Metastasis detection in lymph nodes via microscopic examination of histopathological images is one of the most crucial diagnostic procedures for breast cancer staging. The manual analysis is extremely labor-intensive and time-consuming because of complexities and diversities of histopathology images. Deep learning has been utilized in automatic cancer metastasis detection in recent years. Due to the huge size of whole-slide images, most existing approaches split each image into smaller patches and simply treat these patches independently, which ignores the spatial correlations among them. To solve this problem, this paper proposes an effective spatially sensitive learning framework for cancer metastasis detection in whole-slide images. Moreover, a novel spatial loss function is designed to ensure the consistency of prediction over neighboring patches. Specifically, through incorporating long short-term memory and spatial loss constraint on top of a convolutional
neural network feature extractor, the proposed method can effectively learn both the appearance of each patch and spatial relationships between adjacent image patches. With the standard backpropagation algorithm, the whole framework can be trained in an end-to-end way. Finally, the regions with high tumor probability in the resulting probability map are the metastasis locations. Extensive experiments on the benchmark Camelyon 2016 Grand Challenge dataset show the effectiveness of
the proposed approach with respect to state-of-the-art competitors. The obtained precision, recall, and balanced accuracy are 0.9565, 0.9167, and 0.9458, respectively. It is also demonstrated that the proposed approach can provide more accurate detection results and is helpful for early diagnosis of cancer metastasis.
Metastasis detection in lymph nodes via microscopic examination of histopathological images is one of the most crucial diagnostic procedures for breast cancer staging. The manual analysis is extremely labor-intensive and time-consuming because of complexities and diversities of histopathology images. Deep learning has been utilized in automatic cancer metastasis detection in recent years. Due to the huge size of whole-slide images, most existing approaches split each image into smaller patches and simply treat these patches independently, which ignores the spatial correlations among them.
- deep learning
- convolutional neural network
- long short-term memory
- spatial constraint
1. Introduction
detect the metastasis locations, as shown in Figure 1.
constraint is also imposed on the loss function to further improve performance.
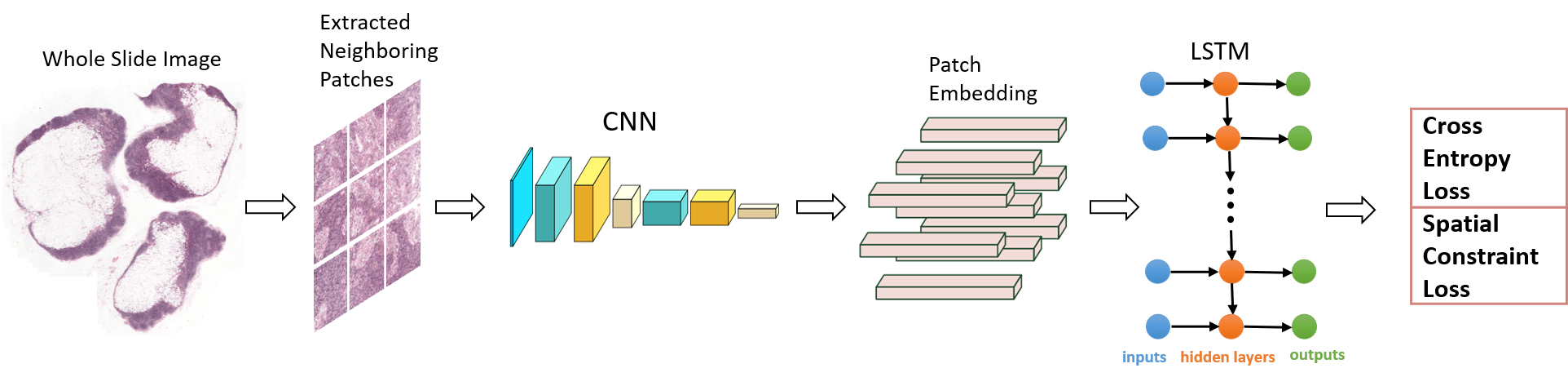
2. Breast Cancer Detection
In earlier years, most approaches employed hand-crafted features for cancer metastasis detection. In Reference [6], authors distinguished the malignant from the benign based on several hand-crafted textural features. Some studies merged two or more hand-crafted features to enhance the accuracy of detection. In Reference [7], graph, haralick, local binary patterns (LBP), and intensity features were used for cancer identification of H&E stained histopathological images. In Reference [8], histopathological images were represented via fusing color histograms, LBP, SIFT, and some kernel features, and the significance of these pattern features was also studied. However, it takes considerable effort to design and validate the hand-made features. In addition, the properties of tissues with great variations in morphology and texture cannot properly be represented. Therefore, the detection performance of these methods based on hand-crafted features is poor. In Reference [16], authors developed a deep learning system to study the stromal properties of breast tissues associated with tumor for classifying whole slide images (WSIs). Authors in Reference [13] utilized AlexNet to categorize breast cancer in histopathological images to be malignant or benign. Authors in Reference [14] developed two distinct CNN architectures to classify breast cancer of pathology images. Single-task CNN was applied to identify malignant tumors. Multi-task CNN was used for analyzing the properties of benign and malignant tumors. The hybrid CNN unit designed in Reference [15] could fully exploit the global and local features of images, and thus obtained superior prediction performance. In References [17[17][18],18], authors proposed a dense and fast screening architecture (ScanNet) to identify metastatic breast cancer in WSIs. Several methods based on transfer learning are also proposed to detect breast cancer [19,20,21][19][20][21]. However, these above approaches deal with each patch of an image individually. In reality, each image patch and its adjacent ones generally share spatial correlations that are crucial for prediction. If one patch is a tumor, its adjacent patches are more likely in the tumor region, as they are situated in the neighboring areas. In order to fully capture the spatial structure information between adjacent patches, authors in Reference [22] applied a conditional random field (CRF) on patch features, which are first obtained from a CNN classifier. However, the method [22] adopted a two-step learning framework, so the spatial relationships between neighboring patches are not available for the CNN feature extractor. Authors in Reference [23] employed 2D long short-term memory (LSTM) on patch features to learn spatial structure information, due to higher computation cost of the end-to-end learning scheme, the two-stage learning strategy was also adopted in their experiments [23].References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424.
- Apple, S.K. Sentinel lymph node in breast cancer: Review article from a pathologist’s point of view. J. Pathol. Transl. Med. 2016, 50, 83–95.
- Ramos-Vara, J.A. Principles and methods of immunohistochemistry. Methods Mol. Biol. 2011, 691, 83–96.
- Humphreys, G.; Ghent, A. World laments loss of pathology service. Bull. World Health Organ. 2010, 88, 564–565.
- Litjens, G.; Sánchez, C.; Timofeeva, N.; Hermsen, M.; Nagtegaal, I.; Kovacs, I.; Kaa, H.; Bult, P.; Ginnneken, B.V.; Laak, J. Deep learning as a tool for increased accuracy and efficiency of histopathological diagnosis. Sci. Rep. 2016, 6, 26286.
- Spanhol, F.A.; Oliveira, L.S.; Petitjean, C.; Heutte, L. A dataset for breast cancer histopathological image classification. IEEE Trans. Biomed. Eng. 2015, 63, 1455–1462.
- Cruz-Roa, A.A.; Ovalle, J.; Madabhushi, A.; Osorio, F. A Deep Learning Architecture for Image Representation, Visual Interpretability and Automated Basal-Cell Carcinoma Cancer Detection. In Proceedings of the 16th International Conference on Medical Image Computing and Computer Assisted Intervention, Nagoya, Japan, 22–26 September 2013; pp. 403–410.
- Kandemir, M.; Hamprecht, F. Computer-aided diagnosis from weak supervision: A benchmarking study. Comput. Med. Imaging Graph. 2015, 42, 44–50.
- Alom, M.Z.; Taha, T.M.; Yakopcic, C.; Westberg, S.; Asari, V.K. The history began from alexnet: A comprehensive survey on deep learning approaches. arXiv 2018, arXiv:1803.01164.
- Litjens, G.; Kooi, T.; Bejnordi, B.E.; Setio, A.A.; Ciompi, F.; Ghafoorian, M.; Laak, J.A.; Ginneken, B.; Sánchez, C.I. A survey on deep learning in medical image analysis. Med. Image Anal. 2017, 42, 60–88.
- Basha, J.; Bacanin, N.; Vukobrat, N.; Zivkovic, M.; Venkatachalam, K.; Hubálovský, S.; Trojovský, P. Chaotic Harris Hawks Optimization with Quasi-Reflection-Based Learning: An Application to Enhance CNN Design. Sensors 2021, 21, 6654.
- Manzo, M.; Pellino, S. Bucket of Deep Transfer Learning Features and Classification Models for Melanoma Detection. J. Imaging 2020, 6, 129.
- Spanhol, F.; Oliveira, L.S.; Cavalin, P.R.; Petitjean, C.; Heutte, L. Deep features for breast cancer histopathological image classification. In Proceedings of the IEEE International Conference on Systems, Man, and Cybernetics, Banff, AB, Canada, 5–8 October 2017; pp. 1868–1873.
- Bayramoglu, N.; Kannala, J.; Heikkilä, J. Deep learning for magnification independent breast cancer histopathology image classification. In Proceedings of the 23rd International Conference on Pattern Recognition (ICPR), Cancun, Mexico, 4–8 December 2016; pp. 2440–2445.
- Guo, Y.; Dong, H.; Song, F.; Zhu, C.; Liu, J. Breast Cancer Histology Image Classification Based on Deep Neural Networks. In International Conference Image Analysis and Recognition; Springer: Cham, Switzerland, 2018; Volume 10882, pp. 827–836.
- Ehteshami Bejnordi, B.; Linz, J.; Glass, B.; Mullooly, M.; Gierach, G.; Sherman, M.; Karssemeijer, N.; van der Laak, J.; Beck, A. Deep learning-based assessment of tumor-associated stroma for diagnosing breast cancer in histopathology images. In Proceedings of the IEEE 14th International Symposium on Biomedical Imaging, Melbourne, VIC, Australia, 18–21 April 2017; pp. 929–932.
- Lin, H.; Chen, H.; Dou, Q.; Wang, L.; Qin, J.; Heng, P.A. ScanNet: A Fast and Dense Scanning Framework for Metastatic Breast Cancer Detection from Whole-Slide Images. In Proceedings of the IEEE Winter Conference on Applications of Computer Vision (WACV), Lake Tahoe, NV, USA, 12–15 March 2018; pp. 539–546.
- Lin, H.; Chen, H.; Graham, S.; Dou, Q.; Rajpoot, N.; Heng, P.A. Fast scannet: Fast and dense analysis of multi-gigapixel whole-slide images for cancer metastasis detection. IEEE Trans. Med. Imaging 2019, 38, 1948–1958.
- Xie, J.; Liu, R.; Luttrell, J.; Zhang, C. Deep Learning Based Analysis of Histopathological Images of Breast Cancer. Front. Genet. 2019, 10, 80.
- de Matos, J.; de Souza Britto, A.; Oliveira, L.; Koerich, A.L. Double Transfer Learning for Breast Cancer Histopathologic Image Classification. In Proceedings of the International Joint Conference on Neural Networks (IJCNN), Budapest, Hungary, 14–19 July 2019; pp. 1–8.
- Kassani, S.H.; Kassani, P.H.; Wesolowski, M.J.; Schneider, K.A.; Deters, R. Breast Cancer Diagnosis with Transfer Learning and Global Pooling. In Proceedings of the International Conference on Information and Communication Technology Convergence (ICTC), Jeju, Korea, 16–18 October 2019; pp. 519–524.
- Zanjani, F.G.; Zinger, S.; With, P. Cancer detection in histopathology whole-slide images using conditional random fields on deep embedded spaces. In Proceedings of the Digital Pathology, Houston, TX, USA, 6 March 2018.
- Kong, B.; Xin, W.; Li, Z.; Qi, S.; Zhang, S. Cancer Metastasis Detection via Spatially Structured Deep Network. In International Conference Image Analysis and Recognition; Springer: Cham, Switzerland, 2017; pp. 236–248.
