Fungal plant pathogens use proteinaceous effectors as well as newly identified secondary metabolites (SMs) and small non-coding RNA (sRNA) effectors to manipulate the host plant’s defense system via diverse plant cell compartments, distinct organelles, and many host genes. However, most molecular studies of plant–fungal interactions have focused on secreted effector proteins without exploring the possibly equivalent functions performed by fungal (SMs) and sRNAs, which are collectively known as “non-proteinaceous effectors”. Fungal SMs have been shown to be generated throughout the plant colonization process, particularly in the early biotrophic stages of infection. The fungal repertoire of non-proteinaceous effectors has been broadened by the discovery of fungal sRNAs that specifically target plant genes involved in resistance and defense responses. Many RNAs, particularly sRNAs involved in gene silencing, have been shown to transmit bidirectionally between fungal pathogens and their hosts.
- SMs
- sRNAs
- interactions
1. Introduction
Plants have developed a broad spectrum of responses to counter pathogen invasion. Likewise, plant pathogens orchestrate a highly calibrated array of pathogenicity strategies in their quest to cause diseases [1]. The recent increased availability of fungal and plant genomes in the public domain has facilitated considerable progress in molecular plant–fungal interaction studies. Using genetic techniques, pathogenicity or virulence factors have been established, and the study of these factors has increased our understanding of the interactions between pathogens and their hosts. During interaction with their hosts, fungal plant pathogens secrete many proteins known as effectors which manipulate the physiology of the host or suppress the host’s immunity to promote infection [2][3]. Most studies on effectors have focused almost exclusively on secreted proteins, without exploring the possibly equivalent functions performed by fungal secondary metabolites (SMs) (chemical effectors) and sRNAs (sRNA effectors) which are collectively referred to as non-proteinaceous effectors [2][4]. Accumulating evidence has indicated that, pathogens use sRNAs (such as siRNAs and microRNAs) and SMs to manipulate host cell functions [5][6][7][8][9]. Fungal SMs and sRNAs have been shown to manipulate host defense-related genes in the same was as proteinaceous effectors [7][810][11]. In general, SM and sRNA effectors are increasingly becoming important targets for studying the pathogenesis mechanisms of fungal pathogens [911]. Furthermore, these recently discovered SM and sRNA effector entities have been shown in a number of studies to be essential in manipulating host immunity and defense-related genes [2]. It is thus important to adopt new experimental methods to elucidate the in-planta biology of SM and sRNA effectors.
2. Fungal Secondary Metabolites Enhance Pathogenicity during Plant-Fungal Pathogen Interactions
Fungal SMs are not required for the growth and development of the fungus, but they have the potential to improve the pathogen’s fitness under certain conditions. Fungal SMs are often divided into polyketides, terpenes, non-ribosomal peptides and alkaloids on the basis of the primary enzymes and precursors that are involved in their biosynthesis [1012][13][1114]. They eplay a role prior to disease by shaping the plant microbial community, allowing producers to be fully adapted. The existence of fungal SMs, which have no discernible effect on the viability of the producer, raises issues about their potential influence on the environment [1215]. SMs production by fungal pathogens and the presence of a host protein that is specifically susceptible to the corresponding toxin determines the ability of the pathogen to infect the host plant. Because host-specific toxin targets are encoded by plant genes, such genes can be referred to as dominant susceptibility genes [1316]. It is generally known that mutualistic or pathogenic interactions between plants and fungal pathogens entail the simultaneous generation of molecular signals [10][17].
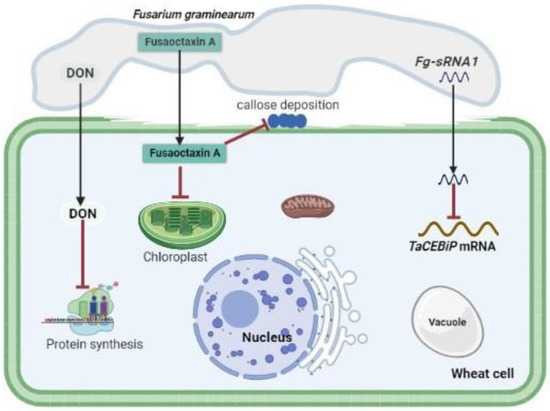
Accumulating evidence indicates that fungal SMs serve as avirulence factors, host defense suppressors, and fungal cell wall hardening factors [810][1418]. Fungal SMs are most effective during the early stage of infection (biotrophic phase), enhancing the fungus’ ability to penetrate and colonize its host without killing its host [2]. Fungal SMs can be host specific or non-host specific and generate necrosis in plant tissue. However, some fungal SMs have functions linked to virulence that are not related to necrosis [1][19][20]. As long as the host plant has the relevant molecular target, such as a resistance gene product, SMs serving as host-specific effectors are thought to play an important role in pathogen virulence [1][21]. Paradoxically, SMs acting as non-host specific effectors have been widely regarded as critical components of pathogen arsenals, despite the fact that they may not be required for pathogenesis [22]. The majority of the fungal SMs have not been defined chemically, and the plants that they are intended to affect are still a mystery. The biological actions that have been reported to be caused by fungal SMs generated in-planta suggest that they have a broad range of plant cellular targets. Some fungi use high affinity iron chelator siderophores synthesized by NRPSs to scavenge environmental iron or to sequester cellular reactive iron [23][24]. These production and transiderophores are essential for fungal growth and development, thus enhancing pathogenicity of various fungal pathogens. Cytochalasans, a diverse group of fungal PKS-NRPS hybrid metabolites, inhibit actin polymerization [25]. The production and transport of proteins are targets of a wide range of fungal SMs [26][1527]. For example, the mycotoxin deoxynivalenol (DON), a member of the type B trichothecenes, produced by Fusarium spp., inhibits protein biosynthesis by binding to the ribosome, resulting in cell signaling, differentiation, reproduction and even teratogenicity disorders in eukaryotes [28][29][30]. A comparative transcriptome analysis of symptomatic and symptomless wheat tissues revealed a substantial induction of TRI genes in symptomless tissues, indicating that DON plays an important role in modulating host defenses and infection establishment [31]. Metabolite profiling of F. graminearum wild-type and the tri5 deletion mutant in infected rachis nodes supports the function of DON in suppressing host defense-related metabolites [32]. DON was demonstrated to modulate programmed cell death (PCD) of host plant cells in a concentration-dependent way [33][34]. A higher concentration of DON may be produced by F. graminearum during infection to trigger hydrogen peroxide (H2O2) production by increasing the size of the hyphal colony. This results in further induction of PCD in wheat [33][34], and thus enhances its switch from biotrophy to necrotrophy [32]. Therefore, it can be unequivocally concluded that, during infection, the mycotoxin DON is produced as a sophisticated strategy of the fungal pathogen to circumvent and hijack the host plant’s defense system.
Cochliobolus species were reported to produce host-specific toxins that enhance pathogen virulence. Victorin, a non-ribosomal peptide produced by Cochliobolus victoriae, is a virulence factor that enhances pathogenicity by inducing PCD during infection of only oat cultivars harboring susceptible genes [35][36][37]. It was also reported that victorin targets the plasma membrane and triggers PCD signaling pathways. HC-toxin, a non-ribosomal peptide produced by Cochliobolus carbonum induces histone hyperacetylation through the inhibition of histone deacetylases, during the infection of only maize varieties harboring susceptible genes [38][39]. Transcriptional activation of host plant defense genes is altered by such histone modifications, thereby enhancing pathogen virulence [39][40]. Alternaria alternata have various pathotypes that produce different host specific toxins that are active only on their corresponding susceptible hosts [41]. Some host specific toxins including destruxin which is produced by Alternaria brassicae are also essential for pathogens in susceptible host plants. A. alternata also produces AAL-toxin, an SM which enhances pathogenicity in tomato varieties harboring susceptible genes by inhibiting ceramide synthase. This will lead to free phytosphingosine and sphinganine accumulation followed by the disruption of plasma membrane [42]. Depudecin is another SM produced by Alternaria brassicicola which also enhances pathogenicity by inhibiting histone deacetylases; its role in pathogenicity is weaker than that of HC-toxin [43]. Tenuazonic acid produced by members of the genus Alternaria and other phytopathogenic fungi inhibits protein biosynthesis on ribosomes [44][45]. It was hypothesized that the Colletotrichum graminicola disease cycle is supported by monorden and monocillins in various ways, initially promoting biotrophic asymptomatic infection by inhibiting Hsp90 chaperons of R-proteins, and disrupting a maize hypersensitive response by enabling a switch to necrotrophy through suppression of basal plant defenses [46]. Fungal SMs such as ophiobolin and herbarumin enhance virulence by inhibiting calmodulin signaling which will disrupt plant regulatory networks [47][48]. It was also demonstrated that Colletotrichum higginsianin SM higginsianin B inhibits jasmonate-mediated plant defenses [49]. Two non-ribosomal octapeptides Fusaoctaxin A and B, which are biosynthesized by the gene cluster fg3_54, were found to be F. graminearum virulence factors [50][51]. Fusaoctaxin A alters the subcellular localization of chloroplasts in coleoptile cells and inhibits callose deposition in plasmodesmata during pathogen infection, thereby facilitating F. graminearum cell-to-cell penetration in wheat cells [50][1651].
3. Small non-coding sRNAs— - The Secret agents in Plant-Fungal Pathogen Interactions
Plant immune responses are tightly regulated by an array of immunity-associated regulators such as sRNAs and some transcription factors [1752]. Based on their biogenesis and structural features, sRNAs can be classified into three categories: short-interfering RNAs (siRNAs), dicer-independent microRNAs (miRNAs) and dicer-independent piwi interacting RNAs (piRNAs) [9][53][1854][1955]. The fundamental sRNA pathway components and other various sRNAs function as critical gene expression regulators to fine-tune the immunity of some cereal plants such as wheat and rice against pathogen invasion [1752]. Normally, when a pathogen attacks its host, these sRNAs are either upregulated or downregulated in order to inhibit expression or to release suppression of their targets [5][2056]. Thus, plant endogenous sRNAs and sRNA pathway components play key roles in regulating and fine-tuning host immune responses to pathogens such as fungi, bacteria, and oomycetes [2157]. Accumulating evidence indicates that sRNAs produced by fungal pathogens can function as effector molecules, modulating host gene expression as a counter-defense mechanism (Table 1) [5][7][9][58][59][60][61].| sRNA | sRNA Origin | Target Origin | Target Genes | Function | Reference |
|---|---|---|---|---|---|
| miR408 | Puccinia striiformis f. sp. tritici (Pst) | T. aestivum | CLP1 | Negatively regulates host immune response by suppressing the expression of CLP1. | [2262] |
| Pst-milR1 | Pst | T. aestivum | PR2 | Represses plant innate immune response by suppressing the expression of PR2. | [7] |
| Pst-milR1 | Pst | T. aestivum | SM638 | Innate immunity. | [7] |
| pt-mil-RNA1 | Pt | T. aestivum | TCP14, CYB5R, and EF2 | Suppresses wheat defense response to Pt by targeting wheat TCP14, CYB5R and EF2. | [189] |
| pt-mil-RNA2 | Pt | T. aestivum | TCP14, CYB5R and EF2 | Suppresses wheat defense response to Pt by targeting wheat TCP14, CYB5R and EF2. | [189] |
| miR398 | Bgh | Barley | HvSOD1 | Negatively regulates host immunity by repressing HvSOD1 accumulation. | [2363] |
| miR9836 | Bgh | Barley | MLA1 | Dampens immune response signaling triggered by host MLA immune receptors. | [2464] |
| Fg-sRNA1 | F. graminearum | Chinese spring wheat | TaCEBiP | Suppresses wheat defense response by targeting and silencing TaCEBiP. | [2560] |
| Fol-milR1 | Fusarium oxysporum | Tomato | SlyFRG4 | Suppresses host immunity by silencing SylFRG4. | [2661] |
| Osa-miR167d | M. oryzae | Rice | ARF12, WRKY45 | Negatively regulates host immunity by downregulating AR12 expression. | [2765] |
| miR156 | M. oryzae | Rice | SPL14 | Enhances host susceptibility by suppressing the expression of SPL14 and WRKY45. | [2866] |
| Osa-miR164a | M. oryzae | Rice | OsNAC60 | Negatively regulates host immunity by suppressing OsNAC60 expression. | [2967] |
| miR168 | M. oryzae | Rice | AGO1 | Negatively regulates host immunity by suppressing AGO1 expression. | [3068] |
| Osa-miR169 | M. oryzae | Rice | NF-YAs | Enhances host susceptibility by suppressing the expression of nuclear factor N-Y (NF-YA) genes. | [3169] |
| miR319 | M. oryzae | Rice | TCP21 | Negatively regulates host immunity by suppressing TCP21 expression. | [3270] |
| miR396 | M. oryzae | Rice | OsGRFs | Negatively regulates host immunity by suppressing the expression of OsGRFs. | [3371] |
| Osa-miR439 | M. oryzae | Rice | Predicted target genes LOC_Os01g23940, LOC_Os01g36270, LOC_Os01g26340 and LOC_Os06g19250 | Enhances host susceptibility by suppressing the expression of predicted target genes LOC_Os01g23940, LOC_Os01g36270, LOC_Os01g26340 and LOC_Os06g19250. | [3472][3573] |
| miR444b.2 | M. oryzae | Rice | MADS-box family genes | Negatively regulates host immunity by suppressing the expression of MADS-box family genes. | [3674] |
| siR109944 | Rhizoctonia solani | Rice | FBL55 | Suppresses host immunity to sheath blight. | [3775] |
| Bc-siR3.2 | Botrytis cinerea (B. cinerea) | A. thaliana | MPK1, MPK2 | Suppresses MPK1, MPK2 function in plant immunity. | [3858] |
| Bc-siR3.1 | B. cinerea | A. thaliana | PRXIIF | Suppresses PRXIIF genes. | [3858] |
| Bc-siR3.2 | B. cinerea | Solanum lycopersicum | MAPKKK4 | Suppresses MAPKKK4 function. | [3858] |
| Bc-siR5 | B. cinerea | A. thaliana | WAK | Suppression the function WAK genes. | [3858] |
| Bc-siR37 | B. cinerea | A. thaliana | WRKY7, PMR6 and FEI2 | Suppresses plant immunity by repressing the expression of WRKY7, PMR6 and FEI2. | [3959] |
4. Cross-Kingdom RNAi Interference during Plant-Fungal Pathogen Interactions
5. Applications of Cross-Kingdom RNAi Technology
6. Translocation of Non-Proteinaceous Effectors across Kingdoms: Extracellular Vesicles as Mediators of Infection
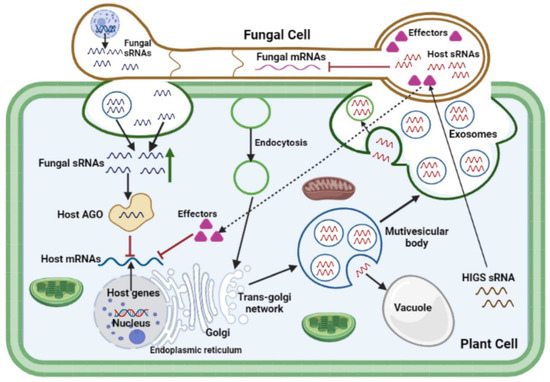
In effector biology, the mechanisms through which fungal effectors, particularly sRNA and SM effectors, are transported into cells of the host plant to their targets remain a matter of speculation. Accumulating evidence from preliminary studies suggests that genetic material may be transferred from the host plant to the infecting fungal pathogen cell through exosomal biogenesis pathways [117]. At fungal penetration sites, multivesicular compartments aggregate around fungal haustorial complexes in the host cytoplasm, allowing differentiated vesicle trafficking across the plant-pathogen cellular interface to occur anterogradely, and possibly retrogradely. These multivesicular bodies consist of several intraluminal vesicles, which are discharged extracellularly as exosomes into the paramural region after fusion with the plasma membrane [117]. Multivesicular body-like compartments were reported to be involved in trafficking processes at intercellular channels known as gap junctions, nanotubes, and even the internalization of plasma membrane sections by neighboring cells [124]. siRNA species generated in the host silencing donor were suggested to be transmitted to the fungal recipient through an exocytic/endocytic exchange process at the haustorial interface [117]. Exosomes and plasma membrane-budded microvesicles have both been identified as extracellular vesicles (EVs) that are secreted by plant cells and found in the cells of fungal pathogens [125]. EVs serve as mediators of infection and defenses during plant–fungal pathogen interactions (Figure 1). Active extracellular vesicular transport, passive transport via trans-cell wall diffusion, binding and internalization through membrane-associated receptors, and other trans-membrane pores or channels are all possible sRNA trafficking mechanisms across the plant-fungal interface [81][86][126][127][128][129][130]. A diverse collection of plant sRNAs, including miRNAs and siRNAs, are selectively loaded into the EVs of plant cells [55][129]. It was suggested that fungal sRNAs delivery is facilitated by EVs, similar to the suggested plant-extracellular vesicle-mediated sRNA transport [131]. To test this hypothesis, EVs isolated from various fungal pathogens including F. oxysporum [132][133], F. graminearum [133][134], Z. tritici [135], and Ustilago maydis [136], were established and this laid a foundation for future study of cross-kingdom RNA transport in plant-fungal pathogen interactions [137]. The secreted EVs included a variety of membrane-trafficking proteins and numerous proteins for substrate transport, indicating that EVs might serve an important role in RNA trafficking [127]. Altogether, accumulating evidence points to the idea that fungal EVs are a viable method of transporting pathogen effector proteins, sRNAs and SMs into host plant cells, and their packaging in membrane-bound compartments protects them from degradation by the host enzymes and dilution by water in the plant apoplast.
7. Tools for the Prediction and Study of SM and sRNA Effectors
In effector biology, genomes, transcriptomes, proteomes, and metabolomes are mined to facilitate the discovery of potential effector genes for molecular or cellular biology, biochemistry, and reverse genetics. Future studies need to focus on the development of integrated approaches for the molecular and functional characterization of fungal SM and sRNA effectors during interactions with their host plants. Deletion mutants are usually studied for pathogenicity or symbiosis. The most significant obstacles in the generation of deletion mutants in fungi continue to be transformation and homologous recombination. This strategy has various difficulties when used for SM and sRNA effectors; therefore, new experimental approaches are needed to overcome them. Phylogenetics and comparative genomics analyses should be performed before experimental research because they are particularly informative since the number of fungal genomes and documented SM pathways is increasing. In silico studies may provide novel insights into the organization of conserved gene clusters, as well as their limits and evolutionary history. These kinds of approaches are extremely useful in locating gene clusters that play a role in the production of SMs that have been characterized in other fungal species. They also make it possible to predict the production of compounds that are either identical or related to those produced by specific fungal species. The increased availability of fungal genome sequences and next-generation genomic technologies enables the assessment of SM gene clusters in an individual fungus. RNA-Seq has revolutionized transcriptome profiling and is utilized to study SM gene cluster expression during infection. RNA-Seq can simultaneously quantify transcripts from many organisms, making it ideal for studying plant-pathogen interactions. Manipulating strain-unique SM genes involved in host-specific pathogenicity facilitates plant-fungal pathogen interactions research. Using BGC expression in heterologous hosts such as Saccharomyces cerevisiae or Aspergillus spp. may help to overcome functional redundancy and in-planta detection limitations. SMs from plant pathogenic fungi have primarily been evaluated using phytotoxicity tests. The utilization of chemical genetic screenings may also discover actions against phytohormone signaling pathways and PTI responses. For high-throughput chemical screening, this technique may utilize A. thaliana transgenic lines that express reporter genes in 96-well microplates [138]. Patterns of gene expression and regulation may be used to decipher the complicated bidirectional interaction between pathogen and host cells [90]. Using RNA-seq on both the pathogen and the host is an effective way to examine both sides of this relationship [139][140]. The recently discovered CRISPR Cas13 system can be used to study sRNA effectors, for instance, through the inactivation or localization of fungal sRNAs [141]. Recent studies have shown that extracellular vesicles play significant roles in host defense and pathogen virulence as well as being essential tools for communication between plants and pathogens. To induce the silencing of fungal genes essential for pathogenicity, plant cells secrete extracellular vesicles containing sRNAs into fungal cells. Transmission electron microscopy following ultra-rapid cryofixation showed EVs in Golovinomyces orontii extrahaustorial matrix. EVs produced by apoplastic pathogens may be detected from plant washing solutions [142]. Such fluids likely include plant and fungal EVs, making it difficult to determine their source of origin. To find out if plant pathogen EVs carry RNAs that are functional inside host plant cells, more research must be done, including the biogenesis of EVs and how specific molecules are sorted and directed towards them.
References
- Rangel, L.I.; Bolton, M.D. The unsung roles of microbial secondary metabolite effectors in the plant disease cacophony. Curr. Opin. Plant Biol. 2022, 68, 102233. https://doi.org/10.1016/j.pbi.2022.102233.
- Jaswal, R.; Kiran, K.; Rajarammohan, S.; Dubey, H.; Singh, P.K.; Sharma, Y.; Deshmukh, R.; Sonah, H.; Gupta, N.; Sharma, T. Effector Biology of Biotrophic Plant Fungal Pathogens: Current Advances and Future Prospects. Microbiol. Res. 2020, 241, 126567. https://doi.org/10.1016/j.micres.2020.126567.
- Mapuranga, J.; Zhang, N.; Zhang, L.; Chang, J.; Yang, W. Infection Strategies and Pathogenicity of Biotrophic Plant Fungal Pathogens. Front. Microbiol. 2022, 13, 799396. https://doi.org/10.3389/fmicb.2022.799396.
- Varden, F.A.; De la Concepcion, J.C.; Maidment, J.H.; Banfield, M.J. Taking the stage: Effectors in the spotlight. Curr. Opin. Plant Biol. 2017, 38, 25–33. https://doi.org/10.1016/j.pbi.2017.04.013.
- Weiberg, A.; Jin, H. Small RNAs—The secret agents in the plant-pathogen interactions. Curr. Opin. Plant Biol. 2015, 26, 87–94. https://doi.org/10.1016/j.pbi.2015.05.033.
- Mueth, N.A.; Ramachandran, S.R.; Hulbert, S.H. Small RNAs from the wheat stripe rust fungus (Puccinia striiformis f.sp. tritici). BMC Genom. 2015, 16, 1–16. https://doi.org/10.1186/s12864-015-1895-4.
- Wang, B.; Sun, Y.; Song, N.; Zhao, M.; Liu, R.; Feng, H.; Wang, X.; Kang, Z. Puccinia striiformis f. sp. tritici mi croRNA ‐like RNA 1 (Pst ‐milR1), an important pathogenicity factor of Pst, impairs wheat resistance to Pst by suppressing the wheat pathogenesis‐related 2 gene. N. Phytol. 2017, 215, 338–350. https://doi.org/10.1111/nph.14577
- Pusztahelyi, T.; Holb, I.J.; Pã³Csi, I. Secondary metabolites in fungus-plant interactions. Front. Plant Sci. 2015, 6, 573. https://doi.org/10.3389/fpls.2015.00573.Kusch, S.; Frantzeskakis, L.; Thieron, H.; Panstruga, R. Small RNAs from cereal powdery mildew pathogens may target host plant genes. Fungal Biol. 2018, 122, 1050–1063. https://doi.org/10.1016/j.funbio.2018.08.008.
- Castro-Moretti, F.R.; Gentzel, I.N.; Mackey, D.; Alonso, A.P. Metabolomics as an Emerging Tool for the Study of Plant–Pathogen Interactions. Metabolites 2020, 10, 52. https://doi.org/10.3390/metabo10020052.Dubey, H.; Kiran, K.; Jaswal, R.; Jain, P.; Kayastha, A.M.; Bhardwaj, S.C.; Mondal, T.K.; Sharma, T.R. Discovery and profiling of small RNAs from Puccinia triticina by deep sequencing and identification of their potential targets in wheat. Funct. Integr. Genom. 2019, 19, 391–407. https://doi.org/10.1007/s10142-018-00652-1.
- Keller, N.P.; Turner, G.; Bennett, J.W. Fungal secondary metabolism—From biochemistry to genomics. Nat. Rev. Genet. 2005, 3, 937–947. https://doi.org/10.1038/nrmicro1286.Pusztahelyi, T.; Holb, I.J.; Pã³Csi, I. Secondary metabolites in fungus-plant interactions. Front. Plant Sci. 2015, 6, 573. https://doi.org/10.3389/fpls.2015.00573.
- Brakhage, A.A. Regulation of fungal secondary metabolism. Nat. Rev. Genet. 2012, 11, 21–32. https://doi.org/10.1038/nrmicro2916.Castro-Moretti, F.R.; Gentzel, I.N.; Mackey, D.; Alonso, A.P. Metabolomics as an Emerging Tool for the Study of Plant–Pathogen Interactions. Metabolites 2020, 10, 52. https://doi.org/10.3390/metabo10020052.
- Rangel, L.I.; Hamilton, O.; Jonge, R.; Bolton, M.D. Fungal social influencers: Secondary metabolites as a platform for shaping the plant‐associated community. Plant J. 2021, 108, 632–645. https://doi.org/10.1111/tpj.15490.Keller, N.P.; Turner, G.; Bennett, J.W. Fungal secondary metabolism—From biochemistry to genomics. Nat. Rev. Genet. 2005, 3, 937–947. https://doi.org/10.1038/nrmicro1286.
- Collemare, J.; Billard, A.; Böhnert, H.; Lebrun, M.-H. Biosynthesis of secondary metabolites in the rice blast fungus Magnaporthe grisea: The role of hybrid PKS-NRPS in pathogenicity. Mycol. Res. 2008, 112, 207–215. https://doi.org/10.1016/j.mycres.2007.08.003.Brakhage, A.A. Regulation of fungal secondary metabolism. Nat. Rev. Genet. 2012, 11, 21–32. https://doi.org/10.1038/nrmicro2916.
- Lo Presti, L.; Lanver, D.; Schweizer, G.; Tanaka, S.; Liang, L.; Tollot, M.; Zuccaro, A.; Reissmann, S.; Kahmann, R. Fungal Effectors and Plant Susceptibility. Annu. Rev. Plant Biol. 2015, 66, 513–545. https://doi.org/10.1146/annurev-arplant-043014-114623.Keller, N.P. Fungal secondary metabolism: Regulation, function and drug discovery. Nat. Rev. Microbiol. 2019, 17, 167–180. https://doi.org/10.1038/s41579-018-0121-1.
- Kwon, C.; Bednarek, P.; Schulze-Lefert, P. Secretory Pathways in Plant Immune Responses. Plant Physiol. 2008, 147, 1575–1583. https://doi.org/10.1104/pp.108.121566.Rangel, L.I.; Hamilton, O.; Jonge, R.; Bolton, M.D. Fungal social influencers: Secondary metabolites as a platform for shaping the plant‐associated community. Plant J. 2021, 108, 632–645. https://doi.org/10.1111/tpj.15490.
- Nielsen, M.E.; Feechan, A.; Böhlenius, H.; Ueda, T.; Thordal-Christensen, H. Arabidopsis ARF-GTP exchange factor, GNOM, mediates transport required for innate immunity and focal accumulation of syntaxin PEN1. Proc. Natl. Acad. Sci. USA 2012, 109, 11443–11448. https://doi.org/10.1073/pnas.1117596109.Collemare, J.; Billard, A.; Böhnert, H.; Lebrun, M.-H. Biosynthesis of secondary metabolites in the rice blast fungus Magnaporthe grisea: The role of hybrid PKS-NRPS in pathogenicity. Mycol. Res. 2008, 112, 207–215. https://doi.org/10.1016/j.mycres.2007.08.003.
- Mapuranga, J.; Zhang, N.; Zhang, L.; Liu, W.; Chang, J.; Yang, W. Harnessing genetic resistance to rusts in wheat and inte-grated rust management methods to develop more durable resistant cultivars. Front. Plant Sci. 2022, 13, 951095. https://doi.org/10.3389/fpls.2022.951095.Kai, K. Bioorganic chemistry of signaling molecules in microbial communication. J. Pestic. Sci. 2019, 44, 200–207. https://doi.org/10.1584/jpestics.j19-02.
- Dubey, H.; Kiran, K.; Jaswal, R.; Jain, P.; Kayastha, A.M.; Bhardwaj, S.C.; Mondal, T.K.; Sharma, T.R. Discovery and profiling of small RNAs from Puccinia triticina by deep sequencing and identification of their potential targets in wheat. Funct. Integr. Genom. 2019, 19, 391–407. https://doi.org/10.1007/s10142-018-00652-1.Lo Presti, L.; Lanver, D.; Schweizer, G.; Tanaka, S.; Liang, L.; Tollot, M.; Zuccaro, A.; Reissmann, S.; Kahmann, R. Fungal Effectors and Plant Susceptibility. Annu. Rev. Plant Biol. 2015, 66, 513–545. https://doi.org/10.1146/annurev-arplant-043014-114623.
- Chapman, E.J.; Carrington, J. Specialization and evolution of endogenous small RNA pathways. Nat. Rev. Genet. 2007, 8, 884–896. https://doi.org/10.1038/nrg2179.Collemare, J.; Lebrun, M. Fungal Secondary Metabolites: Ancient Toxins and Novel Effectors in Plant–Microbe Interactions. 2011, 377–400. https://doi.org/10.1002/9781119949138.ch15.
- Ruiz-Ferrer, V.; Voinnet, O. Roles of Plant Small RNAs in Biotic Stress Responses. Annu. Rev. Plant Biol. 2009, 60, 485–510. https://doi.org/10.1146/annurev.arplant.043008.092111.Stergiopoulos, I.; Collemare, J.; Mehrabi, R.; De Wit, P.J. Phytotoxic secondary metabolites and peptides produced by plant pathogenic Dothideomycete fungi. FEMS Microbiol. Rev. 2013, 37, 67–93. https://doi.org/10.1111/j.1574-6976.2012.00349.x.
- Weiberg, A.; Wang, M.; Bellinger, M.; Jin, H. Small RNAs: A New Paradigm in Plant-Microbe Interactions. Annu. Rev. Phyto-pathol. 2014, 52, 495–516. https://doi.org/10.1146/annurev-phyto-102313-045933.Friesen, T.L.; Faris, J.D.; Solomon, P.S.; Oliver, R.P. Host-specific toxins: Effectors of necrotrophic pathogenicity. Cell. Microbiol. 2008, 10, 1421–1428. https://doi.org/10.1111/j.1462-5822.2008.01153.x.
- Feng, H.; Zhang, Q.; Wang, Q.; Wang, X.; Liu, J.; Li, M.; Huang, L.; Kang, Z. Target of tae-miR408, a chemocyanin-like protein gene (TaCLP1), plays positive roles in wheat response to high-salinity, heavy cupric stress and stripe rust. Plant Mol. Biol. 2013, 83, 433–443. https://doi.org/10.1007/s11103-013-0101-9.Griffiths, S.; Mesarich, C.H.; Overdijk, E.J.R.; Saccomanno, B.; de Wit, P.J.G.M.; Collemare, J. Down‐regulation of cladofulvin biosynthesis is required for biotrophic growth of Cladosporium fulvum on tomato. Mol. Plant Pathol. 2017, 19, 369–380. https://doi.org/10.1111/mpp.12527.
- Xu, W.; Meng, Y.; Wise, R.P. Mla‐ and Rom1 ‐mediated control of microRNA398 and chloroplast copper/zinc superoxide dismutase regulates cell death in response to the barley powdery mildew fungus. N. Phytol. 2013, 201, 1396–1412. https://doi.org/10.1111/nph.12598.Haas, H.; Eisendle, M.; Turgeon, B.G. Siderophores in Fungal Physiology and Virulence. Annu. Rev. Phytopathol. 2008, 46, 149–187. https://doi.org/10.1146/annurev.phyto.45.062806.094338.
- Liu, J.; Cheng, X.; Liu, D.; Xu, W.; Wise, R.; Shen, Q.-H. The miR9863 Family Regulates Distinct Mla Alleles in Barley to At-tenuate NLR Receptor-Triggered Disease Resistance and Cell-Death Signaling. PLOS Genet. 2014, 10, e1004755. https://doi.org/10.1371/journal.pgen.1004755.Bushley, K.E.; Ripoll, D.R.; Turgeon, B.G. Module evolution and substrate specificity of fungal nonribosomal peptide synthetases involved in siderophore biosynthesis. BMC Evol. Biol. 2008, 8, 328. https://doi.org/10.1186/1471-2148-8-328.
- Jian, J.; Liang, X. One Small RNA of Fusarium graminearum Targets and Silences CEBiP Gene in Common Wheat. Microor-ganisms 2019, 7, 425. https://doi.org/10.3390/microorganisms7100425.Skellam, E. The biosynthesis of cytochalasans. Nat. Prod. Rep. 2017, 34, 1252–1263. https://doi.org/10.1039/c7np00036g.
- Ji, H.; Mao, H.; Li, S.; Feng, T.; Zhang, Z.; Cheng, L.; Luo, S.; Borkovich, K.A.; Ouyang, S. Fol ‐milR1, a pathogenicity factor of Fusarium oxysporum, confers tomato wilt disease resistance by impairing host immune responses. N. Phytol. 2021, 232, 705–718. https://doi.org/10.1111/nph.17436.Kwon, C.; Bednarek, P.; Schulze-Lefert, P. Secretory Pathways in Plant Immune Responses. Plant Physiol. 2008, 147, 1575–1583. https://doi.org/10.1104/pp.108.121566.
- Zhao, Z.; Feng, Q.; Cao, X.; Zhu, Y.; Wang, H.; Chandran, V.; Fan, J.; Zhao, J.; Pu, M.; Li, Y.; et al. Osa‐miR167d facilitates infection of Magnaporthe oryzae in rice. J. Integr. Plant Biol. 2019, 62, 702–715. https://doi.org/10.1111/jipb.12816.Nielsen, M.E.; Feechan, A.; Böhlenius, H.; Ueda, T.; Thordal-Christensen, H. Arabidopsis ARF-GTP exchange factor, GNOM, mediates transport required for innate immunity and focal accumulation of syntaxin PEN1. Proc. Natl. Acad. Sci. USA 2012, 109, 11443–11448. https://doi.org/10.1073/pnas.1117596109.
- Zhang, L.-L.; Li, Y.; Zheng, Y.-P.; Wang, H.; Yang, X.; Chen, J.-F.; Zhou, S.-X.; Wang, L.-F.; Li, X.-P.; Ma, X.-C.; et al. Expressing a Target Mimic of miR156fhl-3p Enhances Rice Blast Disease Resistance without Yield Penalty by Improving SPL14 Expression. Front. Genet. 2020, 11, 327. https://doi.org/10.3389/fgene.2020.00327.Pestka, J.J. Deoxynivalenol: Mechanisms of action, human exposure, and toxicological relevance. Arch. Toxicol. 2010, 84, 663–679. https://doi.org/10.1007/s00204-010-0579-8.
- Wang, Z.; Xia, Y.; Lin, S.; Wang, Y.; Guo, B.; Song, X.; Ding, S.; Zheng, L.; Feng, R.; Chen, S.; et al. Osa-miR164a targetsOs-NAC60and negatively regulates rice immunity against the blast fungus Magnaporthe oryzae. Plant J. 2018, 95, 584–597. https://doi.org/10.1111/tpj.13972.Chen, Y.; Kistler, H.C.; Ma, Z. Fusarium graminearum Trichothecene Mycotoxins: Biosynthesis, Regulation, and Management. Annu. Rev. Phytopathol. 2019, 57, 15–39. https://doi.org/10.1146/annurev-phyto-082718-100318.
- Wang, H.; Li, Y.; Chern, M.; Zhu, Y.; Zhang, L.-L.; Lu, J.-H.; Li, X.-P.; Dang, W.-Q.; Ma, X.-C.; Yang, Z.-R.; et al. Suppression of rice miR168 improves yield, flowering time and immunity. Nat. Plants 2021, 7, 129–136. https://doi.org/10.1038/s41477-021-00852-x.Xu, M.; Wang, Q.; Wang, G.; Zhang, X.; Liu, H.; Jiang, C. Combatting Fusarium head blight: Advances in molecular interactions between Fusarium graminearum and wheat. Phytopathol. Res. 2022, 4, 1–16. https://doi.org/10.1186/s42483-022-00142-0.
- Li, Y.; Zhao, S.-L.; Li, J.-L.; Hu, X.-H.; Wang, H.; Cao, X.-L.; Xu, Y.-J.; Zhao, Z.-X.; Xiao, Z.-Y.; Yang, N.; et al. Osa-miR169 Negatively Regulates Rice Immunity against the Blast Fungus Magnaporthe oryzae. Front. Plant Sci. 2017, 8, 2. https://doi.org/10.3389/fpls.2017.00002.Brown, N.A.; Evans, J.; Mead, A.; Hammond-Kosack, K.E. A spatial temporal analysis of the Fusarium graminearum tran-scriptome during symptomless and symptomatic wheat infection. Mol. Plant Pathol. 2017, 18, 1295–1312. https://doi.org/10.1111/mpp.12564.
- Zhang, X.; Bao, Y.; Shan, D.; Wang, Z.; Song, X.; Wang, Z.; Wang, J.; He, L.; Wu, L.; Zhang, Z.; et al. Magnaporthe oryzae Induces the Expression of a MicroRNA to Suppress the Immune Response in Rice. Plant Physiol. 2018, 177, 352–368. https://doi.org/10.1104/pp.17.01665.Bönnighausen, J.; Schauer, N.; Schäfer, W.; Bormann, J. Metabolic profiling of wheat rachis node infection by Fusarium gra-minearum—Decoding deoxynivalenol-dependent susceptibility. N. Phytol. 2018, 221, 459–469. https://doi.org/10.1111/nph.15377.
- Chandran, V.; Wang, H.; Gao, F.; Cao, X.-L.; Chen, Y.-P.; Li, G.-B.; Zhu, Y.; Yang, X.-M.; Zhang, L.-L.; Zhao, Z.; et al. miR396-OsGRFs Module Balances Growth and Rice Blast Disease-Resistance. Front. Plant Sci. 2019, 9, 1999. https://doi.org/10.3389/fpls.2018.01999.Desmond, O.J.; Manners, J.M.; Stephens, A.E.; Maclean, D.J.; Schenk, P.M.; Gardiner, D.M.; Munn, A.L.; Kazan, K. The Fusarium mycotoxin deoxynivalenol elicits hydrogen peroxide production, programmed cell death and defence responses in wheat. Mol. Plant Pathol. 2008, 9, 435–445. https://doi.org/10.1111/j.1364-3703.2008.00475.x.
- Wu, L.; Zhang, Q.; Zhou, H.; Ni, F.; Wu, X.; Qi, Y. Rice MicroRNA Effector Complexes and Targets. Plant Cell 2009, 21, 3421–3435. https://doi.org/10.1105/tpc.109.070938.Diamond, M.; Reape, T.J.; Rocha, O.; Doyle, S.; Kacprzyk, J.; Doohan, F.; McCabe, P.F. The Fusarium Mycotoxin Deoxyniva-lenol Can Inhibit Plant Apoptosis-Like Programmed Cell Death. PLoS ONE 2013, 8, e69542. https://doi.org/10.1371/journal.pone.0069542.
- Junhua, L.; Xuemei, Y.; Jinfeng, C.; Tingting, L.; Zijin, H.; Ying, X.; Jinlu, L.; Jiqun, Z.; Mei, P.; Hui, F.; et al. Osa-miR439 Negatively Regulates Rice Immunity Against Magnaporthe oryzae. Rice Sci. 2021, 28, 156–165. https://doi.org/10.1016/j.rsci.2021.01.005.Navarre, D.A.; Wolpert, T.J. Victorin Induction of an Apoptotic/Senescence–like Response in Oats. Plant Cell 1999, 11, 237–249. https://doi.org/10.1105/tpc.11.2.237.
- Xiao, Z.Y.; Wang, Q.X.; Zhao, S.L.; Wang, H.; Li, J.L.; Fan, J.; Li, Y.; Wang, W.M. MiR444b. 2 regulates resistance to Mag-naporthe oryzae and tillering in rice. Acta Phytopathol. Sin. 2017, 47, 511–522.Tada, Y.; Kusaka, K.; Betsuyaku, S.; Shinogi, T.; Sakamoto, M.; Ohura, Y.; Hata, S.; Mori, T.; Tosa, Y.; Mayama, S. Victorin Triggers Programmed Cell Death and the Defense Response via Interaction with a Cell Surface Mediator. Plant Cell Physiol. 2005, 46, 1787–1798. https://doi.org/10.1093/pcp/pci193.
- Qiao, L.; Zheng, L.; Sheng, C.; Zhao, H.; Jin, H.; Niu, D. Rice siR109944 suppresses plant immunity to sheath blight and impacts multiple agronomic traits by affecting auxin homeostasis. Plant J. 2020, 102, 948–964. https://doi.org/10.1111/tpj.14677.Sweat, T.A.; Lorang, J.M.; Bakker, E.G.; Wolpert, T.J. Characterization of Natural and Induced Variation in the LOV1 Gene, a CC-NB-LRR Gene Conferring Victorin Sensitivity and Disease Susceptibility in Arabidopsis. Mol. Plant-Microbe Interact. 2008, 21, 7–19. https://doi.org/10.1094/mpmi-21-1-0007.
- Weiberg, A.; Wang, M.; Lin, F.-M.; Zhao, H.; Zhang, Z.; Kaloshian, I.; Huang, H.-D.; Jin, H. Fungal Small RNAs Suppress Plant Immunity by Hijacking Host RNA Interference Pathways. Science 2013, 342, 118–123. https://doi.org/10.1126/science.1239705.Brosch, G.; Ransom, R.; Lechner, T.; Walton, J.D.; Loidl, P. Inhibition of maize histone deacetylases by HC toxin, the host-selective toxin of Cochliobolus carbonum. Plant Cell 1995, 7, 1941–1950. https://doi.org/10.1105/tpc.7.11.1941.
- Wang, M.; Weiberg, A.; Dellota, E., Jr.; Yamane, D.; Jin, H. Botrytis small RNA Bc-siR37 suppresses plant defense genes by cross-kingdom RNAi. RNA Biol. 2017, 14, 421–428. https://doi.org/10.1080/15476286.2017.1291112.Ransom, R.F.; Walton, J.D. Histone Hyperacetylation in Maize in Response to Treatment with HC-Toxin or Infection by the Filamentous Fungus Cochliobolus carbonum. Plant Physiol. 1997, 115, 1021–1027. https://doi.org/10.1104/pp.115.3.1021.
- Katiyar-Agarwal, S.; Jin, H. Role of Small RNAs in Host-Microbe Interactions. Annu. Rev. Phytopathol. 2010, 48, 225–246. https://doi.org/10.1146/annurev-phyto-073009-114457.Walton, J.D. Host-selective toxins: Agents of compatibility. Plant Cell 1996, 8, 1723–1733. https://doi.org/10.1105/tpc.8.10.1723.
- Huang, C.-Y.; Wang, H.; Hu, P.; Hamby, R.; Jin, H. Small RNAs–Big Players in Plant-Microbe Interactions. Cell Host Microbe 2019, 26, 173–182. https://doi.org/10.1016/j.chom.2019.07.021.Thomma, B.P.H.J. Alternariaspp.: From general saprophyte to specific parasite. Mol. Plant Pathol. 2003, 4, 225–236. https://doi.org/10.1046/j.1364-3703.2003.00173.x.
- Navarro, L.; Dunoyer, P.; Jay, F.; Arnold, B.; Dharmasiri, N.; Estelle, M.; Voinnet, O.; Jones, J.D.G. A Plant miRNA Contributes to Antibacterial Resistance by Repressing Auxin Signaling. Science 2006, 312, 436–439. https://doi.org/10.1126/science.1126088.Spassieva, S.D.; Markham, J.E.; Hille, J. The plant disease resistance geneAsc-1prevents disruption of sphingolipid metabolism during AAL-toxin-induced programmed cell death. Plant J. 2002, 32, 561–572. https://doi.org/10.1046/j.1365-313x.2002.01444.x.
- Li, Y.; Lu, Y.-G.; Shi, Y.; Wu, L.; Xu, Y.-J.; Huang, F.; Guo, X.-Y.; Zhang, Y.; Fan, J.; Zhao, J.-Q.; et al. Multiple Rice MicroRNAs Are Involved in Immunity against the Blast Fungus Magnaporthe oryzae. Plant Physiol. 2013, 164, 1077–1092. https://doi.org/10.1104/pp.113.230052.Wight, W.D.; Kim, K.-H.; Lawrence, C.B.; Walton, J.D. Biosynthesis and Role in Virulence of the Histone Deacetylase Inhibitor Depudecin from Alternaria brassicicola. Mol. Plant-Microbe Interact. 2009, 22, 1258–1267. https://doi.org/10.1094/mpmi-22-10-1258.
- Knip, M.; Constantin, M.E.; Thordal-Christensen, H. Trans-kingdom Cross-Talk: Small RNAs on the Move. PLOS Genet. 2014, 10, e1004602. https://doi.org/10.1371/journal.pgen.1004602.A Friedman, M.; Aggarwal, V.; E Lester, G. Inhibition of epidermal DNA synthesis by cycloheximide and other inhibitors of protein synthesis. Res. Commun. Chem. Pathol. Pharmacol. 1975, 11, 311–318.
- Weiberg, A.; Bellinger, M.; Jin, H. Conversations between kingdoms: Small RNAs. Curr. Opin. Biotechnol. 2015, 32, 207–215. https://doi.org/10.1016/j.copbio.2014.12.025.Chen, S.; Qiang, S. Recent advances in tenuazonic acid as a potential herbicide. Pestic. Biochem. Physiol. 2017, 143, 252–257. https://doi.org/10.1016/j.pestbp.2017.01.003.
- Romano, N.; Macino, G. Quelling: Transient inactivation of gene expression in Neurospora crassa by transformation with homologous sequences. Mol. Microbiol. 1992, 6, 3343–3353. https://doi.org/10.1111/j.1365-2958.1992.tb02202.x.Wicklow, D.T.; Jordan, A.M.; Gloer, J.B. Antifungal metabolites (monorden, monocillins I, II, III) from Colletotrichum grami-nicola, a systemic vascular pathogen of maize. Mycol. Res. 2009, 113, 1433–1442. https://doi.org/10.1016/j.mycres.2009.10.001.
- Koch, A.; Kogel, K.-H. New wind in the sails: Improving the agronomic value of crop plants through RNAi-mediated gene silencing. Plant Biotechnol. J. 2014, 12, 821–831. https://doi.org/10.1111/pbi.12226.Au, T.; Chick, W.S.; Leung, P. Initial kinetics of the inactivation of calmodulin by the fungal toxin ophiobolin A. Int. J. Biochem. Cell Biol. 2000, 32, 1173–1182. https://doi.org/10.1016/s1357-2725(00)00058-3.
- Bartel, D.P. MicroRNAs: Genomics, Biogenesis, Mechanism, and Function. Cell 2004, 116, 281–297. https://doi.org/10.1016/s0092-8674(04)00045-5.Rivero-Cruz, J.F.; Macías, M.; Cerda-García-Rojas, C.M.; Mata, R. A New Phytotoxic Nonenolide from Phoma herbarum. J. Nat. Prod. 2003, 66, 511–514. https://doi.org/10.1021/np020501t.
- Wang, M.; Weiberg, A.; Jin, H. Pathogen small RNAs: A new class of effectors for pathogen attacks. Mol. Plant Pathol. 2015, 16, 219–223. https://doi.org/10.1111/mpp.12233.Dallery, J.-F.; Zimmer, M.; Halder, V.; Suliman, M.; Pigné, S.; Le Goff, G.; Gianniou, D.D.; Trougakos, I.P.; Ouazzani, J.; Gasperini, D.; et al. Inhibition of jasmonate-mediated plant defences by the fungal metabolite higginsianin B. J. Exp. Bot. 2020, 71, 2910–2921. https://doi.org/10.1093/jxb/eraa061.
- Derbyshire, M.C.; Mbengue, M.; Barascud, M.; Navaud, O.; Raffaele, S. Small RNAs from the plant pathogenic fungus Scle-rotinia sclerotiorum highlight candidate host target genes associated with quantitative disease resistance. bioRxiv 2018, 354076. https://doi.org/10.1101/354076.Jia, L.-J.; Tang, H.-Y.; Wang, W.-Q.; Yuan, T.-L.; Wei, W.-Q.; Pang, B.; Gong, X.-M.; Wang, S.-F.; Li, Y.-J.; Zhang, D.; et al. A linear nonribosomal octapeptide from Fusarium graminearum facilitates cell-to-cell invasion of wheat. Nat. Commun. 2019, 10, 1–20. https://doi.org/10.1038/s41467-019-08726-9.
- Rose, L.E.; Overdijk, E.J.R.; van Damme, M. Small RNA molecules and their role in plant disease. Eur. J. Plant Pathol. 2018, 154, 115–128. https://doi.org/10.1007/s10658-018-01614-w.Tang, Z.; Tang, H.; Wang, W.; Xue, Y.; Chen, D.; Tang, W.; Liu, W. Biosynthesis of a New Fusaoctaxin Virulence Factor in Fusarium graminearum Relies on a Distinct Path to Form a Guanidinoacetyl Starter Unit Priming Nonribosomal Octapeptidyl Assembly. J. Am. Chem. Soc. 2021, 143, 19719–19730. https://doi.org/10.1021/jacs.1c07770.
- Hudzik, C.; Hou, Y.; Ma, W.; Axtell, M.J. Exchange of Small Regulatory RNAs between Plants and Their Pests. Plant Physiol. 2019, 182, 51–62. https://doi.org/10.1104/pp.19.00931.Mapuranga, J.; Zhang, N.; Zhang, L.; Liu, W.; Chang, J.; Yang, W. Harnessing genetic resistance to rusts in wheat and inte-grated rust management methods to develop more durable resistant cultivars. Front. Plant Sci. 2022, 13, 951095. https://doi.org/10.3389/fpls.2022.951095.
- Chapman, E.J.; Carrington, J. Specialization and evolution of endogenous small RNA pathways. Nat. Rev. Genet. 2007, 8, 884–896. https://doi.org/10.1038/nrg2179.
- Axtell, M.J. Classification and Comparison of Small RNAs from Plants. Annu. Rev. Plant Biol. 2013, 64, 137–159. https://doi.org/10.1146/annurev-arplant-050312-120043.
- Cai, Q.; He, B.; Wang, S.; Fletcher, S.; Niu, D.; Mitter, N.; Birch, P.R.J.; Jin, H. Message in a Bubble: Shuttling Small RNAs and Proteins Between Cells and Interacting Organisms Using Extracellular Vesicles. Annu. Rev. Plant Biol. 2021, 72, 497–524. https://doi.org/10.1146/annurev-arplant-081720-010616.
- Ruiz-Ferrer, V.; Voinnet, O. Roles of Plant Small RNAs in Biotic Stress Responses. Annu. Rev. Plant Biol. 2009, 60, 485–510. https://doi.org/10.1146/annurev.arplant.043008.092111.
- Weiberg, A.; Wang, M.; Bellinger, M.; Jin, H. Small RNAs: A New Paradigm in Plant-Microbe Interactions. Annu. Rev. Phyto-pathol. 2014, 52, 495–516. https://doi.org/10.1146/annurev-phyto-102313-045933.
- Weiberg, A.; Wang, M.; Lin, F.-M.; Zhao, H.; Zhang, Z.; Kaloshian, I.; Huang, H.-D.; Jin, H. Fungal Small RNAs Suppress Plant Immunity by Hijacking Host RNA Interference Pathways. Science 2013, 342, 118–123. https://doi.org/10.1126/science.1239705.
- Wang, M.; Weiberg, A.; Dellota, E., Jr.; Yamane, D.; Jin, H. Botrytis small RNA Bc-siR37 suppresses plant defense genes by cross-kingdom RNAi. RNA Biol. 2017, 14, 421–428. https://doi.org/10.1080/15476286.2017.1291112.
- Jian, J.; Liang, X. One Small RNA of Fusarium graminearum Targets and Silences CEBiP Gene in Common Wheat. Microor-ganisms 2019, 7, 425. https://doi.org/10.3390/microorganisms7100425.
- Ji, H.; Mao, H.; Li, S.; Feng, T.; Zhang, Z.; Cheng, L.; Luo, S.; Borkovich, K.A.; Ouyang, S. Fol ‐milR1, a pathogenicity factor of Fusarium oxysporum, confers tomato wilt disease resistance by impairing host immune responses. N. Phytol. 2021, 232, 705–718. https://doi.org/10.1111/nph.17436.
- Feng, H.; Zhang, Q.; Wang, Q.; Wang, X.; Liu, J.; Li, M.; Huang, L.; Kang, Z. Target of tae-miR408, a chemocyanin-like protein gene (TaCLP1), plays positive roles in wheat response to high-salinity, heavy cupric stress and stripe rust. Plant Mol. Biol. 2013, 83, 433–443. https://doi.org/10.1007/s11103-013-0101-9.
- Xu, W.; Meng, Y.; Wise, R.P. Mla‐ and Rom1 ‐mediated control of microRNA398 and chloroplast copper/zinc superoxide dismutase regulates cell death in response to the barley powdery mildew fungus. N. Phytol. 2013, 201, 1396–1412. https://doi.org/10.1111/nph.12598.
- Liu, J.; Cheng, X.; Liu, D.; Xu, W.; Wise, R.; Shen, Q.-H. The miR9863 Family Regulates Distinct Mla Alleles in Barley to At-tenuate NLR Receptor-Triggered Disease Resistance and Cell-Death Signaling. PLOS Genet. 2014, 10, e1004755. https://doi.org/10.1371/journal.pgen.1004755.
- Zhao, Z.; Feng, Q.; Cao, X.; Zhu, Y.; Wang, H.; Chandran, V.; Fan, J.; Zhao, J.; Pu, M.; Li, Y.; et al. Osa‐miR167d facilitates infection of Magnaporthe oryzae in rice. J. Integr. Plant Biol. 2019, 62, 702–715. https://doi.org/10.1111/jipb.12816.
- Zhang, L.-L.; Li, Y.; Zheng, Y.-P.; Wang, H.; Yang, X.; Chen, J.-F.; Zhou, S.-X.; Wang, L.-F.; Li, X.-P.; Ma, X.-C.; et al. Expressing a Target Mimic of miR156fhl-3p Enhances Rice Blast Disease Resistance without Yield Penalty by Improving SPL14 Expression. Front. Genet. 2020, 11, 327. https://doi.org/10.3389/fgene.2020.00327.
- Wang, Z.; Xia, Y.; Lin, S.; Wang, Y.; Guo, B.; Song, X.; Ding, S.; Zheng, L.; Feng, R.; Chen, S.; et al. Osa-miR164a targetsOs-NAC60and negatively regulates rice immunity against the blast fungus Magnaporthe oryzae. Plant J. 2018, 95, 584–597. https://doi.org/10.1111/tpj.13972.
- Wang, H.; Li, Y.; Chern, M.; Zhu, Y.; Zhang, L.-L.; Lu, J.-H.; Li, X.-P.; Dang, W.-Q.; Ma, X.-C.; Yang, Z.-R.; et al. Suppression of rice miR168 improves yield, flowering time and immunity. Nat. Plants 2021, 7, 129–136. https://doi.org/10.1038/s41477-021-00852-x.
- Li, Y.; Zhao, S.-L.; Li, J.-L.; Hu, X.-H.; Wang, H.; Cao, X.-L.; Xu, Y.-J.; Zhao, Z.-X.; Xiao, Z.-Y.; Yang, N.; et al. Osa-miR169 Negatively Regulates Rice Immunity against the Blast Fungus Magnaporthe oryzae. Front. Plant Sci. 2017, 8, 2. https://doi.org/10.3389/fpls.2017.00002.
- Zhang, X.; Bao, Y.; Shan, D.; Wang, Z.; Song, X.; Wang, Z.; Wang, J.; He, L.; Wu, L.; Zhang, Z.; et al. Magnaporthe oryzae Induces the Expression of a MicroRNA to Suppress the Immune Response in Rice. Plant Physiol. 2018, 177, 352–368. https://doi.org/10.1104/pp.17.01665.
- Chandran, V.; Wang, H.; Gao, F.; Cao, X.-L.; Chen, Y.-P.; Li, G.-B.; Zhu, Y.; Yang, X.-M.; Zhang, L.-L.; Zhao, Z.; et al. miR396-OsGRFs Module Balances Growth and Rice Blast Disease-Resistance. Front. Plant Sci. 2019, 9, 1999. https://doi.org/10.3389/fpls.2018.01999.
- Wu, L.; Zhang, Q.; Zhou, H.; Ni, F.; Wu, X.; Qi, Y. Rice MicroRNA Effector Complexes and Targets. Plant Cell 2009, 21, 3421–3435. https://doi.org/10.1105/tpc.109.070938.
- Junhua, L.; Xuemei, Y.; Jinfeng, C.; Tingting, L.; Zijin, H.; Ying, X.; Jinlu, L.; Jiqun, Z.; Mei, P.; Hui, F.; et al. Osa-miR439 Negatively Regulates Rice Immunity Against Magnaporthe oryzae. Rice Sci. 2021, 28, 156–165. https://doi.org/10.1016/j.rsci.2021.01.005.
- Xiao, Z.Y.; Wang, Q.X.; Zhao, S.L.; Wang, H.; Li, J.L.; Fan, J.; Li, Y.; Wang, W.M. MiR444b. 2 regulates resistance to Mag-naporthe oryzae and tillering in rice. Acta Phytopathol. Sin. 2017, 47, 511–522.
- Qiao, L.; Zheng, L.; Sheng, C.; Zhao, H.; Jin, H.; Niu, D. Rice siR109944 suppresses plant immunity to sheath blight and impacts multiple agronomic traits by affecting auxin homeostasis. Plant J. 2020, 102, 948–964. https://doi.org/10.1111/tpj.14677.
- Katiyar-Agarwal, S.; Jin, H. Role of Small RNAs in Host-Microbe Interactions. Annu. Rev. Phytopathol. 2010, 48, 225–246. https://doi.org/10.1146/annurev-phyto-073009-114457.
- Huang, C.-Y.; Wang, H.; Hu, P.; Hamby, R.; Jin, H. Small RNAs–Big Players in Plant-Microbe Interactions. Cell Host Microbe 2019, 26, 173–182. https://doi.org/10.1016/j.chom.2019.07.021.
- Schaefer, L.K.; Parlange, F.; Buchmann, G.; Jung, E.; Wehrli, A.; Herren, G.; Müller, M.C.; Stehlin, J.; Schmid, R.; Wicker, T.; et al. Cross-Kingdom RNAi of Pathogen Effectors Leads to Quantitative Adult Plant Resistance in Wheat. Front. Plant Sci. 2020, 11, 253. https://doi.org/10.3389/fpls.2020.00253.
- Navarro, L.; Dunoyer, P.; Jay, F.; Arnold, B.; Dharmasiri, N.; Estelle, M.; Voinnet, O.; Jones, J.D.G. A Plant miRNA Contributes to Antibacterial Resistance by Repressing Auxin Signaling. Science 2006, 312, 436–439. https://doi.org/10.1126/science.1126088.
- Li, Y.; Lu, Y.-G.; Shi, Y.; Wu, L.; Xu, Y.-J.; Huang, F.; Guo, X.-Y.; Zhang, Y.; Fan, J.; Zhao, J.-Q.; et al. Multiple Rice MicroRNAs Are Involved in Immunity against the Blast Fungus Magnaporthe oryzae. Plant Physiol. 2013, 164, 1077–1092. https://doi.org/10.1104/pp.113.230052.
- Knip, M.; Constantin, M.E.; Thordal-Christensen, H. Trans-kingdom Cross-Talk: Small RNAs on the Move. PLOS Genet. 2014, 10, e1004602. https://doi.org/10.1371/journal.pgen.1004602.
- Weiberg, A.; Bellinger, M.; Jin, H. Conversations between kingdoms: Small RNAs. Curr. Opin. Biotechnol. 2015, 32, 207–215. https://doi.org/10.1016/j.copbio.2014.12.025.
- Chacko, N.; Lin, X. Non-coding RNAs in the development and pathogenesis of eukaryotic microbes. Appl. Microbiol. Biotechnol. 2013, 97, 7989–7997. https://doi.org/10.1007/s00253-013-5160-y.
- Buck, A.H.; Coakley, G.; Simbari, F.; McSorley, H.J.; Quintana, J.F.; Le Bihan, T.; Kumar, S.; Abreu-Goodger, C.; Lear, M.; Harcus, Y.; et al. Exosomes secreted by nematode parasites transfer small RNAs to mammalian cells and modulate innate immunity. Nat. Commun. 2014, 5, 5488. https://doi.org/10.1038/ncomms6488.
- Romano, N.; Macino, G. Quelling: Transient inactivation of gene expression in Neurospora crassa by transformation with homologous sequences. Mol. Microbiol. 1992, 6, 3343–3353. https://doi.org/10.1111/j.1365-2958.1992.tb02202.x.
- Koch, A.; Kogel, K.-H. New wind in the sails: Improving the agronomic value of crop plants through RNAi-mediated gene silencing. Plant Biotechnol. J. 2014, 12, 821–831. https://doi.org/10.1111/pbi.12226.
- Nunes, C.C.; Dean, R.A. Host-induced gene silencing: A tool for understanding fungal host interaction and for developing novel disease control strategies. Mol. Plant Pathol. 2011, 13, 519–529. https://doi.org/10.1111/j.1364-3703.2011.00766.x.
- Bartel, D.P. MicroRNAs: Genomics, Biogenesis, Mechanism, and Function. Cell 2004, 116, 281–297. https://doi.org/10.1016/s0092-8674(04)00045-5.
- Wang, M.; Weiberg, A.; Jin, H. Pathogen small RNAs: A new class of effectors for pathogen attacks. Mol. Plant Pathol. 2015, 16, 219–223. https://doi.org/10.1111/mpp.12233.
- Wang, M.; Weiberg, A.; Lin, F.-M.; Thomma, B.P.H.J.; Huang, H.-D.; Jin, H. Bidirectional cross-kingdom RNAi and fungal uptake of external RNAs confer plant protection. Nat. Plants 2016, 2, 1–10. https://doi.org/10.1038/nplants.2016.151.
- Derbyshire, M.C.; Mbengue, M.; Barascud, M.; Navaud, O.; Raffaele, S. Small RNAs from the plant pathogenic fungus Scle-rotinia sclerotiorum highlight candidate host target genes associated with quantitative disease resistance. bioRxiv 2018, 354076. https://doi.org/10.1101/354076.
- Zhao, J.-P.; Jiang, X.-L.; Zhang, B.-Y.; Su, X.-H. Involvement of microRNA-Mediated Gene Expression Regulation in the Pathological Development of Stem Canker Disease in Populus trichocarpa. PLoS ONE 2012, 7, e44968. https://doi.org/10.1371/journal.pone.0044968.
- Gupta, O.P.; Permar, V.; Koundal, V.; Singh, U.D.; Praveen, S. MicroRNA regulated defense responses in Triticum aestivum L. during Puccinia graminis f.sp. tritici infection. Mol. Biol. Rep. 2011, 39, 817–824. https://doi.org/10.1007/s11033-011-0803-5.
- Xin, M.; Wang, Y.; Yao, Y.; Xie, C.; Peng, H.; Ni, Z.; Sun, Q. Diverse set of microRNAs are responsive to powdery mildew infection and heat stress in wheat (Triticum aestivum L.). BMC Plant Biol. 2010, 10, 123. https://doi.org/10.1186/1471-2229-10-123.
- Lau, S.K.P.; Chow, W.-N.; Wong, A.Y.P.; Yeung, J.M.Y.; Bao, J.; Zhang, N.; Lok, S.; Woo, P.C.Y.; Yuen, K.-Y. Identification of MicroRNA-Like RNAs in Mycelial and Yeast Phases of the Thermal Dimorphic Fungus Penicillium marneffei. PLOS Neglected Trop. Dis. 2013, 7, e2398. https://doi.org/10.1371/journal.pntd.0002398.
- Rose, L.E.; Overdijk, E.J.R.; van Damme, M. Small RNA molecules and their role in plant disease. Eur. J. Plant Pathol. 2018, 154, 115–128. https://doi.org/10.1007/s10658-018-01614-w.
- Hudzik, C.; Hou, Y.; Ma, W.; Axtell, M.J. Exchange of Small Regulatory RNAs between Plants and Their Pests. Plant Physiol. 2019, 182, 51–62. https://doi.org/10.1104/pp.19.00931.
- Sperschneider, J.; Jones, A.W.; Nasim, J.; Xu, B.; Jacques, S.; Zhong, C.; Upadhyaya, N.M.; Mago, R.; Hu, Y.; Figueroa, M.; et al. The stem rust fungus Puccinia graminis f. sp. tritici induces centromeric small RNAs during late infection that are associated with genome-wide DNA methylation. BMC Biol. 2021, 19, 1–25. https://doi.org/10.1186/s12915-021-01123-z.
- Mueth, N.A.; Hulbert, S.H. Small RNAs target native and cross-kingdom transcripts on both sides of the wheat stripe rust interaction. Genomics 2022, 114, 110526. https://doi.org/10.1016/j.ygeno.2022.110526.
- Li, Y.; Cao, X.; Zhu, Y.; Yang, X.; Zhang, K.; Xiao, Z.; Wang, H.; Zhao, J.; Zhang, L.; Li, G.; et al. Osa‐miR398b boosts H 2 O 2 production and rice blast disease‐resistance via multiple superoxide dismutases. N. Phytol. 2019, 222, 1507–1522. https://doi.org/10.1111/nph.15678.
- Campo, S.; Peris‐Peris, C.; Siré, C.; Moreno, A.B.; Donaire, L.; Zytnicki, M.; Notredame, C.; Llave, C.; San Segundo, B. Identi-fication of a novel micro RNA (mi RNA) from rice that targets an alternatively spliced transcript of the N ramp6 (N atural resistance‐associated macrophage protein 6) gene involved in pathogen resistance. N. Phytol. 2013, 199, 212–227. https://doi.org/10.1111/nph.12292.
- Salvador-Guirao, R.; Hsing, Y.-I.; Segundo, B.S. The Polycistronic miR166k-166h Positively Regulates Rice Immunity via Post-transcriptional Control of EIN2. Front. Plant Sci. 2018, 9, 337. https://doi.org/10.3389/fpls.2018.00337.
- Zhu, C.; Liu, J.-H.; Zhao, J.-H.; Liu, T.; Chen, Y.-Y.; Wang, C.-H.; Zhang, Z.-H.; Guo, H.-S.; Duan, C.-G. A fungal effector suppresses the nuclear export of AGO1–miRNA complex to promote infection in plants. Proc. Natl. Acad. Sci. USA 2022, 119, e2114583119. https://doi.org/10.1073/pnas.2114583119.
- Koch, A.; Wassenegger, M. Host‐induced gene silencing–Mechanisms and applications. N. Phytol. 2021, 231, 54–59. https://doi.org/10.1111/nph.17364.
- Nowara, D.; Gay, A.; Lacomme, C.; Shaw, J.; Ridout, C.; Douchkov, D.; Hensel, G.; Kumlehn, J.; Schweizer, P. HIGS: Host-Induced Gene Silencing in the Obligate Biotrophic Fungal Pathogen Blumeria graminis. Plant Cell 2010, 22, 3130–3141. https://doi.org/10.1105/tpc.110.077040.
- Huang, G.; Allen, R.; Davis, E.L.; Baum, T.J.; Hussey, R.S. Engineering broad root-knot resistance in transgenic plants by RNAi silencing of a conserved and essential root-knot nematode parasitism gene. Proc. Natl. Acad. Sci. USA 2006, 103, 14302–14306. https://doi.org/10.1073/pnas.0604698103.
- Panwar, V.; McCallum, B.; Bakkeren, G. Host-induced gene silencing of wheat leaf rust fungus Puccinia triticina pathogenicity genes mediated by the Barley stripe mosaic virus. Plant Mol. Biol. 2013, 81, 595–608. https://doi.org/10.1007/s11103-013-0022-7.
- Qi, T.; Guo, J.; Peng, H.; Liu, P.; Kang, Z.; Guo, J. Host-Induced Gene Silencing: A Powerful Strategy to Control Diseases of Wheat and Barley. Int. J. Mol. Sci. 2019, 20, 206. https://doi.org/10.3390/ijms20010206.
- Koch, A.; Kumar, N.; Weber, L.; Keller, H.; Imani, J.; Kogel, K.-H. Host-induced gene silencing of cytochrome P450 lanosterol C14α-demethylase–encoding genes confers strong resistance to Fusarium species. Proc. Natl. Acad. Sci. USA 2013, 110, 19324–19329. https://doi.org/10.1073/pnas.1306373110.
- Pliego, C.; Nowara, D.; Bonciani, G.; Gheorghe, D.M.; Xu, R.; Surana, P.; Whigham, E.; Nettleton, D.; Bogdanove, A.J.; Wise, R.P.; et al. Host-Induced Gene Silencing in Barley Powdery Mildew Reveals a Class of Ribonuclease-Like Effectors. Mol. Plant-Microbe Interact. 2013, 26, 633–642. https://doi.org/10.1094/mpmi-01-13-0005-r.
- Cheng, W.; Song, X.-S.; Li, H.P.; Cao, L.-H.; Sun, K.; Qiu, X.-L.; Xu, Y.-B.; Yang, P.; Huang, T.; Zhang, J.-B.; et al. Host-induced gene silencing of an essential chitin synthase gene confers durable resistance to Fusarium head blight and seedling blight in wheat. Plant Biotechnol. J. 2015, 13, 1335–1345. https://doi.org/10.1111/pbi.12352.
- Chen, W.; Kastner, C.; Nowara, D.; Oliveira-Garcia, E.; Rutten, T.; Zhao, Y.; Deising, H.B.; Kumlehn, J.; Schweizer, P. Host-induced silencing of Fusarium culmorum genes protects wheat from infection. J. Exp. Bot. 2016, 67, 4979–4991. https://doi.org/10.1093/jxb/erw263.
- Zhu, X.; Qi, T.; Yang, Q.; He, F.; Tan, C.; Ma, W.; Voegele, R.T.; Kang, Z.; Guo, J. Host-Induced Gene Silencing of the MAPKK Gene PsFUZ7 Confers Stable Resistance to Wheat Stripe Rust. Plant Physiol. 2017, 175, 1853–1863. https://doi.org/10.1104/pp.17.01223.
- Panwar, V.; Jordan, M.; McCallum, B.; Bakkeren, G. Host-induced silencing of essential genes in Puccinia triticina through transgenic expression of RNAi sequences reduces severity of leaf rust infection in wheat. Plant Biotechnol. J. 2017, 16, 1013–1023. https://doi.org/10.1111/pbi.12845.
- Qi, T.; Zhu, X.; Tan, C.; Liu, P.; Guo, J.; Kang, Z.; Guo, J. Host-induced gene silencing of an important pathogenicity factor PsCPK1 in Puccinia striiformis f. sp. tritici enhances resistance of wheat to stripe rust. Plant Biotechnol. J. 2017, 16, 797–807. https://doi.org/10.1111/pbi.12829.
- Ridout, C.; Skamnioti, P.; Porritt, O.; Sacristan, S.; Jones, J.; Brown, J.K. Multiple Avirulence Paralogues in Cereal Powdery Mildew Fungi May Contribute to Parasite Fitness and Defeat of Plant Resistance. Plant Cell 2006, 18, 2402–2414. https://doi.org/10.1105/tpc.106.043307.
- Panwar, V.; McCallum, B.; Bakkeren, G. Endogenous silencing of Puccinia triticina pathogenicity genes through in plan-ta-expressed sequences leads to the suppression of rust diseases on wheat. Plant J. 2012, 73, 521–532. https://doi.org/10.1111/tpj.12047.
- Kusch, S.; Singh, M.; Thieron, H.; Spanu, P.D.; Panstruga, R. Site-specific analysis reveals candidate cross-kingdom small RNAs, tRNA and rRNA fragments, and signs of fungal RNA phasing in the barley-powdery mildew interaction. bioRxiv 2022, 501657. https://doi.org/10.1101/2022.07.26.501657.
- Bai, G.; Shaner, G. Management and resistance in wheat and barley to fusarium head blight. Annu. Rev. Phytopathol. 2004, 42, 135–161. https://doi.org/10.1146/annurev.phyto.42.040803.140340.
- Scherm, B.; Balmas, V.; Spanu, F.; Pani, G.; Delogu, G.; Pasquali, M.; Migheli, Q. Fusarium culmorum: Causal agent of foot and root rot and head blight on wheat. Mol. Plant Pathol. 2012, 14, 323–341. https://doi.org/10.1111/mpp.12011.
- Dean, R.; Van Kan, J.A.L.; Pretorius, Z.A.; Hammond-Kosack, K.E.; Di Pietro, A.; Spanu, P.D.; Rudd, J.J.; Dickman, M.; Kahmann, R.; Ellis, J.; et al. The Top 10 fungal pathogens in molecular plant pathology. Mol. Plant Pathol. 2012, 13, 414–430. https://doi.org/10.1111/j.1364-3703.2011.00783.x.
- Zhu, L.; Zhu, J.; Liu, Z.; Wang, Z.; Zhou, C.; Wang, H. Host-Induced Gene Silencing of Rice Blast Fungus Magnaporthe oryzae Pathogenicity Genes Mediated by the Brome Mosaic Virus. Genes 2017, 8, 241. https://doi.org/10.3390/genes8100241.
- Guo, X.-Y.; Li, Y.; Fan, J.; Xiong, H.; Xu, F.-X.; Shi, J.; Shi, Y.; Zhao, J.-Q.; Wang, Y.-F.; Cao, X.-L.; et al. Host-Induced Gene Silencing of MoAP1 Confers Broad-Spectrum Resistance to Magnaporthe oryzae. Front. Plant Sci. 2019, 10, 433. https://doi.org/10.3389/fpls.2019.00433.
- Gibbings, D.; Voinnet, O. Control of RNA silencing and localization by endolysosomes. Trends Cell Biol. 2010, 20, 491–501. https://doi.org/10.1016/j.tcb.2010.06.001.
- Van Niel, G.; D’Angelo, G.; Raposo, G. Shedding light on the cell biology of extracellular vesicles. Nat. Rev. Mol. Cell Biol. 2018, 19, 213–228. https://doi.org/10.1038/nrm.2017.125.
- Chaloner, T.; van Kan, J.; Grant-Downton, R.T. RNA ‘Information Warfare’ in Pathogenic and Mutualistic Interactions. Trends Plant Sci. 2016, 21, 738–748. https://doi.org/10.1016/j.tplants.2016.05.008.
- Hua, C.; Zhao, J.-H.; Guo, H.-S. Trans-Kingdom RNA Silencing in Plant–Fungal Pathogen Interactions. Mol. Plant 2018, 11, 235–244. https://doi.org/10.1016/j.molp.2017.12.001.
- Cai, Q.; He, B.; Weiberg, A.; Buck, A.H.; Jin, H. Small RNAs and extracellular vesicles: New mechanisms of cross-species communication and innovative tools for disease control. PLOS Pathog. 2019, 15, e1008090. https://doi.org/10.1371/journal.ppat.1008090.
- Da Rocha, I.F.M.; Amatuzzi, R.F.; Lucena, A.C.R.; Faoro, H.; Alves, L.R. Cross-Kingdom Extracellular Vesicles EV-RNA Communication as a Mechanism for Host–Pathogen Interaction. Front. Cell. Infect. Microbiol. 2020, 10, 593160. https://doi.org/10.3389/fcimb.2020.593160.
- Liu, G.; Kang, G.; Wang, S.; Huang, Y.; Cai, Q. Extracellular Vesicles: Emerging Players in Plant Defense against Pathogens. Front. Plant Sci. 2021, 12, 757925. https://doi.org/10.3389/fpls.2021.757925.
- Kwon, S.; Tisserant, C.; Tulinski, M.; Weiberg, A.; Feldbrügge, M. Inside-out: From endosomes to extracellular vesicles in fungal RNA transport. Fungal Biol. Rev. 2020, 34, 89–99. https://doi.org/10.1016/j.fbr.2020.01.001.
- Bleackley, M.R.; Samuel, M.; Garcia-Ceron, D.; McKenna, J.A.; Lowe, R.; Pathan, M.; Zhao, K.; Ang, C.-S.; Mathivanan, S.; Anderson, M.A. Extracellular Vesicles from the Cotton Pathogen Fusarium oxysporum f. sp. vasinfectum Induce a Phytotoxic Response in Plants. Front. Plant Sci. 2020, 10, 1610. https://doi.org/10.3389/fpls.2019.01610.
- Garcia‐Ceron, D.; Dawson, C.S.; Faou, P.; Bleackley, M.R.; Anderson, M.A. Size‐exclusion chromatography allows the isolation of EVs from the filamentous fungal plant pathogen Fusarium oxysporum f. sp. vasinfectum (Fov). Proteomics 2021, 21, e2000240. https://doi.org/10.1002/pmic.202000240.
- Koch, A.; Schlemmer, T.; Lischka, R. Elucidating the role of extracellular vesicles in the Barley-Fusarium interaction. Trillium Extracell. Vesicles 2020, 2, 28–35. https://doi.org/10.47184/tev.2020.01.03.
- Hill, E.H.; Solomon, P.S. Extracellular vesicles from the apoplastic fungal wheat pathogen Zymoseptoria tritici. Fungal Biol. Biotechnol. 2020, 7, 1–14. https://doi.org/10.1186/s40694-020-00103-2.
- Kwon, S.; Rupp, O.; Brachmann, A.; Blum, C.; Kraege, A.; Goesmann, A.; Feldbrügge, M. mRNA Inventory of Extracellular Vesicles from Ustilago maydis. J. Fungi 2021, 7, 562. https://doi.org/10.3390/jof7070562.
- Schlemmer, T.; Lischka, R.; Wegner, L.; Ehlers, K.; Biedenkopf, D.; Koch, A. Extracellular vesicles isolated from dsRNA-sprayed barley plants exhibit no growth inhibition or gene silencing in Fusarium graminearum. Fungal Biol. Biotechnol. 2022, 9, 1–14. https://doi.org/10.1186/s40694-022-00143-w.
- Halder, V.; Kombrink, E. Facile high-throughput forward chemical genetic screening by in situ monitoring of glucuroni-dase-based reporter gene expression in Arabidopsis thaliana. Front. Plant Sci. 2015, 6, 13. https://doi.org/10.3389/fpls.2015.00013.
- Westermann, A.; Gorski, S.; Vogel, J. Dual RNA-seq of pathogen and host. Nat. Rev. Genet. 2012, 10, 618–630. https://doi.org/10.1038/nrmicro2852.
- Asai, S.; Rallapalli, G.; Piquerez, S.J.M.; Caillaud, M.-C.; Furzer, O.; Ishaque, N.; Wirthmueller, L.; Fabro, G.; Shirasu, K.; Jones, J.D.G. Expression Profiling during Arabidopsis/Downy Mildew Interaction Reveals a Highly-Expressed Effector That Attenu-ates Responses to Salicylic Acid. PLOS Pathog. 2014, 10, e1004443. https://doi.org/10.1371/journal.ppat.1004443.
- Murugan, K.; Babu, K.; Sundaresan, R.; Rajan, R.; Sashital, D.G. The Revolution Continues: Newly Discovered Systems Expand the CRISPR-Cas Toolkit. Mol. Cell 2017, 68, 15–25. https://doi.org/10.1016/j.molcel.2017.09.007.
- Rutter, B.D.; Innes, R.W. Extracellular Vesicles Isolated from the Leaf Apoplast Carry Stress-Response Proteins. Plant Physiol. 2016, 173, 728–741. https://doi.org/10.1104/pp.16.01253.
