org/10.1007/s10142-018-00652-1.
- Chapman, E.J.; Carrington, J. Specialization and evolution of endogenous small RNA pathways. Nat. Rev. Genet. 2007, 8, 884–896. https://doi.org/10.1038/nrg2179.
- Ruiz-Ferrer, V.; Voinnet, O. Roles of Plant Small RNAs in Biotic Stress Responses. Annu. Rev. Plant Biol. 2009, 60, 485–510. https://doi.org/10.1146/annurev.arplant.043008.092111.
- Weiberg, A.; Wang, M.; Bellinger, M.; Jin, H. Small RNAs: A New Paradigm in Plant-Microbe Interactions. Annu. Rev. Phyto-pathol. 2014, 52, 495–516. https://doi.org/10.1146/annurev-phyto-102313-045933.
- Feng, H.; Zhang, Q.; Wang, Q.; Wang, X.; Liu, J.; Li, M.; Huang, L.; Kang, Z. Target of tae-miR408, a chemocyanin-like protein gene (TaCLP1), plays positive roles in wheat response to high-salinity, heavy cupric stress and stripe rust. Plant Mol. Biol. 2013, 83, 433–443. https://doi.org/10.1007/s11103-013-0101-9.
- Xu, W.; Meng, Y.; Wise, R.P. Mla‐ and Rom1 ‐mediated control of microRNA398 and chloroplast copper/zinc superoxide dismutase regulates cell death in response to the barley powdery mildew fungus. N. Phytol. 2013, 201, 1396–1412. https://doi.org/10.1111/nph.12598.
- Liu, J.; Cheng, X.; Liu, D.; Xu, W.; Wise, R.; Shen, Q.-H. The miR9863 Family Regulates Distinct Mla Alleles in Barley to At-tenuate NLR Receptor-Triggered Disease Resistance and Cell-Death Signaling. PLOS Genet. 2014, 10, e1004755. https://doi.org/10.1371/journal.pgen.1004755.
- Jian, J.; Liang, X. One Small RNA of Fusarium graminearum Targets and Silences CEBiP Gene in Common Wheat. Microor-ganisms 2019, 7, 425. https://doi.org/10.3390/microorganisms7100425.
- Ji, H.; Mao, H.; Li, S.; Feng, T.; Zhang, Z.; Cheng, L.; Luo, S.; Borkovich, K.A.; Ouyang, S. Fol ‐milR1, a pathogenicity factor of Fusarium oxysporum, confers tomato wilt disease resistance by impairing host immune responses. N. Phytol. 2021, 232, 705–718. https://doi.org/10.1111/nph.17436.
- Zhao, Z.; Feng, Q.; Cao, X.; Zhu, Y.; Wang, H.; Chandran, V.; Fan, J.; Zhao, J.; Pu, M.; Li, Y.; et al. Osa‐miR167d facilitates infection of Magnaporthe oryzae in rice. J. Integr. Plant Biol. 2019, 62, 702–715. https://doi.org/10.1111/jipb.12816.
- Zhang, L.-L.; Li, Y.; Zheng, Y.-P.; Wang, H.; Yang, X.; Chen, J.-F.; Zhou, S.-X.; Wang, L.-F.; Li, X.-P.; Ma, X.-C.; et al. Expressing a Target Mimic of miR156fhl-3p Enhances Rice Blast Disease Resistance without Yield Penalty by Improving SPL14 Expression. Front. Genet. 2020, 11, 327. https://doi.org/10.3389/fgene.2020.00327.
- Wang, Z.; Xia, Y.; Lin, S.; Wang, Y.; Guo, B.; Song, X.; Ding, S.; Zheng, L.; Feng, R.; Chen, S.; et al. Osa-miR164a targetsOs-NAC60and negatively regulates rice immunity against the blast fungus Magnaporthe oryzae. Plant J. 2018, 95, 584–597. https://doi.org/10.1111/tpj.13972.
- Wang, H.; Li, Y.; Chern, M.; Zhu, Y.; Zhang, L.-L.; Lu, J.-H.; Li, X.-P.; Dang, W.-Q.; Ma, X.-C.; Yang, Z.-R.; et al. Suppression of rice miR168 improves yield, flowering time and immunity. Nat. Plants 2021, 7, 129–136. https://doi.org/10.1038/s41477-021-00852-x.
- Li, Y.; Zhao, S.-L.; Li, J.-L.; Hu, X.-H.; Wang, H.; Cao, X.-L.; Xu, Y.-J.; Zhao, Z.-X.; Xiao, Z.-Y.; Yang, N.; et al. Osa-miR169 Negatively Regulates Rice Immunity against the Blast Fungus Magnaporthe oryzae. Front. Plant Sci. 2017, 8, 2. https://doi.org/10.3389/fpls.2017.00002.
- Zhang, X.; Bao, Y.; Shan, D.; Wang, Z.; Song, X.; Wang, Z.; Wang, J.; He, L.; Wu, L.; Zhang, Z.; et al. Magnaporthe oryzae Induces the Expression of a MicroRNA to Suppress the Immune Response in Rice. Plant Physiol. 2018, 177, 352–368. https://doi.org/10.1104/pp.17.01665.
- Chandran, V.; Wang, H.; Gao, F.; Cao, X.-L.; Chen, Y.-P.; Li, G.-B.; Zhu, Y.; Yang, X.-M.; Zhang, L.-L.; Zhao, Z.; et al. miR396-OsGRFs Module Balances Growth and Rice Blast Disease-Resistance. Front. Plant Sci. 2019, 9, 1999. https://doi.org/10.3389/fpls.2018.01999.
- Wu, L.; Zhang, Q.; Zhou, H.; Ni, F.; Wu, X.; Qi, Y. Rice MicroRNA Effector Complexes and Targets. Plant Cell 2009, 21, 3421–3435. https://doi.org/10.1105/tpc.109.070938.
- Junhua, L.; Xuemei, Y.; Jinfeng, C.; Tingting, L.; Zijin, H.; Ying, X.; Jinlu, L.; Jiqun, Z.; Mei, P.; Hui, F.; et al. Osa-miR439 Negatively Regulates Rice Immunity Against Magnaporthe oryzae. Rice Sci. 2021, 28, 156–165. https://doi.org/10.1016/j.rsci.2021.01.005.
- Xiao, Z.Y.; Wang, Q.X.; Zhao, S.L.; Wang, H.; Li, J.L.; Fan, J.; Li, Y.; Wang, W.M. MiR444b. 2 regulates resistance to Mag-naporthe oryzae and tillering in rice. Acta Phytopathol. Sin. 2017, 47, 511–522.
- Qiao, L.; Zheng, L.; Sheng, C.; Zhao, H.; Jin, H.; Niu, D. Rice siR109944 suppresses plant immunity to sheath blight and impacts multiple agronomic traits by affecting auxin homeostasis. Plant J. 2020, 102, 948–964. https://doi.org/10.1111/tpj.14677.
- Weiberg, A.; Wang, M.; Lin, F.-M.; Zhao, H.; Zhang, Z.; Kaloshian, I.; Huang, H.-D.; Jin, H. Fungal Small RNAs Suppress Plant Immunity by Hijacking Host RNA Interference Pathways. Science 2013, 342, 118–123. https://doi.org/10.1126/science.1239705.
- Wang, M.; Weiberg, A.; Dellota, E., Jr.; Yamane, D.; Jin, H. Botrytis small RNA Bc-siR37 suppresses plant defense genes by cross-kingdom RNAi. RNA Biol. 2017, 14, 421–428. https://doi.org/10.1080/15476286.2017.1291112.
- Katiyar-Agarwal, S.; Jin, H. Role of Small RNAs in Host-Microbe Interactions. Annu. Rev. Phytopathol. 2010, 48, 225–246. https://doi.org/10.1146/annurev-phyto-073009-114457.
- Huang, C.-Y.; Wang, H.; Hu, P.; Hamby, R.; Jin, H. Small RNAs–Big Players in Plant-Microbe Interactions. Cell Host Microbe 2019, 26, 173–182. https://doi.org/10.1016/j.chom.2019.07.021.
- Navarro, L.; Dunoyer, P.; Jay, F.; Arnold, B.; Dharmasiri, N.; Estelle, M.; Voinnet, O.; Jones, J.D.G. A Plant miRNA Contributes to Antibacterial Resistance by Repressing Auxin Signaling. Science 2006, 312, 436–439. https://doi.org/10.1126/science.1126088.
- Li, Y.; Lu, Y.-G.; Shi, Y.; Wu, L.; Xu, Y.-J.; Huang, F.; Guo, X.-Y.; Zhang, Y.; Fan, J.; Zhao, J.-Q.; et al. Multiple Rice MicroRNAs Are Involved in Immunity against the Blast Fungus Magnaporthe oryzae. Plant Physiol. 2013, 164, 1077–1092. https://doi.org/10.1104/pp.113.230052.
- Knip, M.; Constantin, M.E.; Thordal-Christensen, H. Trans-kingdom Cross-Talk: Small RNAs on the Move. PLOS Genet. 2014, 10, e1004602. https://doi.org/10.1371/journal.pgen.1004602.
- Weiberg, A.; Bellinger, M.; Jin, H. Conversations between kingdoms: Small RNAs. Curr. Opin. Biotechnol. 2015, 32, 207–215. https://doi.org/10.1016/j.copbio.2014.12.025.
- Romano, N.; Macino, G. Quelling: Transient inactivation of gene expression in Neurospora crassa by transformation with homologous sequences. Mol. Microbiol. 1992, 6, 3343–3353. https://doi.org/10.1111/j.1365-2958.1992.tb02202.x.
- Koch, A.; Kogel, K.-H. New wind in the sails: Improving the agronomic value of crop plants through RNAi-mediated gene silencing. Plant Biotechnol. J. 2014, 12, 821–831. https://doi.org/10.1111/pbi.12226.
- Bartel, D.P. MicroRNAs: Genomics, Biogenesis, Mechanism, and Function. Cell 2004, 116, 281–297. https://doi.org/10.1016/s0092-8674(04)00045-5.
- Wang, M.; Weiberg, A.; Jin, H. Pathogen small RNAs: A new class of effectors for pathogen attacks. Mol. Plant Pathol. 2015, 16, 219–223. https://doi.org/10.1111/mpp.12233.
- Derbyshire, M.C.; Mbengue, M.; Barascud, M.; Navaud, O.; Raffaele, S. Small RNAs from the plant pathogenic fungus Scle-rotinia sclerotiorum highlight candidate host target genes associated with quantitative disease resistance. bioRxiv 2018, 354076. https://doi.org/10.1101/354076.
- Rose, L.E.; Overdijk, E.J.R.; van Damme, M. Small RNA molecules and their role in plant disease. Eur. J. Plant Pathol. 2018, 154, 115–128. https://doi.org/10.1007/s10658-018-01614-w.
- Hudzik, C.; Hou, Y.; Ma, W.; Axtell, M.J. Exchange of Small Regulatory RNAs between Plants and Their Pests. Plant Physiol. 2019, 182, 51–62. https://doi.org/10.1104/pp.19.00931.
