Gene elongation is a molecular mechanism consisting of an in-tandem duplication of a gene and divergence and fusion of the two copies, resulting in a gene constituted by two divergent paralogous modules. Several examples of genes with internal sequence repetitions are reported in literature; thus, gene elongation might have shaped the structures of many genes during the first steps of molecular and cellular evolution.
- paralogous modules
- gene duplication
- gene fusion
1. Introduction
Gene elongation consists of an in-tandem gene duplication, which produces two (or more) copies of the same gene, followed by the deletion of the intervening region and the conversion (by mutation) of the stop codon of the first copy into a sense one, resulting in the elongation, by fusion, of the initial gene and its copy. Thus, gene elongation is actually the combination of gene duplication and gene fusion. The newly formed gene is constituted by two paralogous modules, which might independently undergo different mutations and further duplications [3] that, over time, can hide the traces of these early events.
2. Data
As shown in Figure 1, once the two copies have originated, in principle they might follow two different evolutionary pathways in which they can either immediately fuse and then diverge or, vice versa, diverge and then fuse. The final outcome of these two alternative pathways is the same: the formation of a gene constituted by two divergent paralogous modules. Potentially, each module or both of them might undergo further duplication events, leading to a gene coding for more repetitions of amino acid sequences. Gene elongation produces two or more copies of the same protein fold within a single polypeptide chain, often generating a direct repeat of domains and thus a pseudo-symmetrical structure with internal symmetry axes [1]. Many present-day proteins show internal repeats of amino acid sequences, which often correspond to functional or structural domains; since gene elongation has occurred in so many cases, this event must be considered an evolutionary advantage. The biological significance of proteins with repetitive structures might include (i) the improvement of the protein function by increasing the number of active sites; (ii) the acquisition of an additional function by modifying a redundant segment [3], thus obtaining a bifunctional enzyme; and/or (iii) the stabilization of a protein structure, thus increasing the enzyme’s catalytic activity. Several cases of genes with internal sequence repetitions are reported in the literature. For example, in the bacterial ferredoxin, the second half of the amino acid sequence is an almost exact duplicate of the first [8]. Tang et al. [9] observed that the pepsin family of proteases have an intramolecular two-fold symmetry axis that relates two topologically similar domains, and proposed a mechanism for its evolution by gene elongation. The carB gene of Escherichia coli, which encodes a subunit of carbamoyl-phosphate synthetase, was proposed to be formed by the duplication of an ancestral gene, since its amino acid sequence shows a highly significant similarity between the amino- and carboxyl-terminal halves of the protein [10]. Rubin et al. [11] found that the two halves of Gram-negative bacterial tetracycline efflux pumps share a process of tandem gene duplication and divergence. Gupta and Singh [12] suggested a model for the evolution of the heat-shock protein 70 (Hsp70) of Archaea and Bacteria based on gene duplication. Moreover, domain fusions have occurred in the evolution of the ATP binding cassette (ABC) superfamily [13]. However, the most documented case of gene elongation involves the histidine biosynthetic genes hisA and hisF; they are paralogous and very likely originated from the duplication (and the further divergence and fusion of the two resulting copies) of an ancestral gene, half the size of the extant genes [14]. The biological significance of this gene elongation event became clear when the structures of the HisA and HisF proteins were determined. Crystallographic studies showed that these two enzymes are structurally homologous (β/α)8-barrels (TIM-barrels) [37], with a two-fold repeat pattern; the first and second (β/α)4-half barrels of both enzymes are related by a two-fold axis of symmetry. These data led to the proposal of a model for the evolution of the HisA and HisF (β/α)8-barrels via two successive gene duplication events: a first duplication of a single ancestor gene encoding a (β/α)4-half barrel, subsequent fusion and divergent evolution, followed by a second gene duplication and diversification leading to the extant genes encoding the enzymes HisA and HisF [32]. Fani et al. [15] investigated the possibility of an even older gene elongation event, starting from (β/α)-mers smaller than the (β/α)4 units. According to this idea, the ancestor gene might have encoded a single (β/α)-module that, in turn, might have been able to aggregate in a homo-octamer to form a still unstable and thus not efficient complete TIM-barrel. This ancestral TIM-barrel enzyme might have been endowed with a broad specificity, catalyzing different enzymatic reactions. Subsequently, a “cascade” of three gene elongation events would have given rise to the complete ancestor of the extant TIM-barrel coding genes. It is quite possible that a similar evolutionary pathway might have shaped other genes coding for enzymes with internal repeats, such as proteins with repeated trans-membrane domains. The hisA and hisF genes share the same internal organization in all histidine-synthesizing organisms (Archaea, Bacteria, and Eucarya) suggesting that the elongation event(s) leading to the extant hisA and hisF very likely occurred (long) before the appearance of the Last Universal Common Ancestor (LUCA) [15]. This finding may suggest that gene elongation has shaped the structures of many (other) genes during the first steps of molecular and cellular evolution.
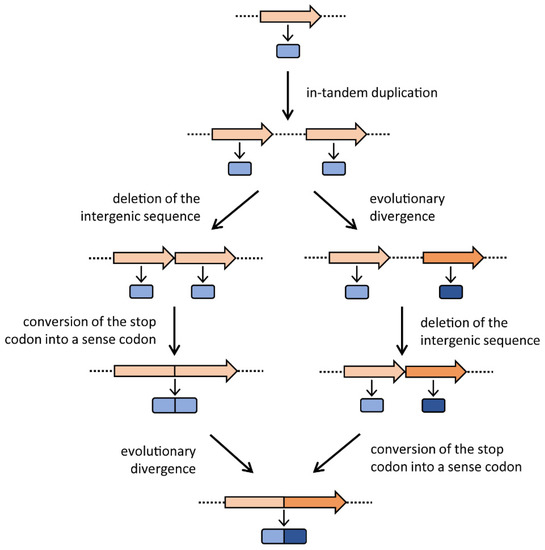
Figure 1. The gene elongation mechanism. The two possible evolutionary routes are depicted. Genes are represented with arrows, and the encoded proteins with rounded rectangles. Representation adapted from Fani and Fondi [3].
