Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is a comparison between Version 3 by Peter Tang and Version 2 by Daniela Ogonczyk-Makowska.
Human metapneumovirus (HMPV) is an important human pathogen that, along with respiratory syncytial virus (RSV), is a major cause of respiratory tract infections in young infants. Recombinant chimeric viruses expressing antigens of other viruses can be generated by reverse genetics and used for simultaneous immunization against more than one pathogen. This approach can result in the development of promising vaccine candidates against HMPV, and several studies have indeed validated viral vectors expressing HMPV antigens.
- human metapneumovirus
- respiratory syncytial virus
- chimeric vaccines
- recombinant vaccines
- viral vectors
- bivalent vaccines
1. Introduction
Acute respiratory tract infections (RTIs) are the 5th leading global cause of mortality among all age groups and the 3rd leading mortality cause in children younger than 5 years old [1]. An important number of RTIs is caused by viruses belonging to the Pneumoviridae family, namely, respiratory syncytial virus (RSV) and human metapneumovirus (HMPV) that are responsible for approximately 31% and 5.5% of RTIs in children, respectively [2].
HMPV is a single-stranded, negative-sense RNA virus belonging to the family of Pneumoviridae, genus Metapneumovirus. The genus has two species: human and avian metapneumovirus (AMPV). HMPVs are divided into two major genetic subtypes: A and B that are further divided into sublineages A1, A2a, A2b, B1, and B2 [3]. Lineages A and B share 80% and 90% of their nucleotide and amino acid sequence identity, correspondingly [4]. The major antigen of HMPV, the F protein, is highly conserved and a high level of cross-neutralization and cross-protection is observed between the two lineages [5,6][5][6]. The clinical signs of HMPV disease do not vary significantly between the different genetic lineages of HMPV [7]. AMPV, previously known as turkey rhinotracheitis virus, is an important poultry pathogen, divided into subgroups A, B, C, and D [8,9,10][8][9][10]. The subgroup C (AMPV_C) shares 80% of the amino acid identity of N, P, M, F, M2-1, and M2-2 proteins with HMPV [11], making it the closest related virus to HMPV.
HMPV is a relatively newly-discovered pathogen, described for the first time by the researchers from the Netherlands in 2001 [11]. Its late identification can be explained by the similarity of symptoms caused by HMPV and RSV, as well as the difficulty to observe HMPV growth in vitro. Both viruses cause pneumonia and bronchiolitis, with variable severity of illness according to age [12]. Although HMPV disease is generally less severe compared to RSV, the incidence of infections caused by HMPV is similar to that of influenza and parainfluenza, and it causes a significant number of hospitalizations every year [13]. Infections with either virus are associated with the development of asthma and its exacerbations, yet the causal relationship has not been established [2,14,15][2][14][15]. Although both pneumoviruses can infect any age group, they cause the greatest disease severity in young infants. HMPV infects mainly children between 6 and 12 months of age, whereas RSV infects infants earlier, within the first 2–3 months after the birth [2,16,17,18,19][2][16][17][18][19]. Virtually every child is infected by RSV by the age of 2 [20] and by HMPV by the age of 5 [21], and the incidence of reinfections with both viruses increases significantly after the age of 50 causing a lot of RTIs in elderly [19,22][19][22]. Another population at risk of HMPV infections is comprised of immunosuppressed patients [23].
1.1. Need for a Vaccine
Despite the high frequency of pneumoviral infections and over 50 years of research in this field, no licensed vaccine against HMPV or RSV is currently available. Among the numerous vaccine candidates against RSV that have been developed, only a few have advanced to clinical trials and most of them failed as a result of insufficient immunogenicity or underattenuation [24]. The only anti-HMPV vaccine advanced to clinical trials until now—a live-attenuated recombinant rHMPV-PA virus, in which HMPV-P protein was exchanged for its counterpart from AMPV_C—proved to be insufficiently immunogenic and overattenuated in seronegative children, thus leaving no advanced candidates for an anti-HMPV vaccine [25].
This lack of effective vaccine candidates against HMPV can be explained by the recent discovery of the virus, but also by the lack of a successful vaccine against closely related RSV that could serve as a base for vaccine design. One of the reasons for the slow progress in this field is the clinical failure of formalin-inactivated RSV-vaccine (FI-RSV) that occurred in the 1960s. The administration of FI-RSV led to the development of an exaggerated immune response to wild type (wt) RSV infection with enhanced pulmonary disease (EPD) and two vaccinated children died [26]. EPD has been further documented in various animal models, indicating that it is not only a human phenomenon [27,28,29,30][27][28][29][30]. A similar effect was observed for a formalin or heat-inactivated HMPV vaccine, which induced the symptoms of EPD in rodents [31,32][31][32]. Another reason for slow vaccine development is the transient immunity provided by natural infection with RSV and HMPV. As documented in RSV-seropositive adults, levels of RSV-neutralizing antibodies correlate with the resistance to subsequent RSV infection, but the protection they confer is incomplete and short-lasting, which results in frequent reinfections [20,33][20][33]. The immune response to primary HMPV infection was described to be weak and aberrant in BALB/c mice, with excessive Th2-cytokines production at later stages of infection, which correlated with airways hyperresponsiveness and the development of asthma [34]. Pneumoviruses have also developed many mechanisms to evade immune responses. The RSV nonstructural proteins NS1 and NS2 suppress the IFN-induced antiviral response in infected cells and removal of these proteins resulted in the induction of high levels of IFN alpha and beta in vitro [35,36,37][35][36][37]. HMPV shares with RSV the same ability to decrease IFN response, despite the lack of NS1 and NS2 proteins. It has been documented that HMPV-G protein has the ability to inhibit the IFN type 1 response in vitro and in vivo [38,39][38][39] and that its SH and M2-2 proteins can also modulate the host’s immune responses [40,41,42][40][41][42]. Both HMPV and RSV can persist in the lungs of an infected animal host, despite the presence of neutralizing antibodies [34,43][34][43]. Such persistence has been documented by RT-PCR detection of viral RNA in the guinea pig and mouse models for RSV [44,45,46][44][45][46] and in mice for HMPV [43]. Another hurdle to effective immunization is the presence of maternal antibodies in the bloodstream of vaccinated infants that can impair the immunogenic effect of the vaccine [47], yet the role of maternal antibodies in RSV infection is not fully understood [48].
1.2. Vaccine Development
The main objectives in pneumovirus vaccine development are to avoid EPD and obtain sufficient immunogenicity without causing disease. Viral protein antigens can be used as subunit vaccines to induce an appropriate immune response. These proteins can be delivered as nanoparticles, virus-like particles (VLPs), or they can be coupled with adjuvants [49]. Another interesting strategy is to immunize with viral nucleic acid coding for viral antigens, as it has been documented for RSV and HMPV vaccine candidates [50,51,52][50][51][52]. An mRNA-based vaccine coding for the F proteins of HMPV and human parainfluenza type 3 virus (PIV3) has been recently advanced to phase 1 clinical trial [53].
Among the three surface proteins of HMPV (F, G, and SH), the F protein constitutes the major HMPV antigen [6,54,55][6][54][55]. Although immunogenic, its G protein does not induce a potent protective immune response [54,56][54][56] and is not indispensable for viral replication in vivo [57]. HMPV-SH protein was shown not to confer any significant protection [54], the same being observed for RSV [58]. The F protein is also the major antigen of RSV, more immunogenic than the G protein [59], although the latter is also able to induce a protective immune response [60,61][60][61] and has been frequently tested as a subunit vaccine [62,63,64][62][63][64]. RSV-F protein stabilized in its pre-fusion form by introducing disulfide bond (DS) and cavity-filling (Cav1) mutations (DS-Cav1) is currently being tested as a subunit vaccine in phase I clinical trials (ClinicalTrials.gov Identifier: NCT03049488). The pre-fusion RSV-F was shown to be more immunogenic than its postfusion conformation, as a result of an exposition of a unique antigenic site Ø that is recognized by very potent RSV-neutralizing antibodies [65,66,67][65][66][67]. Trials with the HMPV-F subunit vaccine showed that the vaccine was immunogenic, but not protective in a rodent model [68]. To increase its immunogenic potential, HMPV-F protein has been stabilized in its pre-fusion state by analogous strategies as for RSV-F, but its immunogenicity was not enhanced [69,70][69][70]. This difference between pre-fusion RSV-F and HMPV-F can be explained by additional glycosylation at the apex of the pre-fusion form of HMPV-F. HMPV-F and RSV-F share some antigenic sites, and antibodies able to cross-neutralize the two pneumoviruses were identified [71,72,73,74,75][71][72][73][74][75]. Grafting of a major protective epitope from RSV-F into HMPV-F protein resulted in the elaboration of a chimeric protein carrying epitopes of both pneumoviruses, an interesting candidate for vaccine design [76,77][76][77].
Several promising nanoparticle vaccines against RSV have been elaborated [78[78][79],79], among which is ResVax, a nanoparticle RSV-F-based vaccine developed by Novavax [80]. ResVax has been recently tested in phase 3 clinical trials in maternal immunization model of lower RTIs prevention in infants (NCT02624947), where it did not meet its clinical endpoint [81]. VLPs are structures composed of the proteins of viral capsid; they do not contain the viral genome and are replication-incompetent. Coexpression of HMPV-F, -G, and -M proteins leads to the formation of VLPs in vitro [82], yet the expression of only F and M proteins has been shown sufficient for VLPs assembly [83]. Several potential VLPs vaccines against HMPV [82[82][83][84][85],83,84,85], and many more against RSV [78,86,87,88,89,90,91,92,93,94,95,96[78][86][87][88][89][90][91][92][93][94][95][96][97][98][99][100][101][102][103][104][105],97,98,99,100,101,102,103,104,105], were developed, but none of them has been advanced to clinical trials. A promising vaccine candidate—a VLP composed of pre-fusion, postfusion RSV-F, or both together, assembled with HMPV-M protein—conferred full protection against RSV infection in immunized mice [96].
On the other hand, immunization with an attenuated replication-competent virus is a very promising approach. In this case, it is possible to obtain a vaccine that presents all viral epitopes and is able to induce both humoral and cellular immune responses [106]. The main disadvantage of live attenuated vaccines is the risk of reversion of the attenuated profile, restoration of infectivity and subsequent development of the disease [107]. Live attenuated viruses can be divided into two groups: non-recombinant and recombinant or chimeric viruses. Non-recombinant viruses are rendered less infectious due to genetic modifications that appear naturally during serial passages in vitro in conditions of environmental stress. Several attenuated strains of RSV and HMPV have been obtained as a result of serial cold passages or chemical mutagenesis, which confer a temperature-sensitive (ts) phenotype to a virus [108,109,110,111,112][108][109][110][111][112]. Apart from the mentioned selection strategy, recombinant attenuated viruses can be generated by reverse genetics, a technique that makes it possible to generate functional particles of a modified virus based on its genetic material [113,114][113][114]. This approach allows not only to introduce attenuating mutations directly into the viral genome [57[57][115],115], but also to express exogenous antigens in the backbone of the virus, leading to the development of polyvalent, chimeric vaccines. The insertion of a foreign gene can have a potentially attenuating effect on its own, for example, by rearranging the order of genes in terms of 3’-5’ expression gradient, as it is observed in paramyxoviruses. Gene order can also be changed in a more invasive way by moving virulence genes from the high-expression position to the low-expression locus. Alternatively, the replication efficiency can be decreased by deletions or silencing of nonessential viral proteins that play an accessory function in the life cycle of the virus. It is also possible to swap some of the genes between strains of different host preferences, thus introducing new host range restrictions. HMPV virus with the P gene exchanged for its AMPV_C counterpart (rHMPV-PA) was more attenuated than wt HMPV, and yet protective against HMPV infection in African green monkeys (AGMs) [116].
1.3. Target Populations
The ultimate goal of HMPV vaccination is the prevention of lower RTIs in populations at risk. Similarly to RSV, the target populations for HMPV vaccination are young children, elderly people, and pregnant women [117]. Early vaccination of infants and young children can potentially prevent HMPV-infections and the transmission of the virus. Replication-competent vaccines, namely, live-attenuated HMPV or recombinant viruses are a good choice for this population. Another vaccine strategy, prime-boost regimen, consists of vaccinating with gene-based/live vaccine first and then with a protein/particle-based vaccine [118]. Immunization of pregnant women can not only provide a passive antibody transfer to their children but also prevent a mother-to-child transmission of HMPV. Live vaccines are considered being too risky to be used in this population; therefore, subunit vaccines or VLPs with standard adjuvants should be considered. The effect of vaccination of adult populations with live vaccines can be hampered by the presence of anti-HMPV antibodies in the bloodstream or in the respiratory tract (RT). For the vaccination of elderly people, who had experienced multiple HMPV infections, subunit or VLPs with a potent adjuvant are recommended [118].
2. Vector-Based Chimeric Vaccines
One of the strategies aimed to circumvent the problem of incomplete immunity provided by natural HMPV infection is to express its protective antigens in the backbone of a more immunogenic virus. The increase in immunogenicity can also be provided by improved antigen expression by the vector, as demonstrated for a recombinant bovine/human PIV3 (rB/HPIV3) expressing either RSV-F or G proteins [60] and for human parainfluenza type 1 virus (HPIV1) expressing HMPV antigens [54]. HMPV is difficult to grow in cell culture; vectoring its antigens with the backbone of a better-replicating virus can facilitate vaccine development and manufacturing. For instance, rB/HPIV3 expressing RSV-F reaches 10–100-fold higher viral titers in vivo compared to wt RSV [60] and replicates more efficiently in the RT of AGMs than wt RSV or HMPV [119,120][119][120]. Expression of a foreign antigen in the backbone of another virus can also mitigate the problem of pre-existing immunity that often diminishes the effect of the vaccine. Using vectors of another host range can facilitate the immunization of seropositive individuals with no risk of causing disease, as it has been described for Newcastle Disease Virus (NDV) expressing RSV-F. The majority of live vaccines against RSV and HMPV up to date have been attenuated by a few codon changes in their genome. Therefore, using more stably attenuated backbone can render the attenuation more reliable, and decrease the risk of the reversion of an attenuated phenotype. The insertion of an additional gene can itself influence the virus’ replicative capacity, thus limiting its ability to spread and cause the disease, but also to create the risk of overattenuation. This subtle balance between attenuation and immunogenicity remains the major challenge of vaccine development.
In general, vector-based chimeric vaccines retain the advantages of live attenuated vaccines and they can be readily generated by reverse genetics. Most of the backbones used so far for vectoring pneumoviral antigens belong to the Paramyxoviridae family. These viruses are closely related, both phylogenetically and structurally, to pneumoviruses. This close relationship increases the chance of an efficient expression and integration of a pneumoviral protein in the background of a paramyxoviral vector. Among paramyxoviruses, the most widely used vaccine backbones are bovine parainfluenza virus type 3 (BPIV3) and its recombinant derivative rB/HPIV3, HPIV1, parainfluenza virus type 5 (PIV5), Newcastle Disease Virus (NDV), and Sendai virus (SeV). The reasons explaining this frequent use of paramyxoviruses as vaccine backbones are numerous: First, their genomes are well-characterized and complete genomic sequences of all known members of this family are easily accessible [121]. Second, their genomes are simple and organized in a modular way. Tandem alignment of the genes, of which the most are transcribed as separate mRNA products, facilitates genetic manipulations [122]. Third, most paramyxoviruses are able to replicate efficiently in cell lines certified for vaccine manufacture, such as Vero cells, which can facilitate vaccine development [122]. Fourth, as they replicate entirely in the cytoplasm and their replication cycle does not involve the integration into the host cell genome, their use is potentially safe [121]. Also, recombination events between mononegaviruses have not been observed in nature and they remain extremely rare in vitro, even in optimized conditions [123[123][124],124], indicating that there is a low probability of gene exchange between engineered vaccine viruses and the pathogens present in the environment. Fifth, the small probability of recombination also contributes to the stability of a genetic insert, providing a stable foreign gene expression system. Paramyxoviruses are able to stably express several exogenes simultaneously and the level of expression can be manipulated by changing the position of gene insertion [125]. Sixth, most paramyxoviruses infect their host via their RT, representing an easy and safe route of administration of the vaccine as well as for induction of both local and systemic immune responses. Last, but not least, there are many animal paramyxoviruses that are naturally attenuated in humans due to a host range restriction. Mutations that render these viruses harmless to people have been identified and characterized, thus making attenuation of different paramyxoviruses possible [122].
3. Recombinant Virus Engineering
3.1. Reverse Genetics
Negative strand viruses can be readily recovered from cell cultures by means of reverse genetics [126]. This technique makes it possible to engineer a fully functional virus starting from its genetic sequence. The genetic material in the form of cDNA can be easily modified according to the vaccine design. The technique is based on transfecting permissive cells with a plasmid coding for the viral genome and satellite plasmids coding for all the proteins necessary for the formation of a ribonucleoprotein complex (RNP) that initiates the transcription of viral genes (Figure 1). The proteins indispensable to form the RNP in paramyxoviruses and pneumoviruses are N, P, and L proteins [127[127][128][129],128,129], yet the addition of M2-1 protein facilitates the recovery HMPV from cDNA [130]. The elaboration of a polyvalent vaccine can be accomplished by either cloning additional genes into the plasmid coding for viral genome or replacing protective antigens of a vector with the ones of another pathogen.
Figure 1. Schematic representation of reverse genetics pipeline. The exogene of interest is cloned into a plasmid containing a complete genome of a vector virus and then transfected, along with satellite plasmids coding for viral proteins indispensable to initiate viral assembly, into a cell line designed for initial virus production. The first progeny of the recombinant virus is harvested and propagated on a permissive cell line.
3.2. Genomic Organization of Paramyxoviridae and Pneumoviridae
The genomes of Paramyxoviridae and Pneumoviridae consist of a simple, nonsegmented, linear, single-stranded, 15,000–19,000-nucleotide-long negative RNA that contains 6–10 genes [121]. The most 3’-proximal region consists of a leader (le)—a 50-nucleotide-long promoter—and each gene is preceded and followed by a short (10–13 nucleotides) conserved sequence named gene start (GS) and gene end (GE). These sequences act like transcription control signals for viral RNA-dependent RNA-polymerase (vRNAP) and they guide the enzyme along the genome [131]. Single genes are separated from each other by short, non-coding intergenic regions and the order of the genes is usually conserved as 3’-Nucleoprotein (N), Phosphoprotein (P), Matrix Protein (M), Glycoprotein (G), Hemagglutinin-Neuraminidase (HN), or Large Polymerase subunit (L)–5’ with the presence and location of additional genes depending on the virus [132] (Figure 2).

Figure 2. Genetic organization of Pneumoviruses and some of Paramyxoviruses used as viral vectors. Pneumoviridae: RSV: Respiratory Syncytial Virus (Orthopneumovirus), HMPV: Human Metapneumovirus (Metapneumovirus). Paramyxoviridae: PIV5: Parainfluenza type 5 virus (Rubulavirus), HPIV3: Human Parainfluenza type 3 virus, BPIV3: Bovine Parainfluenza type 3 virus, HPIV1: Human Parainfluenza type 1 virus and SeV: Sendai Virus (Respirovirus), NDV: Newcastle Disease Virus and APMV3: Avian Paramyxovirus type 3 (Avulavirus). Genes: le: leader, NS1 and NS2: accessory proteins of RSV, N: nucleoprotein, P: phosphoprotein, M: matrix protein, F: fusion protein, SH: small hydrophobic protein, G: attachment glycoprotein, HN: Haemagglutinin-Neuraminidase protein, M2: gene coding for M2-1 and M2-2 proteins, L: large polymerase subunit.
The 5’-end of a genome contains a trailer sequence (tr) of a variable length, ranging from 50 up to 707 nucleotides [121,133][121][133]. The genomic RNA of paramyxoviruses and pneumoviruses does not exist as an unbound RNA-particle: it is always assembled with numerous copies of N protein and forms a helicoidal nucleocapsid. In paramyxoviruses, each N protein molecule is associated with precisely six nucleotides, a feature that is believed to underlay the ‘’rule of six’’, i.e., the length of the genome of paramyxoviruses has to be a multiple of six for an effective viral replication [134,135][134][135]. The stringent adhesion to this rule is observed for SeV [136]; for other viruses, like PIV5, NDV, and HPIV3, it strongly increases the efficacy of replication [137,138,139][137][138][139]. The fact that it does not give any replicative advantage to RSV [140] might be explained by distinct differences in nucleocapsid structure of the two families [141]. Transcription of the viral genome is initiated at the 3’-end and a 3’-to 5’ expression gradient is observed. The complex of vRNAP sporadically fails to resume the synthesis of another distinct mRNA at each gene junction, which results in a gradual loss of transcription-efficacy along the genome [121].
3.3. Principles of Exogene Insertions
The main principles to be taken into consideration while engineering the genome of recombinant paramyxoviruses are the preferential adherence to the rule of six, a coherence of transcription control signals (GS/GE) used to drive the expression of a foreign antigen and exogene positioning in the transcription gradient. As it has been documented in numerous studies, the GS and GE signals of the vector virus can efficiently direct the expression of an exogenous protein [60]. Although GS and GE signals are often highly conserved along the viral genome, there might be variations in their sequences and transcription efficacy [142]. Flanking the sequence of GFP inserted into the 6th genome position of PIV5 with GS/GE specific for either 2nd or 7th gene junctions resulted in large differences in GFP expression levels (Figure 3) [143]. GS/GE characteristics for the 1st junction provided better expression levels than the ones originating from the 7th. It is therefore important to flank the exogene with potent transcription regulators. In accordance with the 3’-5’ transcription gradient, exogenes placed in more 3’-proximal position should be expressed better than 5’-proximal inserts, yet the tendency cannot be described as linear and some deviations are observed [144]. The adherence to this gradient might also be influenced by the type of attenuation of vector virus, as it has been demonstrated for HPIV1 bearing RSV-F protein [145]. The positions of an exogene in the viral genome are described in Figure 3 and the nomenclature used to label chimeric viruses in this work is explained in Figure 4.
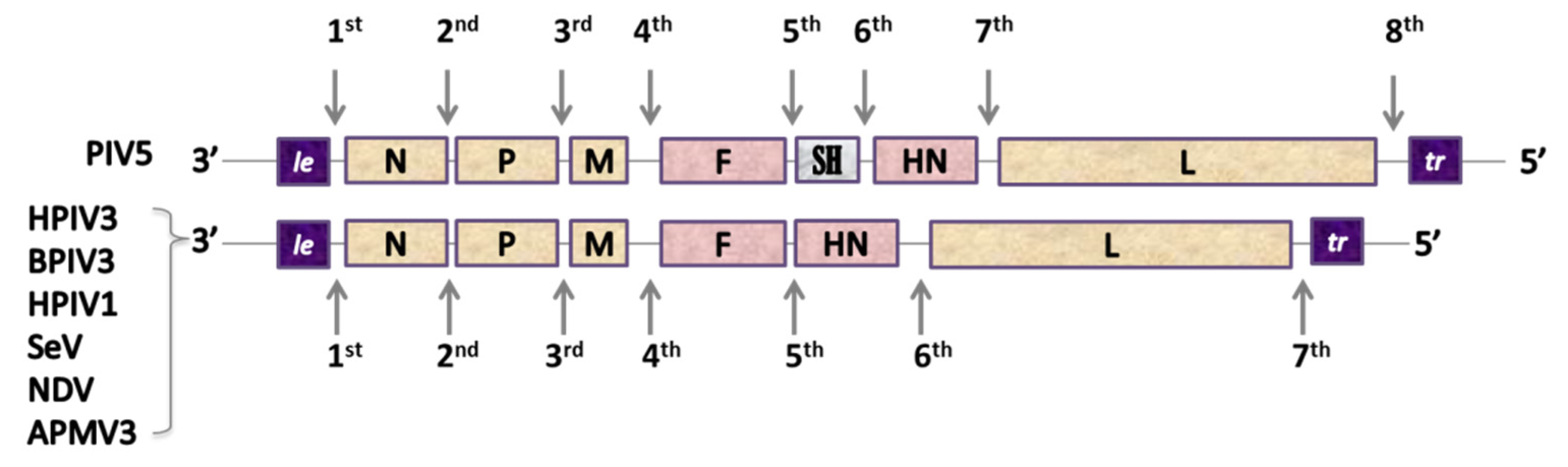
Figure 3.
Insert-positions in the paramyxovirus genomes.

Figure 4. The nomenclature used in this work to describe chimeric viruses expressing additional antigens. The inserts are marked according to their 3’-5’ order of the vector’s genome. The subtype of the virus at the origin of the insert is marked if it is specified in the source. Although discussing different chimeric viruses based on the same vector that are mentioned in the same study, recombinant viruses can be also referred to as F1st, G2nd, etc. constructs. The nomenclature of vector backbones modified by additional mutations, protein swapping, etc. (i.e., rHPIV1-CΔ170, rHMPV-PA) was unchanged in relation to the source publication.
On one hand, a 3’-proximal insertion should provide the best level of expression; on the other hand, it can influence the level of transcription of all downstream genes, leading to a decrease in viral replication. RSV-F insertion into 1st position of rB/HPIV3 genome reduced the expression of downstream genes by 20–45% [146]; 2nd genomic position can provide a good expression, but it can also influence the N:P protein ratio, which plays a decisive role in the replicative capacity of paramyxoviruses, causing an additional attenuating effect [146]. Insertions in either 1st or 2nd genomic positions are the most privileged for PIV3-based vectors, with the 2nd position usually providing better virus recovery, higher viral replication, and better exogene expression [147]. For some vectors, namely, NDV and Avian paramyxovirus serotype 3 (APMV3), the 3rd position is the most advantageous, whereas the insertion at the 2nd one results in delayed viral replication and the largest reduction in virus recovery [148,149,150][148][149][150]. The 4th and the 5th positions are not frequently used, so as to not influence the expression of vector’s surface proteins (F and HN) [146], with the exception of studies on SeV bearing either RSV-F or HMPV-F at the 5th position [151,152,153][151][152][153].
The nature of the insert itself can also influence the vector’s biology. The rB/HPIV3//RSV-F1st virus showed an 8-fold reduced replication in vitro compared to rB/HPIV3, whereas the RSV-G1st insert did not influence viral replication [60]. This decrease in viral growth might have been due to excessive syncytia formation and increased cytopathology resulting from the expression of a second fusion protein. Another reason might have been the size of the insert–a bigger F protein might have influenced the replication more significantly than a smaller G protein. The size of an exogene can significantly change the replicative capacity of the virus, as it has been shown for HPIV3 vector bearing inserts of different sizes [154]. The level of integration of a foreign protein into a vector’s particle can also be a price to pay in the exchange for efficient expression of the exogene. Improved exogene integration into the vector’s backbone can be obtained by the substitution of the transmembrane domain (TM) and cytoplasmic tail (CT) of the inserts with their equivalents from the vector virus. As demonstrated for HPIV1 vector bearing RSV-F protein either in its wt or chimeric TMCT form, packaging of RSV-F(TMCT) was strongly improved, but the chimeric virus was overattenuated and not protective in hamsters [155].
References
- GBD 2015 LRI Collaborators Estimates of the global, regional, and national morbidity, mortality, and aetiologies of lower respiratory tract infections in 195 countries: A systematic analysis for the Global Burden of Disease Study 2015. Lancet Infect. Dis. 2017, 17, 1133–1161.
- García-García, M.L.; Calvo, C.; Rey, C.; Díaz, B.; Molinero, M.D.; Pozo, F.; Casas, I. Human metapnuemovirus infections in hospitalized children and comparison with other respiratory viruses. 2005-2014 prospective study. PLoS ONE 2017, 12, e0173504.
- Huck, B.; Scharf, G.; Neumann-Haefelin, D.; Puppe, W.; Weigl, J.; Falcone, V. Novel Human Metapneumovirus Sublineage. Emerg. Infect. Dis. 2006, 12, 147–150.
- Biacchesi, S.; Skiadopoulos, M.H.; Boivin, G.; Hanson, C.T.; Murphy, B.R.; Collins, P.L.; Buchholz, U.J. Genetic diversity between human metapneumovirus subgroups. Virology 2003, 315, 1–9.
- van den Hoogen, B.G.; Herfst, S.; Sprong, L.; Cane, P.A.; Forleo-Neto, E.; de Swart, R.L.; Osterhaus, A.D.M.E.; Fouchier, R.A.M. Antigenic and Genetic Variability of Human Metapneumoviruses. Emerg. Infect. Dis. 2004, 10, 658–666.
- Skiadopoulos, M.H.; Biacchesi, S.; Buchholz, U.J.; Riggs, J.M.; Surman, S.R.; Amaro-Carambot, E.; McAuliffe, J.M.; Elkins, W.R.; St. Claire, M.; Collins, P.L.; et al. The Two Major Human Metapneumovirus Genetic Lineages Are Highly Related Antigenically, and the Fusion (F) Protein Is a Major Contributor to This Antigenic Relatedness. J. Virol. 2004, 78, 6927–6937.
- Wei, H.-Y.; Tsao, K.-C.; Huang, C.-G.; Huang, Y.-C.; Lin, T.-Y. Clinical features of different genotypes/genogroups of human metapneumovirus in hospitalized children. J. Microbiol. Immunol. Infect. 2013, 46, 352–357.
- Juhasz, K.; Easton, A.J. Extensive Sequence Variation in the Attachment (G) Protein Gene of Avian Pneumovirus: Evidence for Two Distinct Subgroups. J. Gen. Virol. 1994, 75, 2873–2880.
- Seal, B.S. Matrix protein gene nucleotide and predicted amino acid sequence demonstrate that the first US avian pneumovirus isolate is distinct from European strains. Virus Res. 1998, 58, 45–52.
- Arnauld, C.; Bäyon-Auboyer, M.-H.; Eterradossi, N.; Toquin, D. Nucleotide sequences of the F, L and G protein genes of two non-A/non-B avian pneumoviruses (APV) reveal a novel APV subgroup. J. Gen. Virol. 2000, 81, 2723–2733.
- van den Hoogen, B.G.; de Jong, J.C.; Groen, J.; Kuiken, T.; de Groot, R.; Fouchier, R.A.M.; Osterhaus, A.D.M.E. A newly discovered human pneumovirus isolated from young children with respiratory tract disease. Nat. Med. 2001, 7, 719–724.
- Moe, N.; Krokstad, S.; Stenseng, I.H.; Christensen, A.; Skanke, L.H.; Risnes, K.R.; Nordbø, S.A.; Døllner, H. Comparing Human Metapneumovirus and Respiratory Syncytial Virus: Viral Co-Detections, Genotypes and Risk Factors for Severe Disease. PLoS ONE 2017, 12, e0170200.
- Williams, J.V.; Edwards, K.M.; Weinberg, G.A.; Griffin, M.R.; Hall, C.B.; Zhu, Y.; Szilagyi, P.G.; Wang, C.K.; Yang, C.; Silva, D.; et al. Population-Based Incidence of Human Metapneumovirus Infection among Hospitalized Children. J. Infect. Dis. 2010, 201, 1890–1898.
- Peiris, J.S.M.; Tang, W.-H.; Chan, K.-H.; Khong, P.-L.; Guan, Y.; Lau, Y.-L.; Chiu, S.S. Children with Respiratory Disease Associated with Metapneumovirus in Hong Kong. Emerg. Infect. Dis. 2003, 9, 628–633.
- Kusel, M.M.H.; de Klerk, N.H.; Kebadze, T.; Vohma, V.; Holt, P.G.; Johnston, S.L.; Sly, P.D. Early-life respiratory viral infections, atopic sensitization, and risk of subsequent development of persistent asthma. J. Allergy Clin. Immunol. 2007, 119, 1105–1110.
- Edwards, K.M.; Zhu, Y.; Griffin, M.R.; Weinberg, G.A.; Hall, C.B.; Szilagyi, P.G.; Staat, M.A.; Iwane, M.; Prill, M.M.; Williams, J.V. Burden of Human Metapneumovirus Infection in Young Children. N. Engl. J. Med. 2013, 368, 633–643.
- Bosis, S.; Esposito, S.; Niesters, H.G.M.; Crovari, P.; Osterhaus, A.D.M.E.; Principi, N. Impact of human metapneumovirus in childhood: Comparison with respiratory syncytial virus and influenza viruses. J. Med. Virol. 2005, 75, 101–104.
- Anderson, E.J.; Simões, E.A.F.; Buttery, J.P.; Dennehy, P.H.; Domachowske, J.B.; Jensen, K.; Lieberman, J.M.; Losonsky, G.A.; Yogev, R. Prevalence and Characteristics of Human Metapneumovirus Infection Among Hospitalized Children at High Risk for Severe Lower Respiratory Tract Infection. J. Pediatric Infect. Dis. Soc. 2012, 1, 212–222.
- Cattoir, L.; Vankeerberghen, A.; Boel, A.; Van Vaerenbergh, K.; De Beenhouwer, H. Epidemiology of RSV and hMPV in Belgium: A 10-year follow-up. Acta Clin. Belg. 2019, 74, 229–235.
- Glezen, W.P.; Taber, L.H.; Frank, A.L.; Kasel, J.A. Risk of primary infection and reinfection with respiratory syncytial virus. Am. J. Dis. Child. 1986, 140, 543–546.
- Leung, J.; Esper, F.; Weibel, C.; Kahn, J.S. Seroepidemiology of human metapneumovirus (hMPV) on the basis of a novel enzyme-linked immunosorbent assay utilizing hMPV fusion protein expressed in recombinant vesicular stomatitis virus. J. Clin. Microbiol. 2005, 43, 1213–1219.
- Falsey, A.R.; Erdman, D.; Anderson, L.J.; Walsh, E.E. Human Metapneumovirus Infections in Young and Elderly Adults. J. Infect. Dis. 2003, 187, 785–790.
- Englund, J.A.; Boeckh, M.; Kuypers, J.; Nichols, W.G.; Hackman, R.C.; Morrow, R.A.; Fredricks, D.N.; Corey, L. Brief Communication: Fatal Human Metapneumovirus Infection in Stem-Cell Transplant Recipients. Ann. Intern. Med. 2006, 144, 344.
- Mazur, N.I.; Higgins, D.; Nunes, M.C.; Melero, J.A.; Langedijk, A.C.; Horsley, N.; Buchholz, U.J.; Openshaw, P.J.; McLellan, J.S.; Englund, J.A.; et al. The respiratory syncytial virus vaccine landscape: Lessons from the graveyard and promising candidates. Lancet Infect. Dis. 2018, 18, e295–e311.
- Karron, R.A.; San Mateo, J.; Wanionek, K.; Collins, P.L.; Buchholz, U.J. Evaluation of a Live Attenuated Human Metapneumovirus Vaccine in Adults and Children. J. Pediatric Infect. Dis. Soc. 2018, 7, 86–89.
- Murphy, B.R.; Prince, G.A.; Walsh, E.E.; Kim, H.W.; Parrott, R.H.; Hemming, V.G.; Rodriguez, W.J.; Chanock, R.M. Dissociation between serum neutralizing and glycoprotein antibody responses of infants and children who received inactivated respiratory syncytial virus vaccine. J. Clin. Microbiol. 1986, 24, 197–202.
- Waris, M.E.; Tsou, C.; Erdman, D.D.; Zaki, S.R.; Anderson, L.J. Respiratory synctial virus infection in BALB/c mice previously immunized with formalin-inactivated virus induces enhanced pulmonary inflammatory response with a predominant Th2-like cytokine pattern. J. Virol. 1996, 70, 2852–2860.
- Kakuk, T.J.; Soike, K.; Brideau, R.J.; Zaya, R.M.; Cole, S.L.; Zhang, J.-Y.; Roberts, E.D.; Wells, P.A.; Wathen, M.W. A Human Respiratory Syncytial Virus (RSV) Primate Model of Enhanced Pulmonary Pathology Induced with a Formalin-Inactivated RSV Vaccine but Not a Recombinant FG Subunit Vaccine. J. Infect. Dis. 1993, 167, 553–561.
- Porter, D.D.; Prince, G.A.; Yim, K.C.; Curtis, S.J. Vaccine-enhanced respiratory syncytial virus disease in cotton rats following immunization with Lot 100 or a newly prepared reference vaccine. J. Gen. Virol. 2001, 82, 2881–2888.
- Gershwin, L.J.; Schelegle, E.S.; Gunther, R.A.; Anderson, M.L.; Woolums, A.R.; Larochelle, D.R.; Boyle, G.A.; Friebertshauser, K.E.; Singer, R.S. A bovine model of vaccine enhanced respiratory syncytial virus pathophysiology. Vaccine 1998, 16, 1225–1236.
- Yim, K.C.; Cragin, R.P.; Boukhvalova, M.S.; Blanco, J.C.G.; Hamlin, M.-È.; Boivin, G.; Porter, D.D.; Prince, G.A. Human metapneumovirus: Enhanced pulmonary disease in cotton rats immunized with formalin-inactivated virus vaccine and challenged. Vaccine 2007, 25, 5034–5040.
- Hamelin, M.-E.; Couture, C.; Sackett, M.K.; Boivin, G. Enhanced lung disease and Th2 response following human metapneumovirus infection in mice immunized with the inactivated virus. J. Gen. Virol. 2007, 88, 3391–3400.
- Hall, C.B.; Walsh, E.E.; Long, C.E.; Schnabel, K.C. Immunity to and frequency of reinfection with respiratory syncytial virus. J. Infect. Dis. 1991, 163, 693–698.
- Alvarez, R.; Tripp, R.A. The Immune Response to Human Metapneumovirus Is Associated with Aberrant Immunity and Impaired Virus Clearance in BALB/c Mice. J. Virol. 2005, 79, 5971–5978.
- Valarcher, J.-F.; Furze, J.; Wyld, S.; Cook, R.; Conzelmann, K.-K.; Taylor, G. Role of alpha/beta interferons in the attenuation and immunogenicity of recombinant bovine respiratory syncytial viruses lacking NS proteins. J. Virol. 2003, 77, 8426–8439.
- Schlender, J.; Bossert, B.; Buchholz, U.; Conzelmann, K.K. Bovine respiratory syncytial virus nonstructural proteins NS1 and NS2 cooperatively antagonize alpha/beta interferon-induced antiviral response. J. Virol. 2000, 74, 8234–8242.
- Spann, K.M.; Tran, K.-C.; Chi, B.; Rabin, R.L.; Collins, P.L. Suppression of the induction of alpha, beta, and lambda interferons by the NS1 and NS2 proteins of human respiratory syncytial virus in human epithelial cells and macrophages . J. Virol. 2004, 78, 4363–4369.
- Bao, X.; Liu, T.; Shan, Y.; Li, K.; Garofalo, R.P.; Casola, A. Human metapneumovirus glycoprotein G inhibits innate immune responses. Plos Pathog. 2008, 4, e1000077.
- Cheemarla, N.R.; Guerrero-Plata, A. Human Metapneumovirus Attachment Protein Contributes to Neutrophil Recruitment into the Airways of Infected Mice. Viruses 2017, 9, 310.
- Bao, X.; Kolli, D.; Liu, T.; Shan, Y.; Garofalo, R.P.; Casola, A. Human Metapneumovirus Small Hydrophobic Protein Inhibits NF- B Transcriptional Activity. J. Virol. 2008, 82, 8224–8229.
- Ren, J.; Wang, Q.; Kolli, D.; Prusak, D.J.; Tseng, C.-T.K.; Chen, Z.J.; Li, K.; Wood, T.G.; Bao, X. Human Metapneumovirus M2-2 Protein Inhibits Innate Cellular Signaling by Targeting MAVS. J. Virol. 2012, 86, 13049–13061.
- Ren, J.; Liu, G.; Go, J.; Kolli, D.; Zhang, G.; Bao, X. Human Metapneumovirus M2-2 Protein Inhibits Innate Immune Response in Monocyte-Derived Dendritic Cells. PLoS ONE 2014, 9, e91865.
- Alvarez, R.; Harrod, K.S.; Shieh, W.-J.; Zaki, S.; Tripp, R.A. Human Metapneumovirus Persists in BALB/c Mice despite the Presence of Neutralizing Antibodies. J. Virol. 2004, 78, 14003–14011.
- Mejías, A.; Chávez-Bueno, S.; Gómez, A.M.; Somers, C.; Estripeaut, D.; Torres, J.P.; Jafri, H.S.; Ramilo, O. Respiratory Syncytial Virus Persistence: Evidence in the Mouse Model. Pediatric Infect. Dis. J. 2008, 27, S60–S62.
- Schwarze, J.; O’Donnell, D.R.; Rohwedder, A.; Openshaw, P.J.M. Latency and Persistence of Respiratory Syncytial Virus Despite T Cell Immunity. Am. J. Respir. Crit. Care Med. 2004, 169, 801–805.
- Dakhama, A.; Vitalis, T.Z.; Hegel, R.G. Persistence of respiratory syncytial virus (RSV) infection and development of RSV-specific IgG1 response in a guinea-pig model of acute brochiolitis. Eur. Respir. J. 1997, 10, 20–26.
- Englund, J.A. Passive protection against respiratory syncytial virus disease in infants: The role of maternal antibody. Pediatric Infect. Dis. J. 1994, 13, 449–453.
- van Erp, E.A.; van Kasteren, P.B.; Guichelaar, T.; Ahout, I.M.L.; de Haan, C.A.M.; Luytjes, W.; Ferwerda, G.; Wicht, O. In Vitro Enhancement of Respiratory Syncytial Virus Infection by Maternal Antibodies Does Not Explain Disease Severity in Infants. J. Virol. 2017, 91, e00851-17.
- Patton, K.; Aslam, S.; Shambaugh, C.; Lin, R.; Heeke, D.; Frantz, C.; Zuo, F.; Esser, M.T.; Paliard, X.; Lambert, S.L. Enhanced immunogenicity of a respiratory syncytial virus (RSV) F subunit vaccine formulated with the adjuvant GLA-SE in cynomolgus macaques. Vaccine 2015, 33, 4472–4478.
- Cseke, G.; Wright, D.W.; Tollefson, S.J.; Johnson, J.E.; Crowe, J.E.; Williams, J.V. Human Metapneumovirus Fusion Protein Vaccines That Are Immunogenic and Protective in Cotton Rats. J. Virol. 2007, 81, 698–707.
- Smith, T.R.F.; Schultheis, K.; Morrow, M.P.; Kraynyak, K.A.; McCoy, J.R.; Yim, K.C.; Muthumani, K.; Humeau, L.; Weiner, D.B.; Sardesai, N.Y.; et al. Development of an intradermal DNA vaccine delivery strategy to achieve single-dose immunity against respiratory syncytial virus. Vaccine 2017, 35, 2840–2847.
- Ma, Y.; Jiao, Y.-Y.; Yu, Y.-Z.; Jiang, N.; Hua, Y.; Zhang, X.-J.; Fu, Y.-H.; Peng, X.-L.; Zheng, Y.-P.; Anderson, L.; et al. A Built-In CpG Adjuvant in RSV F Protein DNA Vaccine Drives a Th1 Polarized and Enhanced Protective Immune Response. Viruses 2018, 10, 38.
- Shaw, C.; Lee, H.; Knightly, C.; Kalidindi, S.; Zaks, T.; Smolenov, I.; Panther, L. 2754. Phase 1 Trial of an mRNA-Based Combination Vaccine Against hMPV and PIV3. Open Forum Infect. Dis. 2019, 6, S970.
- Skiadopoulos, M.H.; Biacchesi, S.; Buchholz, U.J.; Amaro-Carambot, E.; Surman, S.R.; Collins, P.L.; Murphy, B.R. Individual contributions of the human metapneumovirus F, G, and SH surface glycoproteins to the induction of neutralizing antibodies and protective immunity. Virology 2006, 345, 492–501.
- Melero, J.A.; Mas, V. The Pneumovirinae fusion (F) protein: A common target for vaccines and antivirals. Virus Res. 2015, 209, 128–135.
- Ryder, A.B.; Tollefson, S.J.; Podsiad, A.B.; Johnson, J.E.; Williams, J.V. Soluble recombinant human metapneumovirus G protein is immunogenic but not protective. Vaccine 2010, 28, 4145–4152.
- Biacchesi, S.; Skiadopoulos, M.H.; Yang, L.; Lamirande, E.W.; Tran, K.C.; Murphy, B.R.; Collins, P.L.; Buchholz, U.J. Recombinant Human Metapneumovirus Lacking the Small Hydrophobic SH and/or Attachment G Glycoprotein: Deletion of G Yields a Promising Vaccine Candidate. J. Virol. 2004, 78, 12877–12887.
- Connors, M.; Collins, P.L.; Firestone, C.Y.; Murphy, B.R. Respiratory syncytial virus (RSV) F, G, M2 (22K), and N proteins each induce resistance to RSV challenge, but resistance induced by M2 and N proteins is relatively short-lived. J. Virol. 1991, 65, 1634–1637.
- Olmsted, R.A.; Elango, N.; Prince, G.A.; Murphy, B.R.; Johnson, P.R.; Moss, B.; Chanock, R.M.; Collins, P.L. Expression of the F glycoprotein of respiratory syncytial virus by a recombinant vaccinia virus: Comparison of the individual contributions of the F and G glycoproteins to host immunity. Proc. Natl. Acad. Sci. USA 1986, 83, 7462–7466.
- Schmidt, A.C.; McAuliffe, J.M.; Murphy, B.R.; Collins, P.L. Recombinant Bovine/Human Parainfluenza Virus Type 3 (B/HPIV3) Expressing the Respiratory Syncytial Virus (RSV) G and F Proteins Can Be Used to Achieve Simultaneous Mucosal Immunization against RSV and HPIV3. J. Virol. 2001, 75, 4594–4603.
- Schmidt, A.C.; Wenzke, D.R.; McAuliffe, J.M.; St Claire, M.; Elkins, W.R.; Murphy, B.R.; Collins, P.L. Mucosal Immunization of Rhesus Monkeys against Respiratory Syncytial Virus Subgroups A and B and Human Parainfluenza Virus Type 3 by Using a Live cDNA-Derived Vaccine Based on a Host Range-Attenuated Bovine Parainfluenza Virus Type 3 Vector Backbone. J. Virol. 2002, 76, 1089–1099.
- Li, C.; Zhou, X.; Zhong, Y.; Li, C.; Dong, A.; He, Z.; Zhang, S.; Wang, B. A Recombinant G Protein Plus Cyclosporine A–Based Respiratory Syncytial Virus Vaccine Elicits Humoral and Regulatory T Cell Responses against Infection without Vaccine-Enhanced Disease. J. Immunol. 2016, 196, 1721–1731.
- Fuentes, S.; Klenow, L.; Golding, H.; Khurana, S. Preclinical evaluation of bacterially produced RSV-G protein vaccine: Strong protection against RSV challenge in cotton rat model. Sci. Rep. 2017, 7, 1–13.
- Zhang, S.; Zhao, G.; Su, C.; Li, C.; Zhou, X.; Zhao, W.; Zhong, Y.; He, Z.; Peng, H.; Dong, A.; et al. Neonatal priming and infancy boosting with a novel respiratory syncytial virus vaccine induces protective immune responses without concomitant respiratory disease upon RSV challenge. Hum. Vaccines Immunother. 2019, 1–9.
- McLellan, J.S.; Chen, M.; Joyce, M.G.; Sastry, M.; Stewart-Jones, G.B.E.; Yang, Y.; Zhang, B.; Chen, L.; Srivatsan, S.; Zheng, A.; et al. Structure-Based Design of a Fusion Glycoprotein Vaccine for Respiratory Syncytial Virus. Science 2013, 342, 592–598.
- Liang, B.; Surman, S.; Amaro-Carambot, E.; Kabatova, B.; Mackow, N.; Lingemann, M.; Yang, L.; McLellan, J.S.; Graham, B.S.; Kwong, P.D.; et al. Enhanced Neutralizing Antibody Response Induced by Respiratory Syncytial Virus Prefusion F Protein Expressed by a Vaccine Candidate. J. Virol. 2015, 89, 9499–9510.
- Liang, B.; Ngwuta, J.O.; Herbert, R.; Swerczek, J.; Dorward, D.W.; Amaro-Carambot, E.; Mackow, N.; Kabatova, B.; Lingemann, M.; Surman, S.; et al. Packaging and Prefusion Stabilization Separately and Additively Increase the Quantity and Quality of Respiratory Syncytial Virus (RSV)-Neutralizing Antibodies Induced by an RSV Fusion Protein Expressed by a Parainfluenza Virus Vector. J. Virol. 2016, 90, 10022–10038.
- Herfst, S.; Schrauwen, E.J.A.; de Graaf, M.; van Amerongen, G.; van den Hoogen, B.G.; de Swart, R.L.; Osterhaus, A.D.M.E.; Fouchier, R.A.M. Immunogenicity and efficacy of two candidate human metapneumovirus vaccines in cynomolgus macaques. Vaccine 2008, 26, 4224–4230.
- Battles, M.B.; Más, V.; Olmedillas, E.; Cano, O.; Vázquez, M.; Rodríguez, L.; Melero, J.A.; McLellan, J.S. Structure and immunogenicity of pre-fusion-stabilized human metapneumovirus F glycoprotein. Nat. Commun. 2017, 8, 1–11.
- Pilaev, M.; Shen, Y.; Carbonneau, J.; Venable, M.-C.; Rhéaume, C.; Lavigne, S.; Couture, C.; Guarné, A.; Hamelin, M.-È.; Boivin, G. Evaluation of pre- and post-fusion Human metapneumovirus F proteins as subunit vaccine candidates in mice. Vaccine 2020, in press.
- Corti, D.; Bianchi, S.; Vanzetta, F.; Minola, A.; Perez, L.; Agatic, G.; Guarino, B.; Silacci, C.; Marcandalli, J.; Marsland, B.J.; et al. Cross-neutralization of four paramyxoviruses by a human monoclonal antibody. Nature 2013, 501, 439–443.
- Schuster, J.E.; Cox, R.G.; Hastings, A.K.; Boyd, K.L.; Wadia, J.; Chen, Z.; Burton, D.R.; Williamson, R.A.; Williams, J.V. A Broadly Neutralizing Human Monoclonal Antibody Exhibits In Vivo Efficacy Against Both Human Metapneumovirus and Respiratory Syncytial Virus. J. Infect. Dis. 2015, 211, 216–225.
- Wen, X.; Mousa, J.J.; Bates, J.T.; Lamb, R.A.; Crowe, J.E.; Jardetzky, T.S. Structural basis for antibody cross-neutralization of respiratory syncytial virus and human metapneumovirus. Nat. Microbiol. 2017, 2, 1–7.
- Xiao, X.; Tang, A.; Cox, K.S.; Wen, Z.; Callahan, C.; Sullivan, N.L.; Nahas, D.D.; Cosmi, S.; Galli, J.D.; Minnier, M.; et al. Characterization of potent RSV neutralizing antibodies isolated from human memory B cells and identification of diverse RSV/hMPV cross-neutralizing epitopes. mAbs 2019, 11, 1415–1427.
- Mousa, J.J.; Binshtein, E.; Human, S.; Fong, R.H.; Alvarado, G.; Doranz, B.J.; Moore, M.L.; Ohi, M.D.; Crowe, J.E. Human antibody recognition of antigenic site IV on Pneumovirus fusion proteins. PLoS Pathog. 2018, 14, e1006837.
- Wen, X.; Pickens, J.; Mousa, J.J.; Leser, G.P.; Lamb, R.A.; Crowe, J.E.; Jardetzky, T.S. A Chimeric Pneumovirus Fusion Protein Carrying Neutralizing Epitopes of Both MPV and RSV. PLoS ONE 2016, 11, e0155917.
- Olmedillas, E.; Cano, O.; Martínez, I.; Luque, D.; Terrón, M.C.; McLellan, J.S.; Melero, J.A.; Más, V. Chimeric Pneumoviridae fusion proteins as immunogens to induce cross-neutralizing antibody responses. Embo. Mol. Med. 2018, 10, 175–187.
- Smith, G.; Raghunandan, R.; Wu, Y.; Liu, Y.; Massare, M.; Nathan, M.; Zhou, B.; Lu, H.; Boddapati, S.; Li, J.; et al. Respiratory Syncytial Virus Fusion Glycoprotein Expressed in Insect Cells Form Protein Nanoparticles That Induce Protective Immunity in Cotton Rats. PLoS ONE 2012, 7, e50852.
- Glenn, G.M.; Fries, L.F.; Thomas, D.N.; Smith, G.; Kpamegan, E.; Lu, H.; Flyer, D.; Jani, D.; Hickman, S.P.; Piedra, P.A. A Randomized, Blinded, Controlled, Dose-Ranging Study of a Respiratory Syncytial Virus Recombinant Fusion (F) Nanoparticle Vaccine in Healthy Women of Childbearing Age. J. Infect. Dis. 2016, 213, 411–422.
- Patel, N.; Massare, M.J.; Tian, J.-H.; Guebre-Xabier, M.; Lu, H.; Zhou, H.; Maynard, E.; Scott, D.; Ellingsworth, L.; Glenn, G.; et al. Respiratory syncytial virus prefusogenic fusion (F) protein nanoparticle vaccine: Structure, antigenic profile, immunogenicity, and protection. Vaccine 2019, 37, 6112–6124.
- Novavax Announces Topline Results from Phase 3 PrepareTM Trial of ResVaxTM for Prevention of RSV Disease in Infants via Maternal Immunization. 2019. Available online: https://www.nasdaq.com/press-release/novavax-announces-topline-results-phase-3-preparetm-trial-resvaxtm-prevention-rsv (accessed on 2 November 2019).
- Loo, L.; Jumat, M.; Fu, Y.; Ayi, T.; Wong, P.; Tee, N.W.; Tan, B.; Sugrue, R.J. Evidence for the interaction of the human metapneumovirus G and F proteins during virus-like particle formation. Virol. J. 2013, 10, 294.
- Wen, S.C.; Schuster, J.E.; Gilchuk, P.; Boyd, K.L.; Joyce, S.; Williams, J.V. Lung CD8 + T Cell Impairment Occurs during Human Metapneumovirus Infection despite Virus-Like Particle Induction of Functional CD8 + T Cells. J. Virol. 2015, 89, 8713–8726.
- Lévy, C.; Aerts, L.; Hamelin, M.-È.; Granier, C.; Szécsi, J.; Lavillette, D.; Boivin, G.; Cosset, F.-L. Virus-like particle vaccine induces cross-protection against human metapneumovirus infections in mice. Vaccine 2013, 31, 2778–2785.
- Cox, R.G.; Erickson, J.J.; Hastings, A.K.; Becker, J.C.; Johnson, M.; Craven, R.E.; Tollefson, S.J.; Boyd, K.L.; Williams, J.V. Human Metapneumovirus Virus-Like Particles Induce Protective B and T Cell Responses in a Mouse Model. J. Virol. 2014, 88, 6368–6379.
- McGinnes, L.W.; Gravel, K.A.; Finberg, R.W.; Kurt-Jones, E.A.; Massare, M.J.; Smith, G.; Schmidt, M.R.; Morrison, T.G. Assembly and Immunological Properties of Newcastle Disease Virus-Like Particles Containing the Respiratory Syncytial Virus F and G Proteins. J. Virol. 2011, 85, 366–377.
- Murawski, M.R.; McGinnes, L.W.; Finberg, R.W.; Kurt-Jones, E.A.; Massare, M.J.; Smith, G.; Heaton, P.M.; Fraire, A.E.; Morrison, T.G. Newcastle Disease Virus-Like Particles Containing Respiratory Syncytial Virus G Protein Induced Protection in BALB/c Mice, with No Evidence of Immunopathology. J. Virol. 2010, 84, 1110–1123.
- Quan, F.-S.; Kim, Y.; Lee, S.; Yi, H.; Kang, S.-M.; Bozja, J.; Moore, M.L.; Compans, R.W. Viruslike Particle Vaccine Induces Protection Against Respiratory Syncytial Virus Infection in Mice. J. Infect. Dis. 2011, 204, 987–995.
- Hwang, H.S.; Kwon, Y.-M.; Lee, J.S.; Yoo, S.-E.; Lee, Y.-N.; Ko, E.-J.; Kim, M.-C.; Cho, M.-K.; Lee, Y.-T.; Jung, Y.-J.; et al. Co-immunization with virus-like particle and DNA vaccines induces protection against respiratory syncytial virus infection and bronchiolitis. Antivir. Res. 2014, 110, 115–123.
- Lee, S.; Quan, F.-S.; Kwon, Y.; Sakamoto, K.; Kang, S.-M.; Compans, R.W.; Moore, M.L. Additive protection induced by mixed virus-like particles presenting respiratory syncytial virus fusion or attachment glycoproteins. Antivir. Res. 2014, 111, 129–135.
- Raghunandan, R.; Lu, H.; Zhou, B.; Xabier, M.G.; Massare, M.J.; Flyer, D.C.; Fries, L.F.; Smith, G.E.; Glenn, G.M. An insect cell derived respiratory syncytial virus (RSV) F nanoparticle vaccine induces antigenic site II antibodies and protects against RSV challenge in cotton rats by active and passive immunization. Vaccine 2014, 32, 6485–6492.
- Ko, E.-J.; Kwon, Y.-M.; Lee, J.S.; Hwang, H.S.; Yoo, S.-E.; Lee, Y.-N.; Lee, Y.-T.; Kim, M.-C.; Cho, M.K.; Lee, Y.R.; et al. Virus-like nanoparticle and DNA vaccination confers protection against respiratory syncytial virus by modulating innate and adaptive immune cells. Nanomed. Nanotechnol. Biol. Med. 2015, 11, 99–108.
- McGinnes Cullen, L.; Schmidt, M.R.; Kenward, S.A.; Woodland, R.T.; Morrison, T.G. Murine Immune Responses to Virus-Like Particle-Associated Pre- and Postfusion Forms of the Respiratory Syncytial Virus F Protein. J. Virol. 2015, 89, 6835–6847.
- Walpita, P.; Johns, L.M.; Tandon, R.; Moore, M.L. Mammalian Cell-Derived Respiratory Syncytial Virus-Like Particles Protect the Lower as well as the Upper Respiratory Tract. PLoS ONE 2015, 10, e0130755.
- Kim, K.-H.; Lee, Y.-T.; Hwang, H.S.; Kwon, Y.-M.; Kim, M.-C.; Ko, E.-J.; Lee, J.S.; Lee, Y.; Kang, S.-M. Virus-Like Particle Vaccine Containing the F Protein of Respiratory Syncytial Virus Confers Protection without Pulmonary Disease by Modulating Specific Subsets of Dendritic Cells and Effector T Cells. J. Virol. 2015, 89, 11692–11705.
- Cimica, V.; Boigard, H.; Bhatia, B.; Fallon, J.T.; Alimova, A.; Gottlieb, P.; Galarza, J.M. Novel Respiratory Syncytial Virus-Like Particle Vaccine Composed of the Postfusion and Prefusion Conformations of the F Glycoprotein. Clin. Vaccine Immunol. 2016, 23, 451–459.
- Hwang, H.S.; Lee, Y.-T.; Kim, K.-H.; Park, S.; Kwon, Y.-M.; Lee, Y.; Ko, E.-J.; Jung, Y.-J.; Lee, J.S.; Kim, Y.-J.; et al. Combined virus-like particle and fusion protein-encoding DNA vaccination of cotton rats induces protection against respiratory syncytial virus without causing vaccine-enhanced disease. Virology 2016, 494, 215–224.
- Hwang, H.S.; Lee, Y.-T.; Kim, K.-H.; Ko, E.-J.; Lee, Y.; Kwon, Y.-M.; Kang, S.-M. Virus-like particle vaccine primes immune responses preventing inactivated-virus vaccine-enhanced disease against respiratory syncytial virus. Virology 2017, 511, 142–151.
- Hwang, H.S.; Kim, K.-H.; Lee, Y.; Lee, Y.-T.; Ko, E.-J.; Park, S.; Lee, J.S.; Lee, B.; Kwon, Y.-M.; Moore, M.L.; et al. Virus-like particle vaccines containing F or F and G proteins confer protection against respiratory syncytial virus without pulmonary inflammation in cotton rats. Hum. Vaccines Immunother. 2017, 13, 1031–1039.
- Kim, A.-R.; Lee, D.-H.; Lee, S.-H.; Rubino, I.; Choi, H.-J.; Quan, F.-S. Protection induced by virus-like particle vaccine containing tandem repeat gene of respiratory syncytial virus G protein. PLoS ONE 2018, 13, e0191277.
- Lee, Y.; Lee, Y.-T.; Ko, E.-J.; Kim, K.-H.; Hwang, H.S.; Park, S.; Kwon, Y.-M.; Kang, S.M. Soluble F proteins exacerbate pulmonary histopathology after vaccination upon respiratory syncytial virus challenge but not when presented on virus-like particles. Hum. Vaccines Immunother. 2017, 13, 2594–2605.
- Blanco, J.C.G.; Pletneva, L.M.; McGinnes-Cullen, L.; Otoa, R.O.; Patel, M.C.; Fernando, L.R.; Boukhvalova, M.S.; Morrison, T.G. Efficacy of a respiratory syncytial virus vaccine candidate in a maternal immunization model. Nat. Commun. 2018, 9, 1–10.
- Gilbert, B.E.; Patel, N.; Lu, H.; Liu, Y.; Guebre-Xabier, M.; Piedra, P.A.; Glenn, G.; Ellingsworth, L.; Smith, G. Respiratory syncytial virus fusion nanoparticle vaccine immune responses target multiple neutralizing epitopes that contribute to protection against wild-type and palivizumab-resistant mutant virus challenge. Vaccine 2018, 36, 8069–8078.
- McGinnes Cullen, L.; Schmidt, M.R.; Morrison, T.G. Effect of Previous Respiratory Syncytial Virus Infection on Murine Immune Responses to F and G Protein-Containing Virus-Like Particles. J. Virol. 2019, 93, e00087-19.
- Cullen, L.; Schmidt, M.; Torres, G.; Capoferri, A.; Morrison, T. Comparison of Immune Responses to Different Versions of VLP Associated Stabilized RSV Pre-Fusion F Protein. Vaccines 2019, 7, 21.
- Buchholz, U.J.; Nagashima, K.; Murphy, B.R.; Collins, P.L. Live vaccines for human metapneumovirus designed by reverse genetics. Expert Rev. Vaccines 2006, 5, 695–706.
- Bull, J.J. Evolutionary reversion of live viral vaccines: Can genetic engineering subdue it? Virus Evol. 2015, 1, vev005.
- Randolph, V.B.; Kandis, M.; Stemler-Higgins, P.; Kennelly, M.S.; McMullen, Y.M.; Speelman, D.J.; Weeks-Levy, C. Attenuated temperature-sensitive respiratory syncytial virus mutants generated by cold adaptation. Virus Res. 1994, 33, 241–259.
- Crowe, J. Cold-passaged, temperature-sensitive mutants of human respiratory syncytial virus (RSV) are highly attenuated, immunogenic, and protective in seronegative chimpanzees, even when RSV antibodies are infused shortly before immunization. Vaccine 1995, 13, 847–855.
- Juhasz, K.; Whitehead, S.S.; Bui, P.T.; Biggs, J.M.; Crowe, J.E.; Boulanger, C.A.; Collins, P.L.; Murphy, B.R. The temperature-sensitive (ts) phenotype of a cold-passaged (cp) live attenuated respiratory syncytial virus vaccine candidate, designated cpts530, results from a single amino acid substitution in the L protein. J. Virol. 1997, 71, 5814–5819.
- Whitehead, S.S.; Juhasz, K.; Firestone, C.Y.; Collins, P.L.; Murphy, B.R. Recombinant respiratory syncytial virus (RSV) bearing a set of mutations from cold-passaged RSV is attenuated in chimpanzees. J. Virol. 1998, 72, 4467–4471.
- Herfst, S.; de Graaf, M.; Schrauwen, E.J.A.; Sprong, L.; Hussain, K.; van den Hoogen, B.G.; Osterhaus, A.D.M.E.; Fouchier, R.A.M. Generation of temperature-sensitive human metapneumovirus strains that provide protective immunity in hamsters. J. Gen. Virol. 2008, 89, 1553–1562.
- Herfst, S.; de Graaf, M.; Schickli, J.H.; Tang, R.S.; Kaur, J.; Yang, C.-F.; Spaete, R.R.; Haller, A.A.; van den Hoogen, B.G.; Osterhaus, A.D.M.E.; et al. Recovery of Human Metapneumovirus Genetic Lineages A and B from Cloned cDNA. J. Virol. 2004, 78, 8264–8270.
- Hu, B.; Jiang, J.; Zhan, J.; Li, G.; Jiang, Y.; Guan, X.; Chen, Y.; Fang, Z. Development of a reverse genetics system for respiratory syncytial virus long strain and an immunogenicity study of the recombinant virus. Virol. J. 2014, 11, 142.
- Karron, R.A.; Luongo, C.; Thumar, B.; Loehr, K.M.; Englund, J.A.; Collins, P.L.; Buchholz, U.J. A gene deletion that up-regulates viral gene expression yields an attenuated RSV vaccine with improved antibody responses in children. Sci. Transl. Med. 2015, 7, 312ra175.
- Pham, Q.N.; Biacchesi, S.; Skiadopoulos, M.H.; Murphy, B.R.; Collins, P.L.; Buchholz, U.J. Chimeric Recombinant Human Metapneumoviruses with the Nucleoprotein or Phosphoprotein Open Reading Frame Replaced by That of Avian Metapneumovirus Exhibit Improved Growth In Vitro and Attenuation In Vivo. J. Virol. 2005, 79, 15114–15122.
- Phan, T.; Ren, J.; Bao, X. Recent vaccine development for human metapneumovirus. J. Gen. Virol. 2015, 96, 1515–1520.
- Anderson, L.J.; Dormitzer, P.R.; Nokes, D.J.; Rappuoli, R.; Roca, A.; Graham, B.S. Strategic priorities for respiratory syncytial virus (RSV) vaccine development. Vaccine 2013, 31, B209–B215.
- Tang, R.S.; MacPhail, M.; Schickli, J.H.; Kaur, J.; Robinson, C.L.; Lawlor, H.A.; Guzzetta, J.M.; Spaete, R.R.; Haller, A.A. Parainfluenza Virus Type 3 Expressing the Native or Soluble Fusion (F) Protein of Respiratory Syncytial Virus (RSV) Confers Protection from RSV Infection in African Green Monkeys. J. Virol. 2004, 78, 11198–11207.
- Tang, R.S.; Mahmood, K.; Macphail, M.; Guzzetta, J.M.; Haller, A.A.; Liu, H.; Kaur, J.; Lawlor, H.A.; Stillman, E.A.; Schickli, J.H.; et al. A host-range restricted parainfluenza virus type 3 (PIV3) expressing the human metapneumovirus (hMPV) fusion protein elicits protective immunity in African green monkeys. Vaccine 2005, 23, 1657–1667.
- Fields, B.N.; Knipe, D.M.; Howley, P.M. Fields Virology; Wolters Kluwer/Lippincott Williams & Wilkins Health: Philadelphia, PA, USA, 2013; ISBN 9781451105636.
- Bukreyev, A.; Skiadopoulos, M.H.; Murphy, B.R.; Collins, P.L. Nonsegmented Negative-Strand Viruses as Vaccine Vectors. J. Virol. 2006, 80, 10293–10306.
- Spann, K.M.; Collins, P.L.; Teng, M.N. Genetic recombination during coinfection of two mutants of human respiratory syncytial virus. J. Virol. 2003, 77, 11201–11211.
- Chare, E.R.; Gould, E.A.; Holmes, E.C. Phylogenetic analysis reveals a low rate of homologous recombination in negative-sense RNA viruses. J. Gen. Virol. 2003, 84, 2691–2703.
- Skiadopoulos, M.H.; Surman, S.R.; Riggs, J.M.; Örvell, C.; Collins, P.L.; Murphy, B.R. Evaluation of the Replication and Immunogenicity of Recombinant Human Parainfluenza Virus Type 3 Vectors Expressing up to Three Foreign Glycoproteins. Virology 2002, 297, 136–152.
- Kawaoka, Y. Biology of Negative Strand RNA Viruses: The Power of Reverse Genetics; Springer Science & Business Media: Berlin, Germany, 2013; ISBN 978-3-662-06099-5.
- Beaty, S.M.; Park, A.; Won, S.T.; Hong, P.; Lyons, M.; Vigant, F.; Freiberg, A.N.; tenOever, B.R.; Duprex, W.P.; Lee, B. Efficient and Robust Paramyxoviridae Reverse Genetics Systems. mSphere 2017, 2, e00376-16.
- Yu, Q.; Hardy, R.W.; Wertz, G.W. Functional cDNA clones of the human respiratory syncytial (RS) virus N, P, and L proteins support replication of RS virus genomic RNA analogs and define minimal trans-acting requirements for RNA replication. J. Virol. 1995, 69, 2412–2419.
- Grosfeld, H.; Hill, M.G.; Collins, P.L. RNA replication by respiratory syncytial virus (RSV) is directed by the N, P, and L proteins; transcription also occurs under these conditions but requires RSV superinfection for efficient synthesis of full-length mRNA. J. Virol. 1995, 69, 5677–5686.
- Biacchesi, S.; Skiadopoulos, M.H.; Tran, K.C.; Murphy, B.R.; Collins, P.L.; Buchholz, U.J. Recovery of human metapneumovirus from cDNA: Optimization of growth in vitro and expression of additional genes. Virology 2004, 321, 247–259.
- Noton, S.L.; Fearns, R. Initiation and regulation of paramyxovirus transcription and replication. Virology 2015, 479–480, 545–554.
- Ruigrok, R.W.; Crépin, T.; Kolakofsky, D. Nucleoproteins and nucleocapsids of negative-strand RNA viruses. Curr. Opin. Microbiol. 2011, 14, 504–510.
- Kumar, S.; Nayak, B.; Collins, P.L.; Samal, S.K. Complete genome sequence of avian paramyxovirus type 3 reveals an unusually long trailer region. Virus Res. 2008, 137, 189–197.
- Kolakofsky, D.; Pelet, T.; Garcin, D.; Hausmann, S.; Curran, J.; Roux, L. Paramyxovirus RNA Synthesis and the Requirement for Hexamer Genome Length: The Rule of Six Revisited. J. Virol. 1998, 72, 891–899.
- Alayyoubi, M.; Leser, G.P.; Kors, C.A.; Lamb, R.A. Structure of the paramyxovirus parainfluenza virus 5 nucleoprotein–RNA complex. Proc. Natl. Acad. Sci. USA 2015, 112, E1792–E1799.
- Calain, P.; Roux, L. The rule of six, a basic feature for efficient replication of Sendai virus defective interfering RNA. J. Virol. 1993, 67, 4822–4830.
- Durbin, A.P.; Siew, J.W.; Murphy, B.R.; Collins, P.L. Minimum protein requirements for transcription and RNA replication of a minigenome of human parainfluenza virus type 3 and evaluation of the rule of six. Virology 1997, 234, 74–83.
- Murphy, S.K.; Parks, G.D. Genome nucleotide lengths that are divisible by six are not essential but enhance replication of defective interfering RNAs of the paramyxovirus simian virus 5. Virology 1997, 232, 145–157.
- Marcos, F.; Ferreira, L.; Cros, J.; Park, M.-S.; Nakaya, T.; García-Sastre, A.; Villar, E. Mapping of the RNA promoter of Newcastle disease virus. Virology 2005, 331, 396–406.
- Samal, S.K.; Collins, P.L. RNA Replication by a Respiratory Syncytial Virus RNA Analog Does Not Obey the Rule of Six and Retains a Nonviral Trinucleotide Extension at the Leader End. J. Virol. 1996, 70, 8.
- Bhella, D.; Ralph, A.; Murphy, L.B.; Yeo, R.P. Significant differences in nucleocapsid morphology within the Paramyxoviridae. J. Gen. Virol. 2002, 83, 1831–1839.
- Kato, A.; Kiyotani, K.; Hasan, M.K.; Shioda, T.; Sakai, Y.; Yoshida, T.; Nagai, Y. Sendai virus gene start signals are not equivalent in reinitiation capacity: Moderation at the fusion protein gene. J. Virol. 1999, 73, 9237–9246.
- He, B.; Paterson, R.G.; Ward, C.D.; Lamb, R.A. Recovery of infectious SV5 from cloned DNA and expression of a foreign gene. Virology 1997, 237, 249–260.
- Wignall-Fleming, E.B.; Hughes, D.J.; Vattipally, S.; Modha, S.; Goodbourn, S.; Davison, A.J.; Randall, R.E. Analysis of Paramyxovirus Transcription and Replication by High-Throughput Sequencing. J. Virol. 2019, 93.
- Mackow, N.; Amaro-Carambot, E.; Liang, B.; Surman, S.; Lingemann, M.; Yang, L.; Collins, P.L.; Munir, S. Attenuated Human Parainfluenza Virus Type 1 (HPIV1) Expressing the Fusion Glycoprotein of Human Respiratory Syncytial Virus (RSV) as a Bivalent HPIV1/RSV Vaccine. J. Virol 2015, 89, 10319–10332.
- Liang, B.; Munir, S.; Amaro-Carambot, E.; Surman, S.; Mackow, N.; Yang, L.; Buchholz, U.J.; Collins, P.L.; Schaap-Nutt, A. Chimeric Bovine/Human Parainfluenza Virus Type 3 Expressing Respiratory Syncytial Virus (RSV) F Glycoprotein: Effect of Insert Position on Expression, Replication, Immunogenicity, Stability, and Protection against RSV Infection. J. Virol. 2014, 88, 4237–4250.
- Tang, R.S.; Schickli, J.H.; MacPhail, M.; Fernandes, F.; Bicha, L.; Spaete, J.; Fouchier, R.A.M.; Osterhaus, A.D.M.E.; Spaete, R.; Haller, A.A. Effects of Human Metapneumovirus and Respiratory Syncytial Virus Antigen Insertion in Two 3′ Proximal Genome Positions of Bovine/Human Parainfluenza Virus Type 3 on Virus Replication and Immunogenicity. J. Virol. 2003, 77, 10819–10828.
- Zhao, H. Recombinant Newcastle disease virus as a viral vector: Effect of genomic location of foreign gene on gene expression and virus replication. J. Gen. Virol. 2003, 84, 781–788.
- Carnero, E.; Li, W.; Borderia, A.V.; Moltedo, B.; Moran, T.; Garcia-Sastre, A. Optimization of Human Immunodeficiency Virus Gag Expression by Newcastle Disease Virus Vectors for the Induction of Potent Immune Responses. J. Virol. 2009, 83, 584–597.
- Yoshida, A.; Samal, S.K. Avian Paramyxovirus Type-3 as a Vaccine Vector: Identification of a Genome Location for High Level Expression of a Foreign Gene. Front. Microbiol. 2017, 8, 693.
- Jones, B.G.; Sealy, R.E.; Rudraraju, R.; Traina-Dorge, V.L.; Finneyfrock, B.; Cook, A.; Takimoto, T.; Portner, A.; Hurwitz, J.L. Sendai virus-based RSV vaccine protects African green monkeys from RSV infection. Vaccine 2012, 30, 959–968.
- Zhan, X.; Hurwitz, J.L.; Krishnamurthy, S.; Takimoto, T.; Boyd, K.; Scroggs, R.A.; Surman, S.; Portner, A.; Slobod, K.S. Respiratory syncytial virus (RSV) fusion protein expressed by recombinant Sendai virus elicits B-cell and T-cell responses in cotton rats and confers protection against RSV subtypes A and B. Vaccine 2007, 25, 8782–8793.
- Russell, C.J.; Jones, B.G.; Sealy, R.E.; Surman, S.L.; Mason, J.N.; Hayden, R.T.; Tripp, R.A.; Takimoto, T.; Hurwitz, J.L. A Sendai virus recombinant vaccine expressing a gene for truncated human metapneumovirus (hMPV) fusion protein protects cotton rats from hMPV challenge. Virology 2017, 509, 60–66.
- Skiadopoulos, M.H.; Surman, S.R.; Durbin, A.P.; Collins, P.L.; Murphy, B.R. Long Nucleotide Insertions between the HN and L Protein Coding Regions of Human Parainfluenza Virus Type 3 Yield Viruses With Temperature-Sensitive and Attenuation Phenotypes. Virology 2000, 272, 225–234.
- Liu, X.; Liang, B.; Ngwuta, J.; Liu, X.; Surman, S.; Lingemann, M.; Kwong, P.D.; Graham, B.S.; Collins, P.L.; Munir, S. Attenuated Human Parainfluenza Virus Type 1 Expressing the Respiratory Syncytial Virus (RSV) Fusion (F) Glycoprotein from an Added Gene: Effects of Prefusion Stabilization and Packaging of RSV F. J. Virol. 2017, 91, e01101-17.
More
