Hepatitis C Virus (HCV) is an enveloped virus with an RNA genome of positive polarity. HCV replicates mainly in the liver and can cause liver disease, cirrhosis and cancer. Translation of the HCV RNA genome is regulated by the Internal Ribosome Entry Site (IRES) and other cis-elements in the viral RNA genome. The viral RNA usurps cellular ribosomes using a variety of viral RNA elements as well as by recruiting cellular RNA binding proteins. Here, we give a comprehensive overview over the determinants involved in the regulation of HCV translation.
- HCV
- Hepacivirus
- Internal Ribosome Entry Site
- IRES
- Initiation
- Ribosome
- 40S
- eIF3
- ITAF
- stress response
1. Introduction
Hepatitis C virus (HCV) is an enveloped positive strand RNA virus that preferentially replicates in the liver [1], and it is classified in the genus Hepacivirus in the family Flaviviridae. Worldwide, about 71 million people are infected with HCV [2]. The infection is usually noticed only when coincidentally diagnosed by routine testing, for example, during hospitalization, or when the liver disease becomes acute. In the latter case, liver damage by virus replication and the resulting immune responses can lead to impaired bilirubin conjugation in the liver, and unconjugated bilirubin deposits can then be noticed as a yellowish color (called jaundice), often first in the sclera in the eyes and when more severe also in the skin. An acute infection can result in severe liver damage, in rare cases even resulting in death [3][4]. However, most HCV infections remain inapparent [5][6], and the virus infection can become chronic in about 60% to 70 % of all infections [7], often without being noticed. Chronic infection can, in the long run, result in liver cirrhosis and liver cancer (hepatocellular carcinoma, HCC) [8][9][10], while a metabolic reprogramming of the infected cells according to the “Warburg effect” like in cancer cells can be observed only a few days after the onset of HCV replication [11]. Moreover, inapparent replication of the virus usually results in unnoticed spread of the virus to other individuals, a fact that is a major challenge for surveillance, health care, and treatment [12]. Meanwhile, very effective treatment regimens using direct acting antivirals (DAAs) are available, although they are still very expensive [13][14]. Although the error rate of the viral replicase is high and can, in principle, easily give rise to resistance mutations, the conserved nature of the replicase active center and the often occurring reduced fitness of mutants result in the rare appearance of resistance mutations against nucleoside inhibitors such as sofosbuvir [15][16]. An effective vaccination is not yet available, also partially due to the high variability of the viral RNA genome. Thus, further research on HCV is urgently required to combat HCV infections, and despite much progress in the understanding of HCV replication, the molecular mechanisms of HCV replication are still far from being completely understood [12].
2. Hepatitis C virus (HCV)
The HCV RNA genome is about 9600 nucleotides (nts) long and codes for one long polyprotein that is co- and post-translationally processed into the mature gene products [17][18][19]. Unlike most cellular mRNAs, the 5′-end of the HCV genomic RNA has no cap nucleotide attached which would govern efficient cap-dependent translation initiation [20][21][22]. Instead, HCV translation is mediated by virtue of an internal ribosome entry site (IRES) [23][24][25][26][27], which is located largely in the 5′UTR but also slightly spans into the coding region (Figure 1). While the resulting low efficient translation coincides with the “undercover” strategy of HCV replication that often leads to chronic infection and further unnoticed spread of the virus to uninfected individuals, the use of such IRES elements has two more big advantages. The first advantage is that in particular the very ends of the RNA genome do not need to serve functions in translation control such as in capped and polyadenylated cellular mRNAs. Instead, RNA signals that are involved in genome replication can be directly placed at the very genome ends [28][18][29]. The second benefit of cap-independent translation is that the virus escapes antiviral countermeasures of the cell in terms of the downregulation of cap-dependent translation, which is largely conferred by phosphorylation of eIF2 and the resulting inhibition of cap-dependent translation initiation [30].
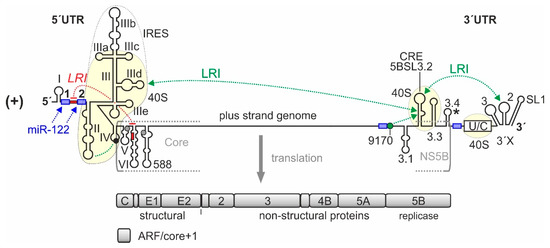
Figure 1.
cis
-Elements in the hepatitis C virus (HCV) RNA genome that are involved in translation regulation. The HCV plus strand RNA genome. The internal ribosome entry site (IRES) in the HCV 5′-untranslated region (5′UTR), the entire 3′UTR and the
cis
-acting replication element (CRE) in the NS5B coding region are involved in translation regulation
. Those regions of the 5′UTR, 3′UTR, and CRE that bind to the ribosomal 40S subunit are underlayed in light yellow. Stem-loops (SLs) in the 5′UTR are numbered by roman numerals. The region of the IRES is surrounded by a dotted line. The IRES includes SLs II–IV of the 5′UTR but spans into the core protein coding region. The canonical AUG start codon in SL IV of the 5′UTR (black circle) gives rise to translation of the polyprotein which is cleaved to yield structural proteins and non-structural (NS-) proteins, including the viral replicase NS5B. The 3′UTR contains the variable region, a poly(U/C) tract (U/C), and the so-called 3′X region including SLs 1, 2, and 3. The stem-loop 5BSL3.2 in the 3′-region of the NS5B coding region is the CRE, flanked by upstream stem-loop 5BSL3.1 and downstream 5BSL3.3. The polyprotein stop codon is located in the stem-loop 5BSL3.4 (asterisk). Some other start codons which give rise to the alternative reading frame (ARF) in the core+1 reading frame
are shown in grey. Positively and negatively acting long-range RNA–RNA interactions (LRIs) are shown in green or red, respectively, with the sequence “9170” shown as green circle. Selected binding sites for microRNA-122 (miR-122) are shown in blue.
The HCV IRES is located in the 5´-untranslated region (5´UTR), part of the core protein coding sequence, and by the 3´UTR. The 5´UTR has some highly conserved structural regions, while others can assume different conformations. The IRES can bind to the ribosomal 40S subunit with high affinity without any other factors. Nevertheless, IRES activity is modulated by additional cis sequences in the viral genome, including the 3´UTR and the cis-acting replication element (CRE). Canonical translation initiation factors (eIFs) are involved in HCV translation initiation, including eIF3, eIF2, eIF1A, eIF5 and eIF5B. Alternatively, under stress conditions and limited eIF2-Met-tRNAiMet availability, alternative initiation factors like eIF2D, eIF2A and/or eIF5B can substitute for eIF2 to allow HCV translation even when cellular mRNA translation is downregulated. In addition, several IRES trans-acting factors (ITAFs) modulate IRES activity by building large networks of RNA-protein and protein-protein interactions, also connecting 5´- and 3´-ends of the viral RNA. Moreover, some ITAFs may act as RNA chaperones that help to position the viral AUG start codon in the ribosomal 40S subunit entry channel. Finally, the liver-specific microRNA-122 (miR-122) stimulates HCV IRES-dependent translation, most likely by stabilizing a certain structure of the IRES that is required for initiation.
In this entreviewy, we focus on the sequences, cellular factors, and molecular mechanisms involved in the regulation of translation by the HCV IRES. Thereby, we touch the functions of miR-122 specifically only with regard to translation regulation.
References
- M P Busch; J C Wilber; Hepatitis C Virus Replication. New England Journal of Medicine 1992, 326, 64-66, 10.1056/nejm199201023260114.
- WHO World Health Organization (WHO). Fact Sheets—Hepatitis C. Available online: http://www.who.int/en/news-room/fact-sheets/detail/hepatitis-c (accessed on 20 February 2020).
- Patrizia Farci; Harvey J. Alter; Atsushi Shimoda; Sugantha Govindarajan; Ling C. Cheung; Jacqueline C. Melpolder; Ronald A. Sacher; James W. Shih; R H Purcell; Hepatitis C Virus–Associated Fulminant Hepatic Failure. New England Journal of Medicine 1996, 335, 631-634, 10.1056/nejm199608293350904.
- Tatsuo Kanda; Osamu Yokosuka; Fumio Imazeki; Hiromitsu Saisho; Acute hepatitis C virus infection, 1986-2001: a rare cause of fulminant hepatitis in Chiba, Japan.. Hepatogastroenterology 2004, 51, 556–558.
- R. Buchanan; Theresa Hydes; S. I. Khakoo; Innate and adaptive genetic pathways in HCV infection. Tissue Antigens 2015, 85, 231-240, 10.1111/tan.12540.
- Barbara Rehermann; Robert Thimme; Insights From Antiviral Therapy Into Immune Responses to Hepatitis B and C Virus Infection. Gastroenterology 2019, 156, 369-383, 10.1053/j.gastro.2018.08.061.
- Alessia Virzì; Armando Andres Roca Suarez; Thomas F. Baumert; Joachim Lupberger; Rewiring Host Signaling: Hepatitis C Virus in Liver Pathogenesis. Cold Spring Harbor Perspectives in Medicine 2019, 10, a037366, 10.1101/cshperspect.a037366.
- R. Waziry; B. Hajarizadeh; Jason Grebely; J. Amin; M. Law; M. Danta; J. George; G.J. Dore; No evidence for higher risk of hepatocellular carcinoma occurrence or recurrence following direct-acting antiviral HCV therapy: a systematic review, meta-analyses, and meta-regression. Journal of Hepatology 2017, 66, S12, 10.1016/s0168-8278(17)30286-6.
- C. Hayes; Peiyi Zhang; Yizhou Zhang; Kazuaki Chayama; Molecular Mechanisms of Hepatocarcinogenesis Following Sustained Virological Response in Patients with Chronic Hepatitis C Virus Infection. Viruses 2018, 10, 531, 10.3390/v10100531.
- Seong Kyun Na; Byung-Cheol Song; Development and surveillance of hepatocellular carcinoma in patients with sustained virologic response after antiviral therapy for chronic hepatitis C. Clinical and Molecular Hepatology 2019, 25, 234-244, 10.3350/cmh.2018.0108.
- Gesche Gerresheim; Roeb; Audrey M Michel; Michael Niepmann; Elke Roeb; Hepatitis C Virus Downregulates Core Subunits of Oxidative Phosphorylation, Reminiscent of the Warburg Effect in Cancer Cells. Cells 2019, 8, 1410, 10.3390/cells8111410.
- Ralf Bartenschlager; Thomas Baumert; Jens Bukh; Michael Houghton; Stanley M. Lemon; Brett D. Lindenbach; Volker Lohmann; Darius Moradpour; Thomas Pietschmann; Charles M. Rice; Robert Thimme; Takaji Wakita; Critical challenges and emerging opportunities in hepatitis C virus research in an era of potent antiviral therapy: Considerations for scientists and funding agencies. Virus Research 2018, 248, 53-62, 10.1016/j.virusres.2018.02.016.
- Nathalie Alazard-Dany; Solène Denolly; Bertrand Boson; François-Loïc Cosset; Overview of HCV Life Cycle with a Special Focus on Current and Possible Future Antiviral Targets. Viruses 2019, 11, 30, 10.3390/v11010030.
- Yueran Zhuo; Tomoyuki Hayashi; Qiushi Chen; Rakesh Aggarwal; Yvan Hutin; Jagpreet Chhatwal; Estimating the price at which hepatitis C treatment with direct-acting antivirals would be cost-saving in Japan. Scientific Reports 2020, 10, 1-9, 10.1038/s41598-020-60986-4.
- Evguenia S. Svarovskaia; Hadas Dvory-Sobol; Neil Parkin; Christy Hebner; Viktoria Gontcharova; Ross Martin; Wen Ouyang; Bin Han; Simin Xu; Karin Ku; Sophia Chiu; Edward Gane; Ira M. Jacobson; David R. Nelson; Eric Lawitz; David L. Wyles; Neby Bekele; Diana Brainard; William T. Symonds; John G. McHutchison; Michael D. Miller; Hongmei Mo; Infrequent development of resistance in genotype 1-6 hepatitis C virus-infected subjects treated with sofosbuvir in phase 2 and 3 clinical trials.. Clinical Infectious Diseases 2014, 59, 1666-74, 10.1093/cid/ciu697.
- Eric F. Donaldson; Patrick R. Harrington; Julian J. O'rear; Lisa K. Naeger; Clinical evidence and bioinformatics characterization of potential hepatitis C virus resistance pathways for sofosbuvir. Hepatology 2014, 61, 56-65, 10.1002/hep.27375.
- Darius Moradpour; F Penin; Hepatitis C Virus Proteins: From Structure to Function. Tuberculosis 2013, 369, 113-142, 10.1007/978-3-642-27340-7_5.
- Choongho Lee; Roles of phosphoinositides and phosphoinositides kinases in hepatitis C virus RNA replication. Archives of Pharmacal Research 2012, 35, 1701-1711, 10.1007/s12272-012-1001-2.
- Christopher Neufeldt; Mirko Cortese; Eliana G. Acosta; Ralf Bartenschlager; Rewiring cellular networks by members of the Flaviviridae family. Nature Reviews Genetics 2018, 16, 125-142, 10.1038/nrmicro.2017.170.
- Alan G. Hinnebusch; Structural Insights into the Mechanism of Scanning and Start Codon Recognition in Eukaryotic Translation Initiation. Trends in Biochemical Sciences 2017, 42, 589-611, 10.1016/j.tibs.2017.03.004.
- William C. Merrick; Graham Pavitt; Protein Synthesis Initiation in Eukaryotic Cells. Cold Spring Harbor Perspectives in Biology 2018, 10, a033092, 10.1101/cshperspect.a033092.
- Nikolay E. Shirokikh; Thomas Preiss; Translation initiation by cap-dependent ribosome recruitment: Recent insights and open questions. Wiley Interdisciplinary Reviews: RNA 2018, 9, e1473, 10.1002/wrna.1473.
- Shelton S. Bradrick; Simardeep Nagyal; Hilary Novatt; A miRNA-responsive cell-free translation system facilitates isolation of hepatitis C virus miRNP complexes. RNA 2013, 19, 1159-1169, 10.1261/rna.038810.113.
- K Tsukiyama-Kohara; N Iizuka; M Kohara; A Nomoto; Internal ribosome entry site within hepatitis C virus RNA.. Journal of Virology 1992, 66, 1476-1483.
- C Wang; P Sarnow; A Siddiqui; Translation of human hepatitis C virus RNA in cultured cells is mediated by an internal ribosome-binding mechanism.. Journal of Virology 1993, 67, 3338-3344.
- Sung Key Jang; Internal initiation: IRES elements of picornaviruses and hepatitis c virus. Virus Research 2006, 119, 2-15, 10.1016/j.virusres.2005.11.003.
- Christian M. T. Spahn; Hepatitis C Virus IRES RNA-Induced Changes in the Conformation of the 40S Ribosomal Subunit. Science 2001, 291, 1959-1962, 10.1126/science.1058409.
- Michael Niepmann; Lyudmila A. Shalamova; Gesche K. Gerresheim; Oliver Rossbach; Signals Involved in Regulation of Hepatitis C Virus RNA Genome Translation and Replication. Frontiers in Microbiology 2018, 9, , 10.3389/fmicb.2018.00395.
- Cristina Romero-López; Alfredo Berzal-Herranz; The 5BSL3.2 Functional RNA Domain Connects Distant Regions in the Hepatitis C Virus Genome. Frontiers in Microbiology 2017, 8, 395, 10.3389/fmicb.2017.02093.
- Wek, R.C. Role of eIF2alpha Kinases in Translational Control and Adaptation to Cellular Stress. Cold Spring Harb. Perspect. Biol. 2018, 10, a032870.
- Cristina Romero-López; Alfredo Berzal-Herranz; The Role of the RNA-RNA Interactome in the Hepatitis C Virus Life Cycle. International Journal of Molecular Sciences 2020, 21, 1479, 10.3390/ijms21041479.
- José L. Walewski; Toby R. Keller; Decherd D. Stump; Andrea D. Branch; Evidence for a new hepatitis C virus antigen encoded in an overlapping reading frame.. RNA 2001, 7, 710-721, 10.1017/s1355838201010111.
- Zhenming Xu; Jinah Choi; T.S.Benedict Yen; Wen Lu; Anne Strohecker; Sugantha Govindarajan; David Chien; Mark J. Selby; Jing‐Hsiung Ou; Synthesis of a novel hepatitis C virus protein by ribosomal frameshift. The EMBO Journal 2001, 20, 3840-3848, 10.1093/emboj/20.14.3840.
- Agoritsa Varaklioti; Niki Vassilaki; Urania Georgopoulou; Penelope Mavromara; Alternate Translation Occurs within the Core Coding Region of the Hepatitis C Viral Genome. Journal of Biological Chemistry 2002, 277, 17713-17721, 10.1074/jbc.m201722200.
- Arnab Basu; Robert Steele; Ranjit Ray; Ratna B. Ray; Functional properties of a 16 kDa protein translated from an alternative open reading frame of the core-encoding genomic region of hepatitis C virus. Journal of General Virology 2004, 85, 2299-2306, 10.1099/vir.0.80028-0.
