Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is a comparison between Version 1 by Chundong Yu and Version 2 by Rita Xu.
Histone demethylase JMJD2D is a multifunctional epigenetic factor coordinating androgen receptor activation, DNA damage repair, DNA replication, cell cycle regulation, and inflammation modulation. JMJD2D is also a well-established epigenetic facilitator in the progression of multiple malignant tumors, especially in colorectal cancer (CRC) and hepatocellular cancer (HCC).
- JMJD2D
- KDM4D
- H3K9me3
1. Introduction
Genetic materials are tightly condensed in a core of positively charged histones that surround negatively charged DNA wraps. The theory of epigenetics was first proposed by Conrad h. Waddington in 1942, which has been recognized as epigenetic modifications mediating cellular phenotype or gene expression through DNA or histone covalent modifications, chromatin remodeling, non-coding RNA, etc., without altering the sequence of DNA [1][2][1,2]. The core histones constituting nucleosomes contain a Lys- and Arg-rich tail or side chain and are extensively modulated by posttranslational modifications (PTMs) [3]. The acetylation, methylation, and phosphorylation modifications of histone have been widely reported, while it can also be modified by other means such as O-acetyl glycosylation, formylation, and ADP-ribosylation [4][5][6][7][8][4,5,6,7,8].
Cancer progression is closely related to the changes in histone PTMs, and the abnormal control of PTMs-related enzymes, including histone methylase, demethylase, acetylase, and acetyltransferase, is an important predisposing factor for tumors [9]. These epigenetic changes may silence multiple tumor suppressor genes or activate oncogenes, leading to the reprogramming of oncogenes in the genome. Allfrey et al. first reported that histone methylation is related to gene transcriptional regulation [10], and then the modifications on Lys, Arg, Ser, Thr, Tyr, and His residues have been reported [11]. The most widely studied modified site is Lys, which can be monomethyl, dimethyl, or trimethyl (i.e., me1, me2, or me3). There is a relatively large lysine methyltransferase family [12]; however, the same type of methylation modification cannot be catalyzed by all enzymes, which have substrate sequence preference. For example, EZH2 can mediate the monomethylation, dimethylation, and trimethylation of H3K27, while G9a (EHMT2) and MMSET (NSD2) mediate the monomethylation and dimethylation of H3K9 and H3K36, respectively, but not trimethylation [13].
Histone methylation modification was once considered to be an irreversible process, which is overturned by the discovery of LSD1 and KDM demethylase family with Jumonji C (JmjC) domain [14][15][16][14,15,16]. Unlike LSD1, KDMs are oxygenases that target multiple sites including H3K9, H3K27, and H3K36, and utilize 2-ketoglutarate (2-OG) and Fe2+ as cofactors to realize the demethylation of substrates through the mechanism of generating formaldehyde-free radicals [17]. The KDM demethylase JMJD2D (also known as KDM4D) belongs to JMJD2 subfamily that contains five members (JMJD2A-E) and can recognize the dimethyl and trimethylation of H3K9 and H3K36, as well as the trimethylation of H1.4K26 [9][18][9,18]. H3K9 and H1.4K26 trimethylations are related to transcriptional inhibition or heterochromatin formation, while H3K36 methylation is related to the activation of genes; and methylation modification at these sites is not only involved in gene transcription control but also related to DNA replication and repair [19][20][19,20]. Therefore, the dynamic transition of histone methylation regulated by the JMJD2 family may have a far-reaching impact on various cellular physiological activities, especially on tumorigenesis. The research on the function of JMJD2A, JMJD2B, and JMJD2C is relatively comprehensive, while that of JMJD2D is ongoing, but JMJD2E is unformed.
2. JMJD2D Promotes Gene Transcription by Antagonizing H3K9 Methylation
KDM4/JMJD2 histone demethylase family members are very similar in overall protein structure, which contains the featured JmjN and JmjC domain. JMJD2A, JMJD2B, and JMJD2C contain two plant homeodomains (PHD) and two tudor domains, while JMJD2D and JMJD2E are smaller and lack of the C-terminal PHD and tudor domains [9][18][21][9,18,21] (Figure 1A). Intriguingly, Shim et al. demonstrated that the truncation mutants of JMJD2A and JMJD2C that lack the tudor or both the tudor and PHD domains can still demethylate H3K9 and H3K36 [22]. Methylation of some lysine residues in H3 histones can activate or inactivate the transcriptional activity of genes, including H3K4, H3K9, H3K17, H3K27, H3K36, and H3K79 [23], while the dimethylation or trimethylation of H3K9 (H3K9me2/3) and H3K27 (H3K27me2/3) primarily associate with heterochromatin and gene repression [24][25][26][27][28][29][24,25,26,27,28,29]. The full-length JMJD2D, which localizes in the human chromosome 11q21 [18][30][18,30], demethylates the histone residues H3K9me2/3 and H1.4K26me3 to the monomethyl state, although H3K9me2/3 is the preferred substrate [9][31][9,31]. The JmjC domain of JMJD2D is the catalytic core, which facilitates a dioxygenase reaction requiring Fe2+, O2, and 2-oxoglutarate (2-OG) to demethylate histones. The oxygen atom is incorporated into the methyl group during oxidative decarboxylation; subsequently, an unstable intermediate imine product is formed and then converted to formaldehyde and demethylated product (Figure 1B) [32]. As expected, the JmjC-domain-lacking JMJD2 protein is deficient in demethylation activity [22]. The histidine 192 on the JmjC domain of JMJD2D is essential for its histone demethylase function and mutation of this residue severely compromises the catalytic activity of JMJD2D, leading to the loss of its ability to stimulate the mouse mammary tumor virus (MMTV) promoter [33]. The serine 200 on the JmjC domain is also essential for the histone demethylase function of JMJD2D and mutating this residue to methionine (S200M) generates a demethylase-dead mutant [34][35][34,35]. The JmjN domain of JMJD2D may be responsible for the integrity of structure and serves as a dimerization interface [9][18][21][36][9,18,21,36]. Deletion of the JmjN domain also abrogates the histone demethylase activity in the cell, due to the nuclear exclusion of JMJD2D [22].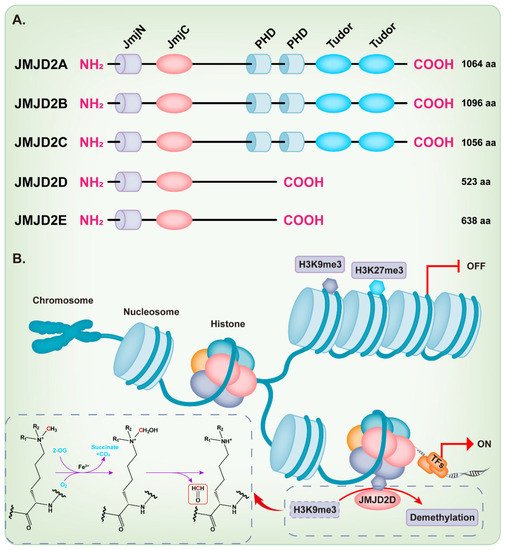
Figure 1. Structure and function of the JMJD2 histone demethylase subfamily. (A) The domain structure diagram of the JMJD2 histone demethylase subfamily. (B) JMJD2D facilitates a dioxygenase reaction requiring Fe2+, O2, and 2-oxoglutarate (2-OG) to demethylate H3K9me3.
3. JMJD2D Has Multiple Biological Functions
As mentioned earlier, JMJD2D promotes gene transcription by antagonizing H3K9 methylation, while it can serve as a coactivator to enhance the transcriptional activities of multiple transcription factors (TFs). JMJD2D is a well-established epigenetic regulator in a variety of biological functions, including androgen receptor (AR) activation, DNA damage repair, DNA replication, cell cycle regulation, inflammation modulation, and tumorigenesis promotion (Figure 2).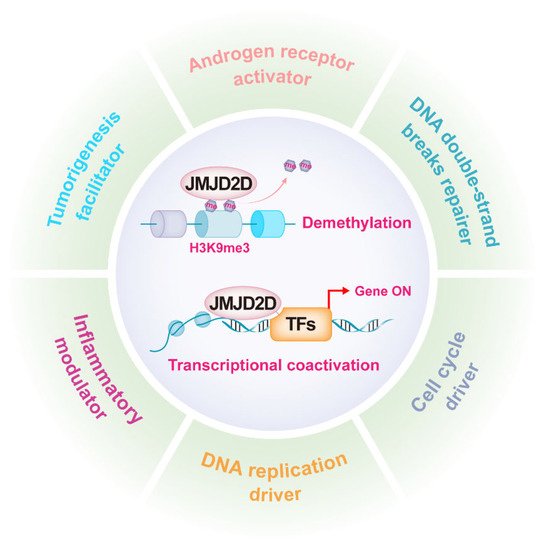
Figure 2. JMJD2D is a multifunctional epigenetic regulator. JMJD2D can promote gene transcription by antagonizing H3K9 methylation and cooperating with multiple transcription factors. JMJD2D has a variety of biological functions, including AR activation, DNA damage repair, DNA replication, cell cycle regulation, inflammation modulation, and tumorigenesis promotion.
