Bile-duct cancers (BDC) are a group of solid tumors arising from the biliary tree. Despite their classification as rare cancers, the incidence of BDC is increasing worldwide. Poor prognosis is a common feature of this type of cancer and is mainly determined by the following factors: late diagnosis, lack of effective therapeutic approaches, and resistance to conventional treatments. In the past few years, next-generation sequencing technologies has allowed us to study the genome, exome, and transcriptome of BDC deeper, revealing a previously underestimated class of RNA: the noncoding RNA (ncRNA). MicroRNAs (miRNAs) are small ncRNAs that play an important regulatory role in gene expression. The aberrant expression of miRNAs and their pivotal role as oncogenes or tumor suppressors in biliary carcinogenesis has been widely described in BDC. Due to their ability to regulate multiple gene networks, miRNAs are involved in all cancer hallmarks, including sustaining proliferative signaling, evading growth suppressors, resisting cell death, enabling replicative immortality, inducing/accessing vasculature, activating invasion and metastasis, reprogramming cellular metabolism, and avoiding immune destruction. Their use as diagnostic, prognostic, and predictive biomarkers has been widely explored in several human cancers, including BDC. Furthermore, miRNA-based therapeutic strategies are currently the subject of numerous clinical trials that are providing evidence of their efficacy as potent anticancer agents.
- non-coding RNA
- microRNA
- bile duct cancer
- precision medicine
1. Introduction
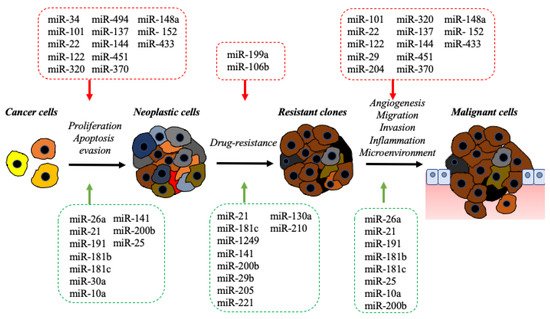
2. The Impact of the microRNA in Molecular Pathology of BDC
|
miRNA |
Target Gene |
Mechanism |
References |
||||
|---|---|---|---|---|---|---|---|
|
miR-26a |
GSK-3β; KRT19 |
Proliferation, migration, and invasion |
|||||
] | [ | ,33] |
miR-21 |
PTEN; PDCD4; TIMP3; PTPN14; 15-PGDH/HPGD |
|||
|
miR-101 |
EZH2, COX-2, APP, MCL-1, VEGF |
Proliferation, apoptosis, EMT, inflammation |
Proliferation, apoptosis; angiogenesis, inflammation; transcriptional repression |
||||
] | [ |
miR-191 |
TET |
Proliferation, invasion, and migration | |||
|
miR-22 | [23] |
||||||
SIRT1, CDK6, SP1, HDAC6 |
Proliferation, senescence, invasion, metastasis; ciliogenesis, histone modifications |
miR-181b-5p |
PARK2 |
Proliferation, migration, and invasion |
|||
|
miR-122 | [ | ||||||
ALDOA, CLIC1 |
Proliferation and invasion |
miR-181c |
NDRG2 |
Proliferation, drug-resistance, and metastasis |
[27] |
||
|
MiR-30a-5p |
SOCS3 |
Proliferation |
[28] |
||||
|
miR-25 |
DR4 |
Proliferation, invasion, and apoptosis |
|||||
|
miR-10a-5p |
PTEN |
Proliferation |
[31] |
|
miRNA |
Target Gene |
Mechanism |
References |
|---|---|---|---|
|
miR-34 |
MYC, MET, CDK4/6, BCL2, CD44, NOTCH1, NOTCH2, JAGGED1 |
Proliferation, apoptosis |
|
, | |||
, | ] |
||
|
miR-29-3p |
ITGA6, ITGB1 |
Cell migration and invasion |
[43] |
|
miR-204 |
SLUG |
Cell migration, invasion, EMT |
|
|
miR-320 |
VEGF, NRP-1 |
Proliferation, invasion, EMT, tumor migration, and metastasis |
|
|
miR-494 |
CCNB1, CDK2, CDK4, CDK6, CCND1, CCNE2, HDAC1, RB1, PLK1, PTTG1, TOP2A |
Proliferation, cell cycle |
[48] |
|
miR-137 |
WNT2B |
Proliferation, migration, and invasion |
[49] |
|
miR-144-5p/miR-451 |
ST8SIA4 |
Proliferation, migration, and invasion |
[50] |
|
miR-370 |
MAP3K8 |
Proliferation, inflammation, tumor microenvironment |
|
|
miR-148a |
RASSF1 |
Proliferation, inflammation, tumor microenvironment |
|
|
miR-152 |
CDKN2A |
Proliferation, inflammation, tumor microenvironment |
