The Y chromosome is one of the sex chromosomes found in males of animals of different taxa, including insects and mammals. Among all chromosomes, the Y chromosome is characterized by a unique chromatin landscape undergoing dynamic evolutionary change. Being entirely heterochromatic, the Y chromosome as a rule preserves few functional genes, but is enriched in tandem repeats and transposons. Due to difficulties in the assembly of the highly repetitive Y chromosome sequence, deep analyses of Y chromosome evolution, structure, and functions are limited to a few species, one of them being Drosophila melanogaster. Here we survey comparative evolutionary history of the fly and human Y chromosomes, peculiarities of transcription of giant genes, such as genes, encoding fertility factors in Drosophila, differential expression of sex-linked rDNA loci, and functions of Y-linked piRNA clusters ensuring sex-specific piRNA silencing. Our comparative analysis will provide further insight into the properties of the Y chromosomes in both insects and mammals.
- Drosophila
- Y chromosome
- piRNA pathway
- rDNA
- intron gigantism
- azoospermia
- transposable elements
1. Introduction
2. Comparative Evolutionary History of the Fly and Human Y Chromosomes
2.1. Y Chromosome Differentiation and Functions in Flies
The Y chromosome is a sex chromosome found in males of different groups of animals, including mammals and Diptera. Whereas in mammals the development of an organism according to the male type is determined by the presence of a functional Y chromosome, in Drosophila, sex is determined by the ratio of the number of X chromosomes to the number of the autosomes: normally, the presence of two X chromosomes triggers development according to the female type, and one—according to the male type [8][9]. Thus, individuals with the XXY genotype are female in flies and male in mammals, while X0 are female in mammals and male in flies. In Drosophila with the X0 karyotype, there are no severe structural or functional body disorders, except for male sterility [6]. Fly Y chromosome is not involved in sex determination. Several genes functioning in sex determination of flies, such as Sex-lethal (Sxl), transformer (tra), transformer-2 (tra2), and others, have been found to date. Sex-specific mRNA splicing of major Drosophila sex-determining genes is a complex process providing female-specific transcripts triggering female-type development [8][9]. In Diptera, the Y chromosomes arose from the autosomes repeatedly (Figure 2a), which provides good material for studying parallel processes of convergent evolutionary development [10][11][12]. While the gene content of the autosomes in Diptera is conserved and originates from a common ancestor, gene composition of the Y chromosomes varies significantly, and the rate of acquisition of new genes often significantly exceeds the rate of their loss [13]. In Diptera, the transfer of genes from other chromosomes contributes to the evolution of the Y chromosome. Relatively young, newly formed Y chromosomes of flies, being formed from tens of thousands of years to a few million years ago, maintain the structure and genes of the ancestral autosome, while most of the genes on the old fly Y chromosomes have been acquired subsequently due to a transfer from the autosomes or the X chromosome. Old Y chromosomes such as in D. melanogaster that presumably have been persisted for long time (several decades of millions of years) are often highly heterochromatic, contain a large amount of repetitive DNA, and their genes undergo degeneration [10][12].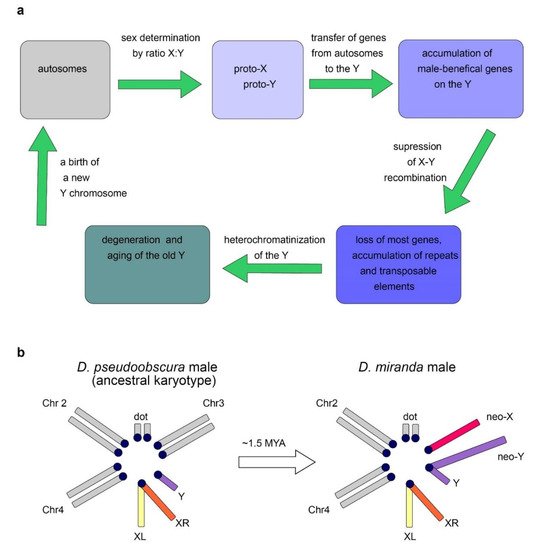
2.2. Origin of the Y Chromosome in Mammals and Sex Determination
In many animals, sex is determined by a pair of heteromorphic X and Y chromosomes. According to modern concepts, sex chromosomes originate from an ancestral pair of autosomes, one of which acquires a sex-specific gene, which starts the process of differentiation of the sex chromosomes. In mammals, this event occurred only once in the common ancestor of marsupials and placentals prior to their splitting, about 160–180 MYA [19][20][21][22]. The proto-Y chromosome of all mammals (from kangaroo to human) arose from a single autosome in which one of the alleles of the SOX3 gene, as a result of a mutation, became the sex-determining gene SRY [23][24]. It is important to note that this gene is not responsible for all sex characteristics alone. The product of the sex determination gene only provides a switch, triggering a certain pathway of development. Unique evolutionary forces facilitated the selection and accumulation of male-beneficial mutations around the SRY locus, and the linkage between them was supported by selective pressure to avoid crossing over between the proto-Y and proto-X [25]. As a rule, if the dominant allele causes the development of a male, then the chromosome in which it is located becomes the Y chromosome (and its homolog is called the X). In birds, males are the homogametic sex (ZZ) and females are the heterogametic (ZW) [21]. Currently two main hypotheses about sex determination in birds are presented. One of them postulates the presence of the key gene controlling ovarian development or inhibiting testis differentiation in the W chromosome, while the other one proposes the number of Z chromosomes as a key sex-determining factor [21][26]. In last case, sex determination is thought to be provided by a sex chromosome gene dosage mechanism, and the most likely sex determinant is the Z chromosomal gene DMRT1 encoding transcription factor. Recently it was shown that male chicken (ZZ) with a single functional copy of DMRT1 (other was deleted by a CRISPR-Cas9-based monoallelic targeting approach) developed ovaries in place of testes. It indicated that DMRT1 is the key sex determination switch in birds essential for testis development [26]. In addition, it was found that the synthesis of estrogen is also an essential factor in primary sex determination in chicken, and that estrogen production is controlled by expression of DMRT1 [26]. These data support the second hypothesis that the dosage of genes on the Z chromosome determines the sexual differentiation in birds [26].2.3. Evolutionary Factors and Forces Determining the Structure and Functional Specialization of the Y Chromosome
The loss of recombination leads to the inefficiency of natural selection and causes the ensuing accumulation of Y-linked loss-of-function mutations, chromosome-wide gene decay, and amplification of repetitive DNAs [27][28][29][30]. In parallel to the loss of genes, Y chromosomes have accumulated large amounts of DNA repeats, and the D. melanogaster old Y chromosome mainly consists of heterochromatin (Figure 2a) [4][21]. Despite the human Y chromosome having undergone a rapid decay early in evolution, its massive degeneration then dramatically stopped. Genes that remained intact currently show remarkable stability, and no human Y-linked genes have been lost during the last 44 million years [22][31]. The maintenance of human Y-linked genes is mainly associated with two functional categories: genes essential for male reproductive functions and dosage-sensitive ubiquitous housekeepers [32]. Studies of males with Y deletions have allowed researchers to identify three ‘azoospermia factor’ (AZF) regions, AZFa, AZFb, and AZFc, and partially map within them the genes essential for spermatogenesis [33]. The AZFa deletions affecting the DBY gene cause the most severe azoospermia phenotype, exhibiting a complete loss of testis germline cells accompanied by the maintenance of somatic Sertoli cells (the so-called Sertoli Cell-Only Syndrome; SCOS) [34][35][36]. As in fruit flies, mammalian Y chromosomes also exhibit gene amplification, with the amplicon structures predominantly containing genes with testis-specific functions [37]. The structure of such genes is maintained by intra-chromosomal gene conversion. The amplicon region of the human Y chromosome contains eight massive palindromes ranging in length from 9 kb to 1.45 Mb with nucleotide identity of the arms over 99.9% [38][39][40]. Due to the presence of repeating structures, local intra-chromosomal gene conversion is possible, as well as intra- and inter-chromatid exchange. These mechanisms partially compensate for the lack of recombination with the X chromosome by eliminating harmful mutations. At the same time, inter-chromatid recombination can in some cases lead to the formation of isodicentric chromosomes formed by homologous crossing over between opposing arms of palindromes on sister chromatids. This may be accompanied by the loss of certain regions containing genes essential for spermatogenesis, and in some cases can lead to the loss of the Y chromosome during cell division with clinical consequences ranging from spermatogenic failure to sex reversal and Turner syndrome [41]. The loss of the ability to recombine plays a key role in establishing the structure of the Y chromosome, because recombination could lead to a disruption of sex determination and the formation of infertile intermediate variants [30][42]. Conversely, mutations that prevent recombination between proto-X and proto-Y, such as inversions, deletions, or accumulation of repeats, are supported by selection. Reducing the ability of recombination with the homologous X chromosome dramatically accelerated the evolution of the Y chromosome preventing the elimination of emerging mutations via crossing over, while the X chromosome has retained the ability to cross over in the homogametic sex. This led to the degeneration of most of the original Y-chromosomal genes, and multiple deletions caused a significant size decrease with a relative increase in the proportion of non-coding heterochromatic regions. The rapid evolutionary degeneration of the Y chromosomes, typical in a wide range of species, leads to the hypothesis that in the future the human Y chromosome may disappear altogether. This hypothesis is based not only on extrapolation, but is also indirectly supported by precedents in the evolution of some species including multiple fishes, reptiles, grasshoppers, cockroaches, and dragonflies [43][44][45]. However, other researchers claim that human Y degeneration stopped millions of years ago and currently nothing threatens Y chromosome survival [46].2.4. Dosage Compensation System Contributes to Y-Linked Gene Maintenance
As a rule, a single gene copy appears to be enough to provide development and life-cycle maintenance of diploid animals; however, a small cohort of genes exhibits a high sensitivity in case of decreased gene dosage. This phenomenon is known as haploinsufficiency, and it is associated with many developmental disorders in human [47][48][49]. Comparing the evolution of flies and humans, one could assume that the Y-chromosomal genes, which have homologues on the X chromosome and do not directly contribute to the functioning of the male reproductive system, are relics that will disappear over time, as has apparently happened in flies. However, their maintenance can be determined by the peculiarities of the dosage compensation system in mammals. In male flies, the genes of the only X chromosome are overactivated in somatic tissues, eliminating the problem of haploinsufficiency and potentially lethal imbalance between the X and autosome transcriptional level in the two sexes. Therefore, in flies, X activation may eventually compensate for haploinsufficient homologous genes lost on Y, which is impossible in mammals. In contrast, in female mammals, inactivation of one of the two X chromosomes occurs. However, according to various estimates and in distinct types of human cells, 20–30% of genes of inactive X chromosome escape the inactivation [50][51]. In mammals, haploinsufficient Y-chromosomal genes have X-chromosomal homologues that avoid inactivation during dosage compensation in females, which indicates the need for their expression on both sex chromosomes to ensure normal functions in the body. Thus, in males, these dosage-sensitive genes cannot disappear from the Y chromosome without negative consequences, and they can survive under selective pressure [31]. This hypothesis is supported by the maintenance of functional X-Y gene pairs associated with housekeeping regulatory functions such as lysine demethylation, stem cell self-renewal, splicing, translation initiation, and deubiquitylation [31][32][50][52]. Strict dosage requirements for sex-linked genes are demonstrated in the case of Turner syndrome (exhibiting X0 karyotype or mosaicism) and Klinefelter syndrome (XXY), since such genes have been haploinsufficient or overexpressed, respectively, in these karyotypes [51]. Turner syndrome is a genetic condition caused by complete or partial loss of the second sex chromosome in human. Half the patients with Turner syndrome have the X0 karyotype (monosomy of the X chromosome), the other half exhibits mosaicism or a presence of the fragmented X or Y chromosomal material and other more complex karyotypes [53][54]. Studies of manifestations of this syndrome indicate that the functions of the Y chromosome consist not only of ensuring the normal functioning of the male reproductive system. Due to the absence of the SRY gene, which is the key to triggering male-type development, patients with this syndrome are exclusively female, with multiple body disorders: small size, rudimentary ovaries and infertility, pathologies of the cardiovascular system, autoimmune disorders, increased risk of developing diabetes, and cognitive impairment [54]. Individuals with Klinefelter syndrome are infertile as a result of excess gene dosage of X escape genes, and abnormal meiotic pairing of the sex chromosomes. An atypical number of X or Y chromosomes (XXY, XXX, or X) contributes to spatial chromosome conformation changes and leads to disruption of DNA methylation patterns of autosomal genes, causing distinct disease phenotypes: mental illness, cancer, and disrupted fertility [51].2.5. Convergent Nature of the Evolution of Y Chromosomes
Despite their independent evolutionary origins in different species, Y chromosomes in species with heterogametic males have a number of similar features: they are usually smaller than X chromosomes, contain significantly fewer genes, most of which are related to the male reproductive system, and also have a relatively large number of repeats and significant areas occupied by heterochromatin. It is worth noting that in species with heterogametic females, such as birds, the sex-specific chromosome W also resembles the Y chromosome in structure, and is characterized by relatively small size, heterochromatinization, and fewer genes [21]. Such common patterns indicate the convergently evolved structural features of these chromosomes. It has been proposed that such convergent evolution is due to the similar nature of the selection pressure. Another common feature—the acquisition of repetitive sequences and the loss of most of the original genes—is associated with accelerated Y evolution due to the loss of recombination with the X chromosome [27]. The difference between the evolution of the Y chromosome in mammals and Diptera is mainly that in Diptera the acquisition of new genes often significantly prevails over the loss of the original ones; although, both processes take place in both groups. Presumably due to slower changes in mammals, the evolutionary processes have not yet reached the point where the Y chromosome has lost all homology with the X chromosome. Attempts to understand how these patterns are generated can be important not only for fundamental evolutionary biology, but also for biomedical challenges, since Y-chromosomal pathologies in humans differ from other genetic anomalies due to the unique nature of the Y chromosome. It is convenient to study the Y chromosome evolution in species with a rapid generation turnover, as in the Drosophila species.3. Fertility Factors and Peculiarities of Giant Gene Transcription
3.1. Fertility Factors and Y-Loop Formation in Drosophila
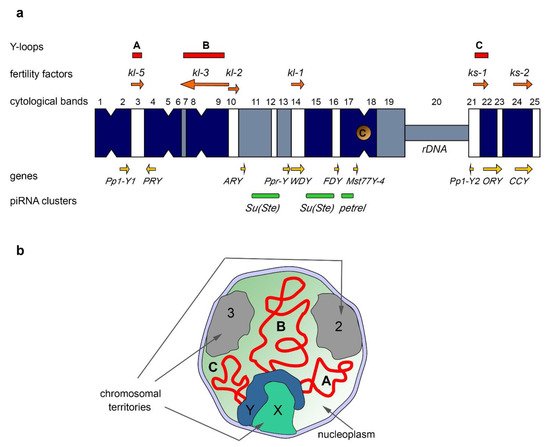
The Y chromosome of D. melanogaster, in its current state, contains a few protein-coding genes primarily expressed in the testes. With the aid of classical genetic methods, including X-irradiation, chromosome deficiencies, X-Y translocations, P-element insertion mutagenesis, and complementation assays, it has been discovered that the Y chromosome contains at least six distinct loci required for spermatogenesis and male fertility [55][56][57][58][59][60][61][62][63] (Figure 1a). These six loci encoding the so-called male fertility factors are located both on the long arm (kl-5, kl-3, kl-2, and kl-1) and the short arm of the Y chromosome (ks-1 and ks-2) [57][60][64][65]. The kl-2, kl-3, and kl-5 genes encode dynein heavy chain proteins that are essential for proper axoneme building in elongating spermatids. Kl-2 is an inner dynein arm heavy chain protein and Kl-3 and Kl-5 are outer dynein arm heavy chain proteins [66][67][68]. Note that D. melanogaster has several other dynein heavy chain genes, located on the chromosomes X, 2, and 3 [69]. The axoneme is the microtubule-based main part of the sperm flagellum, a specific motile organelle of spermatids and mature sperm. In the motile flagellum, dynein ATPase motor proteins provide sliding motions between adjacent microtubules, which together produce well-ordered movements [70]. Deficiency of the kl-3, kl-5 or kl-2 genes leads to loss of the outer dynein structure of the axoneme, and these mutants do not produce motile sperm, resulting in male sterility [65][67][71]. kl-2, kl-3, and kl-5 mutant males exhibit clear defects in spermatid morphology and development with the loss of synchronization of their individualization complexes; in addition, they contain short and curled spermatids, nuclei of which are scattered instead of remaining tightly clustered, as in wild-type flies [65]. The gene sequences of ks-1, ks-2, and kl-1 had not been identified for a long time, complicating their disruption and studying. However, according to a recent study, the CCY gene located near the telomere of the Y short arm is thought to encode male fertility factor ks-2. RNAi-knockdown of CCY results in short and curled nuclei of elongating spermatids and in male sterility [65][72]. In addition, RNAi-knockdown of the WDY gene from the kl-1 locus leads to male sterility, supporting the conclusion that WDY encodes the Kl-1 fertility factor [72]. According to publicly available RNA-seq data, most Y-linked genes begin their expression in third-instar larvae and continue to be expressed throughout the pupal stage with the exception of the FDY gene, which is expressed during all developmental stages. Several Y chromosome genes are found to be expressed in imaginal discs, fat bodies, accessory glands, and all of the genes are expressed in the testes [65]. In the D. melanogaster testes, generation and transcription of Y-loops take place during primary spermatocyte maturation, coinciding with the data that Y expression peaks in spermatocytes [73]. Transcription of the kl-2, kl-3, and kl-5 genes starts in primary spermatocytes and continues during the whole 80–90 h of meiotic G2 phase. Their spliced transcripts are stored in cytoplasmic RNP particles, called kl-granules, in mature spermatocytes along with the ATPase proteins Reptin and Pontin [74][75]. These RNP granules segregate during the meiotic divisions and the stored dynein transcripts undergo delayed translation, occurring post-meiotically. Then the dynein proteins are incorporated into the axoneme during the spermatid elongation process [75]. Fertility factor genes contain unusually large, megabase-sized introns filled with simple satellite repeats in the cases of kl-5, kl-3, and ks-1 genes [1][64][76][77][78]. For instance, the kl-3 gene spans at least 4.3 Mb, while its coding sequence contains only approximately 14 kb [64][76]. The giant introns comprise more than 99% of the whole kl-3 locus. Transcription from kl-3, kl-5, and ks-1 loci in spermatocytes leads to the appearance of lampbrush-like nucleoplasmic structures named Y-loops A (kl-5), B (kl-3), and C (ks-1) (Figure 1a,b), that are visible with the aid of phase-contrast microscopy, and are analogous to those in amphibian oocytes [76]. The kl-5 locus contains four different satellite repeats; the loop-forming site of kl-5 accumulates (AAGAG)n, (AAGAC)n, and (AAGAGAG)n repeats, while the kl-5 non-loop-forming region contains (AATAT)n repeats. Similarly, the ks-1 locus encompasses (AAGAG)n and (AAGAC)n repeats mapping to the loop-forming region in contrast to (AATAAAC)n and (AAGAG)n repeats occupying the non-loop-forming regions. The kl-3 loop is composed of a thinner filament and exhibits a rather diffuse appearance. Only (AATAT)n repeats have been mapped to the kl-3 loop-forming region [1]. Y-loop generation reflects the high transcription levels of the underlying genes. Proper transcription of giant genes requires high processivity of RNA polymerase II (Pol II). The presence of long satellite arrays in the introns can lead to the slowing of elongation or frequent premature dissociations of RNA polymerase. Satellite repeats can form high-order DNA, RNA, or DNA–RNA hybrid structures, which may inhibit transcription elongation. Thus, transcription of gigantic intron-containing genes requires precise regulation, and Y-loops contain chromatin associated with a large number of transcripts and regulatory proteins [77][79]. Three RNA-binding factors, Blanks, Hephaestus (Heph), and Maca, are found to be enriched specifically in the chromatin of the Y-loops [74][80][81]. The corresponding mutations lead to male sterility owing to defects in sperm individualization, similar to the ones observed in males with knockdowns of the kl-5, kl-3, and kl-2 genes [74][81]. These proteins are required for transcription or proper processing of the Y-loop gene transcripts. Blanks is found to be located to the Y-loop B (comprising mainly kl-3 introns) and is needed for proper kl-3 mRNA expression [74][80]. It has been assumed that Blanks maintains Pol II processivity by binding to the nascent transcripts of kl-3 [74]. The Heph protein is found to colocalize with Y-loops A and C. It can regulate the expression of kl-3, kl-5, and ks-1 mRNAs in spermatocytes. Heph may be involved in the processing of kl-5 transcripts, including splicing, or in preventing their premature degradation [74]. Maca is essential for kl-2 and kl-3 transcription and proper splicing of kl-3 transcripts, because in maca knockdown testes, the skipping of exon 13 causes an internal deletion in Kl-3 protein [81]. Recently described testis-enriched transcription regulators, tPlus3a and tPlus3b, appear to be required for the expression of fertility factors kl-3 and kl-5 [82] via an unknown mechanism. Some other RNA-binding proteins are enriched on the Y-loops in the spermatocyte nuclei, such as Pasilla and Boule [79][83], Rb97D [84], and several proteins encoded by unannotated genes (lup-3, lup-4, dolly-1, and dolly-2) [85]. However, their functions in Y-loop transcription await further investigation. Y-loop generation is a conservative feature across the Drosophila genus, including Drosophila simulans, Drosophila yakuba, Drosophila pseudoobscura, Drosophila littoralis, and Drosophila hydei [86][87][88][89][90]. In the spermatocytes of D. hydei, clearly cytologically visible Y-loops are found, and early studies have uncovered that their transcription is associated with the huge DNA repeats [91][92][93]. Although the functional relevance of the gigantic introns still remains unclear, according to some assumptions long lasting transcription of fertility factor genes (around 80–90 h), due to the presence of the gigantic introns, appears to function as a ‘developmental timer’ for spermatocyte growth and differentiation [74][81]. Intron size could also play a critical role in the regulation of gene expression. It has been shown for the Ultrabithorax (Ubx) gene in the early Drosophila embryo that its large size causes abortive Ubx transcription during the syncytial divisions, blocking expression of Ubx protein at the syncytial stage [94].3.2. Intron Gigantism in Humans
The phenomenon of ‘intron gigantism’ occurs across multiple species, including vertebrates, however, little data are available about its functional significance. Several human neuronal and muscle genes are known to bear giant introns. The best-known largest human gene is dystrophin comprising nearby 0.1% of the whole genome, containing 79 exons and spanning 2.2 Mb, with only 11 kb of coding sequence [95][96]. Its gigantic introns, also rich in repetitive DNA sequences, are reminiscent of those of Y-linked Drosophila fertility factors. Dystrophin is located on the p21 region of the X chromosome and codes the causative gene for Duchenne Muscular Dystrophy (DMD) and Becker Muscular Dystrophy (BMD) [95][97]. Dystrophin is a major scaffolding component of normal muscle, which links cytoskeletal actin, tubulin, and intermediate filaments to the extracellular matrix, and stabilizes the plasma membrane of striated muscle cells. Loss-of-function mutations of the dystrophin gene trigger instability of the plasma membrane and lead to myofiber loss [97][98]. Transcription of dystrophin takes place from several promoters in a tissue-specific manner. Full-length Dystrophin is expressed in all striated skeletal, smooth, and cardiac muscles. Shorter isoforms are expressed in brain and retina cells. In case of frameshift mutations, deficiency of the Dystrophin protein leads to severe DMD disease. In case of in-frame mutations, Dystrophin is expressed as a set of mutated proteins either with missense substitutions or deletions or duplications of its internal part, leading to the weaker BMD disease [97]. Exon skipping, with the aid of antisense oligonucleotides to skip the problem exons containing premature stop codon mutations or reading-frameshift mutations, is currently used as an approach for DMD therapy [99]. Transcription of these extremely large genes and the processing of their transcripts, including splicing, has a high metabolic cost for cells. The study of genes possessing giant introns using the Drosophila model provides a useful insight into the problems of expression of such genes in humans and the pathologies associated with their improper transcription or splicing. Study of the molecular mechanisms of maintenance, transcription, and processing of Y-loop genes in Drosophila may improve understanding of the origin, selection, and regulation of genes with similar structures in other species.4. Differential Expression of rDNA Loci of Drosophila Sex Chromosomes
4.1. Nucleolar Dominance as a Widespread Phenomenon
In eukaryotes, there is a known phenomenon of a different level of expression of genes represented in the genome by two or more alleles [100][101]. Some of these alleles are expressed at a high level, while the expression of the rest of them is completely suppressed. One of the most striking examples of this phenomenon is the regulation of expression of loci encoding ribosomal RNA (rRNA), called nucleolar dominance. This phenomenon was initially discovered in interspecies hybrids of different taxonomic groups of animals [102][103][104]. In interspecies hybrids between D. melanogaster and D. simulans, rRNA genes from the D. melanogaster genome are predominantly expressed, while these genes from the D. simulans genome are suppressed [105]. However, later this process was also found within species. Nucleolar dominance has been observed in both the plant and animal kingdoms and is generally characterized by the dominant transcription of rRNA loci residing in only one chromosome [106][107][108]. Among the reasons for this phenomenon, DNA cytosine methylation, histone methylation and deacetylation, small RNA functions, and different affinities between transcription factors and promoter sequences of ribosomal DNA (rDNA) loci are suggested [109][110]. However, the exact mechanism of this phenomenon remains unclear to date. rDNA loci are arranged as tandem repetitive rRNA gene clusters flanked by intergenic spacer sequences (IGSs) [109][110]. They are transcribed by the RNA polymerase I machinery as long precursor transcripts, subsequently processed into mature ribosomal RNAs (18S, 5.8S, and 28S). The transcriptional activity of these loci is high and achieves about 50–60% of the total transcription of metabolically active cells [102][111]. rRNAs are highly conserved, but the loci that encode them are among the most unstable elements of the genome due to their repetitive nature and high transcriptional activity. The number of copies of rRNA genes varies from 100 to 1000 in different organisms, and they are often distributed over many chromosomes, including ten loci in humans [112]. In mice, about 200 rDNA repeats grouped into NORs (nucleolar organizer regions) are distributed among the short arms of six acrocentric chromosomes [102]. rDNA can undergo intrachromatid recombination, which can lead to a loss of rDNA copies or to the formation of circular non-genomic units-extrachromosomal circular rDNAs (ERCs) accumulating in aging yeast cells [113]. In addition, active transcription of rDNA occurs even during the S-phase of the cell cycle, which can cause a conflict between replication and transcription. Such conflicts lead to frequent double-strand breaks and rDNA instability [114]. Given the multifactor nature of rDNA instability, the number of rDNA copies in the loci can vary greatly even within populations of the same species. For instance, in D. melanogaster strains, the variation in the number of rDNA copies can reach a sixfold range [115]. Similar differences in the number of rDNA copies have been shown for a number of other organisms, including mice and humans [116]. Decreased rDNA copy number leads to so-called replicative senescence in yeast [113][117][118]. Nevertheless, despite significant variations, there are mechanisms that maintain the number of rDNA copies both in populations and in the process of transmission to subsequent generations. Therefore, it is important to study the mechanisms of maintenance of rRNA gene copies at a level necessary for survival.4.2. Y-Based Nucleolar Dominance in D. melanogaster Males
In D. melanogaster, rDNA loci reside on the X and Y chromosomes, each containing from 100 to 360 copies of rDNA genes. The presence of rDNA loci on the sex chromosomes greatly simplifies their study compared to other model organisms in which such loci are numerous and distributed over a large number of chromosomes. In D. melanogaster males, intraspecies epigenetic silencing of X chromosomal rDNA in males was shown by two research groups in 2012 [107][108]. However, these studies of nucleolar dominance were based on larval neuroblasts and total RNA preparations from adult flies. While rDNA gene transcripts on the X and Y chromosomes are highly homologous, some genes contain insertions of non-LTR retrotransposons R1 and R2 [119]. These retrotransposons are able to specifically recognize a 30 bp target sequence in the transcribed region of 28S rDNA and integrate into this region preventing correct transcription of the whole cistron. It has been suggested that the nucleolar dominance of Y-linked rDNA loci over those in the X chromosome is partly due to the different number of transposon insertions in the rDNA loci. X-chromosomal rDNA loci contain a higher proportion of genes disrupted by the transposons than Y-chromosomal ones [120]. For instance, the wild-type X chromosome contains insertions in 80% of rDNA units out of 100, while the wild-type Y chromosome contains insertions in 60% of rDNA units out of 360 [107]. The Y chromosome, with approximately the same number of insertions in the rDNA loci, does not show complete dominance over the X chromosome, but provides codominance (some expression of rDNA from the X chromosome occurs). Thus, there must be other factors that, together with the abundance of insertions, contribute to the nucleolar dominance of the Y chromosome. Interestingly, all analyzed lines of Drosophila whose Y chromosome did not exhibit complete dominance carried mutations in the genes of heterochromatin proteins Su(var)2–5 (HP1) and Su(var)3–9 (encoding histone H3K9 methyltransferase). However, the authors failed to establish an unambiguous relationship between mutations of heterochromatin protein genes and derepression of rDNA loci in the X chromosome [107]. Previously, it has been shown that in the absence of H3K9 methylation (in the case of a Su(var)3–9 mutation) and upon disruption of the siRNA pathway (a dcr-2 mutation) disorganization of nucleoli, rDNA loci, and adjacent satellite DNAs were observed [121]. Recently, nucleolar dominance was analyzed in detail in dcr-2 and Su(var)3–9 mutants. Males with the dcr-2 mutation showed no significant change in Y chromosome dominance during fly development, while males with the Su(var)3-9 mutation demonstrated a significant decrease in Y dominance in nervous tissue of larvae, imaginal discs, and germline stem cells (GSCs) of adult males [122]. Thus, heterochromatin-mediated repression of rDNA loci may contribute to the mechanism that regulates of their activity. A recent work describes the investigation of nucleolar dominance of Y-linked rDNA loci in male GSCs of D. melanogaster [123]. Although in the testes of young males, most GSCs contain a single spherical nucleolus 2 µm in diameter; however, during aging, the proportion of GSCs with normal nucleolus morphology gradually decreased, while the proportion of GSCs with atypical morphology increased. Atypical nucleolus morphology was manifested both in the fragmentation of the nucleoli into several foci, or in the altered nucleolar shape. Authors found that only Y-linked rDNA loci are associated with the nucleolus with typical nucleolar morphology, while X-linked loci are not, regardless of age. These results suggest that Y-chromosomal rDNA is actively transcribed, while X rDNA is not, which is consistent with Y nucleolar dominance. However, atypical nucleolar morphology that occurs in GSCs with aging is associated with the activation of the silent rDNA loci on the X, and leads to the transcription of rDNA from two separate chromosomes, each of which forms a separate nucleolus. Activation of X rDNA probably compensates for the decrease in the number of active copies of Y-linked rDNA, which decreases during aging owing to conflicts between transcription and replication machineries causing rDNA instability [114][123]. GSC nucleolar morphology and rDNA copy number reduction is heritable and passed to male offspring from old fathers. Strikingly, the authors find that nucleolar morphology can be recovered in individual GSCs of these F1 sons during the first 10 days after eclosion to restore normal Y dominant state of rDNA transcription. This indicates the existence of a mechanism to maintain the number of rDNA copies across generations. This mechanism may be adaptive for the following reasons: firstly, rRNA expression from only one chromosome can prevent rDNA deletions on the other chromosome, transcription from which is suppressed; secondly, having intact rDNA loci present may allow GSCs to prolong their lifespan [123]. Recently, the SNP in situ hybridization method was used to analyze in detail the transcription of rDNA clusters from the X and Y chromosomes of D. melanogaster [122]. Throughout Drosophila male development, the codominance of X and Y rDNA loci changes to the dominance of those on the Y chromosome. The manifestation of Y dominance in most types of larval tissues, such as nervous tissue, imaginal discs, fat body, and enterocytes of the anterior part of the midgut, has been found. However, salivary glands containing a large number of polytenized chromosomes showed only a modest manifestation of Y dominance. In females, using the SNP method, the codominance between the two X chromosomes was confirmed. Drosophila females with the XXY genotype also exhibit Y dominance, suggesting that the presence of the Y chromosome is necessary and sufficient for the dominance. However, in the ovaries of adult females, codominance is also observed in GSCs and cystoblasts, while in the nurse cells Y dominance is found. Thus, in the case of the presence of the Y chromosome, nucleolar dominance predominantly occurs independently of the sex of the cell. This leads to the assumption that the Y chromosome must contain a specific nucleotide sequence that allows dominance to occur. In general, the sequences in rDNA loci of the Y chromosome and/or its proximal regions may be essential for the nucleolar dominance. Moreover, these results do not exclude the possibility that the long arm of the Y chromosome is involved in this process [122].4.3. Non-Random Segregation of Sister Chromatids of Sex Chromosomes in Drosophila
Recent studies also point to the involvement of rDNA loci in the nonrandom segregation of sister chromatids during cell division. The intergenic spacer repeats are responsible for X-Y pairing in D. melanogaster males [124]. Sister chromatids are not always completely identical due to the presence of epigenetic marks that distinguish them. The asymmetric arrangement of these marks, as well as kinetochore proteins, can lead to selective recognition of chromatids. The divergence of such sister chromatids is apparently one of the causes of asymmetric cell division. Recent studies have shown that non-random sister chromatid segregation is mediated by rDNA loci [125]. Earlier it was shown that non-random segregation in Drosophila is characteristic of the sister chromatids of the X and Y chromosomes, but not the autosomes [126]. Researchers used the Drosophila strain carrying a deletion of 80% of heterochromatin in the wild-type X chromosome (Df(1)bb158). In GSCs of such fly males, random segregation of X chromosome chromatids occurred, while the segregation of the Y chromosomes remained non-random, suggesting that a chromosomal element deleted in the bb158 strain acts in cis to mediate non-random sister chromatid segregation. This indicates that the genetic elements necessary for this phenomenon are present in the deleted heterochromatin. It can be concluded that this genetic element is located in the rDNA loci. A more detailed study revealed that IGS sequences and the protein that binds to these sequences, Indra, are responsible for the non-random segregation of sister chromatids [125]. Unequal sister chromatid exchange can be proposed as a possible mechanism to increase rDNA copy number on one sister chromatid for restoration of the number of rDNA copies disrupted in GSCs by aging.4.4. Differential Expression of rDNA Loci in Human
The phenomenon of nucleolar dominance appears to be common across multiple species. It has not been shown directly in humans, due to the distribution of rDNA loci in multiple autosomal regions making them difficult to analyze owing to their highly repetitive nature. However, only a part of rDNA loci is actively transcribed in human cell lines [127][128][129], suggesting that these loci may also undergo activation or suppression. To date, the principles of silencing or activation of rDNA loci in humans remain unknown. Recently, with the aid of Oxford Nanopore sequencing technology, obvious differences between methylated and unmethylated rDNA gene arrays in human cells have been revealed. The ratio of transcriptionally active unmethylated copies versus methylated ones has been found to be lower in individuals with higher rDNA copy abundance, indicating a possible mechanism for maintenance of a stable number of active rDNA copies [129].5. Drosophila Y Chromosome in Studying of piRNA Biogenesis and Functioning of piRNA-Clusters
5.1. Brief Description of the piRNA System
The piRNA pathway provides both innate and adaptive immune system defense against the activity of transposable elements (TEs) leading to the protection of genome integrity in germinal tissues. It also participates in the maintenance of germline stem cells, regulation of protein-coding gene expression, the establishment of embryonic patterning (in Diptera), and transgenerational epigenetic inheritance [130][131][132]. Small non-coding piRNAs 23-35 nt in length associated with proteins of the PIWI subfamily are present in animals from fungi to humans [133][134][135]. piRNAs are generated from piRNA clusters, which are long precursors that are transcribed from heterochromatic regions containing fragments of transposons. piRNA precursors are processed to generate small piRNAs in perinuclear nuage granules. Mature piRNAs loaded into the proteins of the PIWI subfamily, forming piRNA-induced RNA silencing complexes (piRISCs). The generation of primary piRNAs triggers production of secondary piRNAs via an amplification system called the ping-pong cycle (Figure 3) [136]. The piRNA pathway is active, as a rule, in gonads and plays an essential role in fertility maintenance, preventing transposon activity and repressing harmful protein-coding genes. Transcripts of harmful genomic elements can be silenced post-transcriptionally via recognition and cleavage of complementary RNA-targets by piRISC complexes in the nuage. There is also a co-transcriptional repression mechanism, where recognition of nascent transcripts by piRISCs loaded with guide piRNAs leads to the establishment of heterochromatin in the corresponding genomic regions. Most of the known piRNA clusters in Drosophila are bidirectional and transcribed with the participation of a specific Rhino–Deadlock–Cutoff (RDC) complex (Figure 3) [137][138][139]. Due to the Rhino chromodomain the RDC complex recognizes H3K9me3 histone modifications enriched in the chromatin of piRNA clusters and recruits to them the transcription initiation factor Moonshiner to promote non-canonical transcription. The RDC complex allows a skipping of transcription termination sites and inhibits splicing of piRNA precursor transcripts [137][138][139]. In contrast, in mammals, the mechanisms of piRNA cluster expression seem to be indistinguishable from canonical Pol II transcription and include regular splicing and polyadenylation of the transcripts [131]. However, the role of piRNAs may not be limited to germinal tissues. The involvement of the piRNA pathway in processes associated with disease, such as tumors of various etiologies, and aging in humans, has recently been shown [140][141].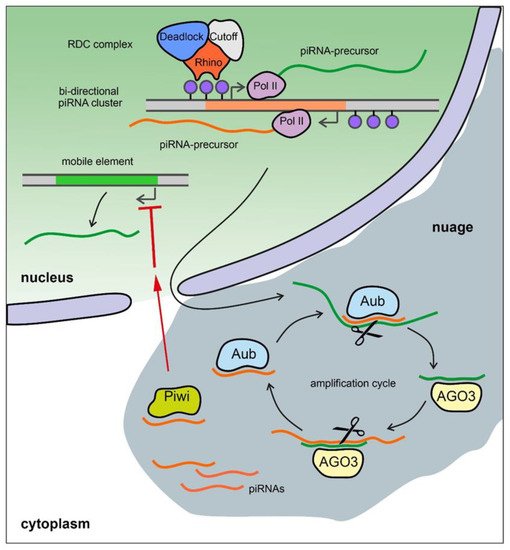
5.2. The Y Chromosome as a Major piRNA-Producing Genomic Region in the Fly Testes
The piRNA system in D. melanogaster exhibits a strong sexual dimorphism. TE-mapping piRNAs are known as the most abundant class of piRNAs in the ovaries, whereas only about 40% of piRNAs map to TEs in the testes, and the largest cohort of piRNAs map to protein-coding genes [7][142]. The genomic origin of most piRNAs between the two sexes is also different. In the testes of Drosophila, almost half of all piRNAs originate from the piRNA clusters located on the Y chromosome (Figure 1a) [7]. The largest number of piRNAs is generated from the Y-linked Suppressor of Stellate (Su(Ste)) repeats directed to silencing of the homologous tandem Stellate genes residing on the X chromosome [142][143][144]. The organization of the Su(Ste) loci has been studied in detail. The number of Su(Ste) repeats comprises more than 500 tandemly ordered copies residing in two cytolocations on the Y (Figure 1a) [2][4][7]. The size of a typical Su(Ste) repeat is about 28 000 nt, consisting of three main parts: region homologous to the Stellate genes, the AT-rich Y-specific region, and the insertion of transposon hoppel (1360) into the promoter. Su(Ste) repeats are transcribed and processed to polyadenylated mRNAs; however, they contain numerous frameshift mutations owing to the presence of point mutations and deletions, and they are not translated [145]. The insertion of the defective transposon hoppel is responsible for the initiation of antisense transcription of Su(Ste) repeats and their acquisition of piRNA cluster functions [143]. Stellate derepression in the case of deletion of most of Su(Ste) repeats or disruption of the piRNA system leads to the accumulation of needle-like protein aggregates in spermatocytes, disturbances of meiosis, and, as a result, a decrease in male fertility [143][146]. The Stellate/Su(Ste) system is species- and sex-specific for D. melanogaster. Earlier, it was proposed that Stellates are selfish genes involved in meiotic drive [147]; however, no experimental pieces of evidence of this assumption have been found to date. Recently, it was is shown that Stellate genes participate in male hybrid sterility of F1 progeny of crosses between D. melanogaster females and D. mauritiana males. The hybrid males possess maternal X-linked Stellate genes, but their paternal Y chromosome does not contain Su(Ste) repeats and the corresponding piRNAs are not generated. Derepression of Stellates in the testes of hybrid males leads to a meiotic catastrophe and complete sterility [142][146]. The contribution of the Stellate/Su(Ste) system to reproductive isolation may explain the fixation and maintenance of this system in the D. melanogaster genome. The acquisition of the Stellate/Su(Ste) system by a part of the ancient fruit fly population could have been a causative factor of hybrid sterility in crosses of females with males that do not possess Su(Ste) repeats on the Y. According to another speculation, the Stellate/Su(Ste) system is similar to toxin–antitoxin systems, which are widespread in prokaryotes [148]. Considering the amplification processes as an inherent property of the Y chromosomes, it can be assumed that some amplified repeats can be licensed as piRNA clusters during the evolution of a species. Given that not all piRNAs map to TEs, piRNAs produced by regions of the Y could exert sex-specific functions to regulate the expression of protein-coding genes besides Stellate. If a gene has a positive effect on the processes occurring in the ovaries or other tissues, but, at the same time, reduces the efficiency of spermatogenesis, mechanisms for its testis-specific suppression can be developed, including piRNA-mediated silencing. The existence of a similar mechanism has been recently confirmed for the X-chromosomal pirate/CG12717 gene, encoding a SUMO-isopeptidase. The Y chromosome of D. melanogaster contains the petrel locus (Figure 1a), which is a source of multiple piRNAs highly complementary to pirate, providing strong testis-specific silencing of this gene [7]. However, the functional significance of the repression of pirate in the testes remains unclear to date. It appears that both in the cases of the Stellate/Su(Ste) and pirate/petrel pairs, their current evolutionary relationships are initially based on parallel acquisition or co-amplification of homologous genes on the sex chromosomes. Note that in the Drosophila testes, the expression and activity of the RDC complex is mainly limited to early stages of spermatogenesis, including GSCs and spermatogonial cells [149]. The transcription of Y-linked Su(Ste) and petrel piRNA clusters takes place in primary spermatocytes, and it is independent from the RDC complex [149]. The complex mosaic structure of petrel repeats [7] makes their further study as a functional piRNA cluster difficult. In the case of Su(Ste), its sense transcription performs in the canonical manner from its own promoter, and the antisense transcription is initiated from several sites within the inserted transposon hoppel [143][150], which makes it similar to mammalian piRNA clusters. It has been assumed that in Drosophila, maternal piRNAs, which are stored at the posterior pole of the oocyte during oogenesis, ensure the initiation of piRNA biogenesis from long piRNA precursors. However, this does not apply to Y chromosomal piRNA clusters due to the absence of the Y chromosome itself in females. According to a recent study, the determination of long RNAs as primary piRNA sources can also occur due to the recognition of specific cis-regulatory 100-nt elements in piRNA precursor sequences, as in the case of the long non-coding RNA flamenco and 3′ UTR of tj mRNA in ovarian somatic cells [151]. In many animals, including humans, the induction of germ cell precursors occurs from somatic pluripotent epiblast cells during embryogenesis, as a result of which all piRNA clusters are determined de novo [132][135]. On the whole, the mechanism of determination of genomic regions as piRNA clusters is poorly resolved. Future studies of Y-chromosomal piRNA clusters in Drosophila could allow us to elucidate these mechanisms both in Drosophila and in mammals.5.3. The Y Chromosome in Other Species as a Source of piRNAs
The suppression of genes harmful for spermatogenesis appears to be one of the main functions of piRNAs originating from the Y chromosome of D. melanogaster. In mouse testes, novel polyadenylated non-coding RNAs called Pirmy and Pirmy-like transcribed from the long arm of the Y chromosome have recently been discovered [152]. Multiple splice variants of Pirmy encoded by a single locus have been identified experimentally; however, each exon of Pirmy has been also found to be amplified in multiple copies on the Y chromosome. The 28 Pirmy-like RNA variants present various combinations of these exons that are distributed in multiple different loci on the mouse Y chromosome. Morphology- and sperm motility-related abnormalities have been found in two strains of Y-deleted mice with disrupted expression of Pirmy and Pirmy-like RNAs. The Pirmy and Pirmy-like RNAs serve as sources of piRNAs that are complementary to 5′- and 3′-UTRs of several autosomal genes, such as FABP9, Spink2, superoxide dismutase (SOD), and calreticulin, and also genes that presumably contribute to sex ratio maintenance in the progeny. The proteins expressed from these autosomal genes are up-regulated in the sperm of Y-deleted mice and appear to be responsible for the disruption of sperm morphology and motility [152]. In Bombyx mori, females are the heterogametic sex (ZW), and the W chromosome is heterochromatinized and consists almost entirely of transposon sequences. piRNA from the Fem locus on the W chromosome functions as a suppressor of the Masc gene, which regulates sex-specific splicing of the doublesex (dsx) gene, which is necessary for sex determination in many insects [153]. Thus, small piRNAs from the Y or W chromosome can potentially be involved in sex determination, the resolution of intragenomic conflicts, reproductive isolation, and the regulation of gene expression for ensuring spermatogenesis [7][16][142][151][152][153][154]. Y-linked piRNA clusters and their functions in humans remain poorly understood [155][156]. High-throughput sequencing of piRNAs from three human adult testis samples and subsequent data analysis have revealed 28 putative piRNA-cluster candidate regions on the Y [157]. However, among them, only one uni-directional cluster with coordinates chrY:3231747-3235845 contains a significant number of mapped piRNAs (45.4 rpkm). This locus includes remnants of SINE, LINE, and LTR TEs, and has a highly homologous region of the same size on the X chromosome. The remaining 27 piRNA clusters predicted on the Y are rather small and remain unexplored. It should also be noted that due to the high level of heterochromatinization and a large number of repetitive elements, the human Y chromosome is not perfectly assembled, and data about piRNA clusters are not complete.5.4. The Y Chromosome and TEs
Degeneration of the Y chromosome has been accompanied by the acquisition of transposable elements. The old Y chromosome of D. melanogaster is strongly enriched with retrotransposons of different families. Earlier, in some laboratory strains of D. melanogaster, active Gypsy elements restricted to the Y chromosome have been found [158]. Using the latest genome assembly, it has been uncovered that Dm412, Gypsy, Het-A, Doc, TART, Mdg1, Mdg3, blood, and FW TEs are prevalent on the Y chromosome of D. melanogaster [4]. Y chromosomes of the D. simulans clade are similarly enriched in retrotransposons relative to the rest part of the genomes; however, Y chromosomes from even closely related species accumulate distinct TE sets [5]. Whether TE accumulation is coupled with beneficial developmental processes remains to be determined. The stability and non-random localization of TEs throughout the Y speaks in favor of their putative functional role in the host [2][159]. Recently the organization of functional centromeres of D. melanogaster has been resolved in detail, due to the mapping of CENP-A-occupied regions of all chromosomes. It has been found that CENP-A mapped DNA is mainly composed of retrotransposons and is often flanked and inserted by large blocks of satellite repeats [160]. However, satellites are practically not found in the Y centromere, despite the fact that the whole Y chromosome is strongly enriched in simple tandem repeats [4]. For instance, the Y centromere region consists of a tandem array of non-LTR mobile element Jockey-3 [160], and its stability and active state is required for the maintenance of the centromere. The presence of telomere-specific Het-A, TAHRE, and TART non-LTR retrotransposons in the pericentromeric region of the D. melanogaster Y chromosome and in closely related species [2][161] remains mysterious and can potentially be associated with their involvement in telomere maintenance [162]. Recent data indicate that the Y chromosome of D. melanogaster appears to be a cryptic library of active copies of TEs. The preference of TEs for insertion into the Y chromosome can potentially be beneficial for the host, providing immunity against active TEs in the testes by producing piRNAs. However, by analyzing testis single-cell sequencing data, Lawlor and colleagues found an unexpected burst of activity of TEs residing on the Y in early spermatocytes of D. melanogaster [163]. This event occurs during a specific developmental period that coincides with the up-regulation of Y-chromosomal fertility factors and spermatocyte-specific transcription of many Y-linked genes [73], as well as with a decreasing level of several components of the piRNA pathway. Piwi expression is not detected in primary spermatocytes [164][165], nor is the RDC complex [149]. As mentioned above, the RDC complex is responsible for transcription of bi-directional piRNA clusters providing the bulk of piRNAs for the suppression of TEs. Indeed, in primary spermatocytes, the piRNA machinery switches to the production of piRNAs from the non-canonical Su(Ste) and petrel clusters, ensuring the silencing of protein-coding genes [7][142][143]. Note that moderate activation of TEs at this stage of spermatogenesis can, in a certain percentage of cases, lead to transposon insertions and mutations in functional genomic regions. Given that spermatogenesis is a highly redundant process, some TE activity leading to detrimental effects with a low frequency may be inconsequential. Eventually, some mutations can be adaptive for an individual and, will subsequently be fixed in the population. Thus, TE mobilization in this narrow developmental window leads to newly arising genetic variability important for the evolutionary adaptation of the population to changing environmental conditions.6. Conclusions
Despite the fact that in some animal taxa Y chromosomes are completely absent, in most heterosexual eukaryotes Y chromosomes are maintained and perform various essential functions. These include sex determination, ensuring male fertility; correct segregation of meiotic chromosomes; regulation of the activity of rDNA repeats; epigenetic regulation of harmful elements, including TEs and protein-coding genes; contribution to interspecies hybrid sterility, and other responsibilities. The Y chromosome life cycle progresses through a series of stages common to many organisms, such as birth, accumulation of genes necessary for spermatogenesis, cessation of the recombination process, degeneration of the bulk of acquired sequences, and aging. Whereas in mammals the appearance of the Y chromosome has occurred once in a common ancestor of marsupials and placentals before their splitting, presumably 160-180 MYA, in Diptera and some fishes, Y chromosomes have arisen and disappeared several times during their evolutionary history. Studies of model organisms, Drosophila and mice, have fundamental significance for uncovering the shared properties of Y chromosomes of multiple species. The convergent nature of evolution of the Y chromosome allows researchers to consider that the data obtained in model organisms can be useful to a certain extent for the prediction of the human Y chromosome behavior in the future, as well as in understanding how the specific structure of this chromosome reflects its functions in normal and pathological conditions.References
- Bonaccorsi, S.; Lohe, A. Fine mapping of satellite DNA sequences along the Y chromosome of Drosophila melanogaster: Relationships between satellite sequences and fertility factors. Genetics 1991, 129, 177–189.
- Hoskins, R.A.; Carlson, J.W.; Wan, K.H.; Park, S.; Mendez, I.; Galle, S.E.; Booth, B.W.; Pfeiffer, B.D.; George, R.A.; Svirskas, R.; et al. The Release 6 reference sequence of the Drosophila melanogaster genome. Genome Res. 2015, 25, 445–458.
- Carvalho, A.B.; Vicoso, B.; Russo, C.A.; Swenor, B.; Clark, A.G. Birth of a new gene on the Y chromosome of Drosophila melanogaster. Proc. Natl. Acad. Sci. USA 2015, 112, 12450–12455.
- Chang, C.H.; Larracuente, A.M. Heterochromatin-Enriched Assemblies Reveal the Sequence and Organization of the Drosophila melanogaster Y Chromosome. Genetics 2019, 211, 333–348.
- Chang, C.H.; Gregory, L.E.; Gordon, K.E.; Meiklejohn, C.D.; Larracuente, A.M. Unique structure and positive selection promote the rapid divergence of Drosophila Y chromosomes. eLife 2022, 11, e75795.
- Piergentili, R. Multiple roles of the Y chromosome in the biology of Drosophila melanogaster. Sci. World J. 2010, 10, 1749–1767.
- Chen, P.; Kotov, A.A.; Godneeva, B.K.; Bazylev, S.S.; Olenina, L.V.; Aravin, A.A. piRNA-mediated gene regulation and adaptation to sex-specific transposon expression in D. melanogaster male germline. Genes Dev. 2021, 35, 914–935.
- Salz, H.K.; Erickson, J.W. Sex determination in Drosophila: The view from the top. Fly 2010, 4, 60–70.
- Bachtrog, D.; Mank, J.E.; Peichel, C.L.; Kirkpatrick, M.; Otto, S.P.; Ashman, T.L.; Hahn, M.W.; Kitano, J.; Mayrose, I.; Ming, R.; et al. Tree of Sex Consortium. Sex determination: Why so many ways of doing it? PLoS Biol. 2014, 12, e1001899.
- Carvalho, A.B.; Koerich, L.B.; Clark, A.G. Origin and evolution of Y chromosomes: Drosophila tales. Trends Genet. 2009, 25, 270–277.
- Vicoso, B.; Bachtrog, D. Numerous transitions of sex chromosomes in Diptera. PLoS Biol. 2015, 13, e1002078.
- Mahajan, S.; Bachtrog, D. Convergent evolution of Y chromosome gene content in flies. Nat. Commun. 2017, 8, 785.
- Koerich, L.B.; Wang, X.; Clark, A.G.; Carvalho, A.B. Low conservation of gene content in the Drosophila Y chromosome. Nature 2008, 456, 949–951.
- Bachtrog, D.; Mahajan, S.; Bracewell, R. Massive gene amplification on a recently formed Drosophila Y chromosome. Nat. Ecol. Evol. 2019, 3, 1587–1597.
- Bachtrog, D.; Charlesworth, B. Reduced adaptation of a non-recombining neo-Y chromosome. Nature 2002, 416, 323–326.
- Bachtrog, D. The Y Chromosome as a Battleground for Intragenomic Conflict. Trends Genet. 2020, 36, 510–522.
- Carvalho, A.B.; Clark, A.G. Y chromosome of D. pseudoobscura is not homologous to the ancestral Drosophila Y. Science 2005, 307, 108–110.
- Ricchio, J.; Uno, F.; Carvalho, A.B. New Genes in the Drosophila Y Chromosome: Lessons from D. willistoni. Genes 2021, 12, 1815.
- Ohno, S. Sex Chromosomes and Sex-Linked Genes; Monographs on Endocrinology; Springer: Berlin/Heidelberg, Germany, 1967.
- Potrzebowski, L.; Vinckenbosch, N.; Marques, A.C.; Chalmel, F.; Jégou, B.; Kaessmann, H. Chromosomal gene movements reflect the recent origin and biology of therian sex chromosomes. PLoS Biol. 2008, 6, e80.
- Veyrunes, F.; Waters, P.D.; Miethke, P.; Rens, W.; McMillan, D.; Alsop, A.E.; Grützner, F.; Deakin, J.E.; Whittington, C.M.; Schatzkamer, K.; et al. Bird-like sex chromosomes of platypus imply recent origin of mammal sex chromosomes. Genome Res. 2008, 18, 965–973.
- Subrini, J.; Turner, J. Y chromosome functions in mammalian spermatogenesis. eLife 2021, 10, e67345.
- Foster, J.W.; Graves, J.A. An SRY-related sequence on the marsupial x chromosome: Implications for the evolution of the mammalian testis-determining gene. Proc. Natl. Acad. Sci. USA 1994, 91, 1927–1931.
- Graves, J.A. Evolution of the mammalian Y chromosome and sex-determining genes. J. Exp. Zool. 1998, 281, 472–481.
- Larson, E.L.; Kopania, E.E.K.; Good, J.M. Spermatogenesis and the evolution of mammalian sex chromosomes. Trends Genet. 2018, 34, 722–732.
- Ioannidis, J.; Taylor, G.; Zhao, D.; Liu, L.; Idoko-Akoh, A.; Gong, D.; Lovell-Badge, R.; Guioli, S.; McGrew, M.J.; Clinton, M. Primary sex determination in birds depends on DMRT1 dosage, but gonadal sex does not determine adult secondary sex characteristics. Proc. Natl. Acad. Sci. USA 2021, 118, e2020909118.
- Rice, W.R. The accumulation of sexually antagonistic genes as a selective agent promoting the evolution of reduced recombination between primitive sex chromosomes. Evolution 1987, 41, 911–914.
- Rice, W.R. Genetic hitchhiking and the evolution of reduced genetic activity of the Y sex chromosome. Genetics 1987, 116, 161–167.
- Bachtrog, D. Y-chromosome evolution: Emerging insights into processes of Y-chromosome degeneration. Nat. Rev. Genet. 2013, 14, 113–124.
- Charlesworth, B.; Charlesworth, D. The degeneration of Y chromosomes. Philos. Trans. R. Soc. Lond. B. Biol. Sci. 2000, 355, 1563–1572.
- Bellott, D.W.; Hughes, J.F.; Skaletsky, H.; Brown, L.G.; Pyntikova, T.; Cho, T.J.; Koutseva, N.; Zaghlul, S.; Graves, T.; Rock, S.; et al. Mammalian Y chromosomes retain widely expressed dosage-sensitive regulators. Nature 2014, 508, 494–499.
- Lahn, B.T.; Page, D.C. Functional coherence of the human Y chromosome. Science 1997, 278, 675–680.
- Vogt, P.H. AZF deletions and Y chromosomal haplogroups: History and update based on sequence. Hum. Reprod. Update 2005, 11, 319–336.
- Kamp, C.; Huellen, K.; Fernandes, S.; Sousa, M.; Schlegel, P.N.; Mielnik, A.; Kleiman, S.; Yavetz, H.; Krause, W.; Küpker, W.; et al. High deletion frequency of the complete AZFa sequence in men with Sertoli-cell-only syndrome. Mol. Hum. Reprod. 2001, 7, 987–994.
- Lardone, M.C.; Parodi, D.A.; Valdevenito, R.; Ebensperger, M.; Piottante, A.; Madariaga, M.; Smith, R.; Pommer, R.; Zambrano, N.; Castro, A. Quantification of DDX3Y, RBMY1, DAZ and TSPY mRNAs in testes of patients with severe impairment of spermatogenesis. Mol. Hum. Reprod. 2007, 13, 705–712.
- Kotov, A.A.; Olenkina, O.M.; Godneeva, B.K.; Adashev, V.E.; Olenina, L.V. Progress in understanding the molecular functions of DDX3Y (DBY) in male germ cell development and maintenance. Biosci. Trends 2017, 11, 46–53.
- Bhowmick, B.K.; Satta, Y.; Takahata, N. The origin and evolution of human ampliconic gene families and ampliconic structure. Genome Res. 2007, 17, 441–450.
- Skaletsky, H.; Kuroda-Kawaguchi, T.; Minx, P.J.; Cordum, H.S.; Hillier, L.; Brown, L.G.; Repping, S.; Pyntikova, T.; Ali, J.; Bieri, T.; et al. The male-specific region of the human Y chromosome is a mosaic of discrete sequence classes. Nature 2003, 423, 825–837.
- Betrán, E.; Demuth, J.P.; Williford, A. Why chromosome palindromes? Int. J. Evol. Biol. 2012, 2012, 207958.
- Bonito, M.; D’Atanasio, E.; Ravasini, F.; Cariati, S.; Finocchio, A.; Novelletto, A.; Trombetta, B.; Cruciani, F. New insights into the evolution of human Y chromosome palindromes through mutation and gene conversion. Hum. Mol. Genet. 2021, 30, 2272–2285.
- Lange, J.; Skaletsky, H.; van Daalen, S.K.; Embry, S.L.; Korver, C.M.; Brown, L.G.; Oates, R.D.; Silber, S.; Repping, S.; Page, D.C. Isodicentric Y chromosomes and sex disorders as byproducts of homologous recombination that maintains palindromes. Cell 2009, 138, 855–869.
- Hughes, J.F.; Page, D.C. The Biology and Evolution of Mammalian Y Chromosomes. Annu. Rev. Genet. 2015, 49, 507–527.
- Traut, W.; Winking, H. Meiotic chromosomes and stages of sex chromosome evolution in fish: Zebrafish, platyfish and guppy. Chromosome Res. 2001, 9, 659–672.
- Graves, J.A. Sex chromosome specialization and degeneration in mammals. Cell 2006, 124, 901–914.
- Blackmon, H.; Ross, L.; Bachtrog, D. Sex Determination, Sex Chromosomes, and Karyotype Evolution in Insects. J. Hered. 2017, 108, 78–93.
- Griffin, D.K. Is the Y chromosome disappearing?—Both sides of the argument. Chromosome Res. 2012, 20, 35–45.
- Fisher, E.; Scambler, P. Human haploinsufficiency—One for sorrow, two for joy. Nat. Genet. 1994, 7, 5–7.
- Johnson, A.F.; Nguyen, H.T.; Veitia, R.A. Causes and effects of haploinsufficiency. Biol. Rev. 2019, 94, 1774–1785.
- Zug, R. Developmental disorders caused by haploinsufficiency of transcriptional regulators: A perspective based on cell fate determination. Biol. Open. 2022, 11, bio058896.
- Carrel, L.; Brown, C.J. When the Lyon(ized chromosome) roars: Ongoing expression from an inactive X chromosome. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2017, 372, 20160355.
- Fang, H.; Disteche, C.M.; Berletch, J.B. X Inactivation and Escape: Epigenetic and Structural Features. Front. Cell Dev. Biol. 2019, 7, 219.
- Cortez, D.; Marin, R.; Toledo-Flores, D.; Froidevaux, L.; Liechti, A.; Waters, P.D.; Grützner, F.; Kaessmann, H. Origins and functional evolution of Y chromosomes across mammals. Nature 2014, 508, 488–493.
- Hook, E.B.; Warburton, D. Turner syndrome revisited: Review of new data supports the hypothesis that all viable 45, X cases are cryptic mosaics with a rescue cell line, implying an origin by mitotic loss. Hum. Genet. 2014, 133, 417–424.
- Gravholt, C.H.; Viuff, M.H.; Brun, S.; Stochholm, K.; Andersen, N.H. Turner syndrome: Mechanisms and management. Nat. Rev. Endocrinol. 2019, 15, 601–614.
- Stern, C. Untersuchungen über Aberrationen desY-chromosoms von Drosophila melanogaster. Zeitschrift für Induktive Abstammungs-und Vererbungslehre 1929, 51, 253–353. (In German)
- Shen, T.H. Zytologische Untersuchungen über Sterilität bei Männchen von Drosophila melanogaster und bei F1 Männchen der Kreuzung zwischen D. simulans-Weibchen und D. melanogaster-Männchen. Zeitschrift für Zellforschung und Mikroskopische Anatomie 1932, 15, 547–580. (In German)
- Brosseau, G.E. Genetic analysis of the male fertility factors on the Y chromosome of Drosophila melanogaster. Genetics 1960, 45, 257–274.
- Ayles, G.B.; Sanders, T.; Kiefer, B.; Suzuki, D. Temperature sensitive mutations in Drosophila melanogaster: XI. Male sterile mutants of the Y chromosome. Dev. Biol. 1973, 32, 239–257.
- Hardy, R.W.; Tokuyasu, K.T.; Lindsley, D.L. Analysis of spermatogenesis in Drosophila melanogaster bearing deletions for Y-chromosome fertility genes. Chromosoma 1981, 83, 593–617.
- Kennison, J.A. The genetic and cytological organization of the Y chromosome of Drosophila melanogaster. Genetics 1981, 98, 529–548.
- Hazelrigg, T.; Fornili, P.; Kaufman, T.C. A cytogenetic analysis of X-ray induced male steriles on the Y chromosome of Drosophila melanogaster. Chromosoma 1982, 87, 535–559.
- Gatti, M.; Pimpinelli, S. Functional elements in Drosophila melanogaster heterochromatin. Annu. Rev. Genet. 1992, 26, 239–275.
- Zhang, P.; Stankiewicz, R.L. Y-linked male sterile mutations induced by P element in Drosophila melanogaster. Genetics 1998, 150, 735–744.
- Gatti, M.; Pimpinelli, S. Cytological and genetic analysis of the Y chromosome of Drosophila melanogaster. Chromosoma 1983, 88, 349–373.
- Zhang, J.; Luo, J.; Chen, J.; Dai, J.; Montell, C. The Role of Y Chromosome Genes in Male Fertility in Drosophila melanogaster. Genetics 2020, 215, 623–633.
- Carvalho, A.B.; Lazzaro, B.P.; Clark, A.G. Y chromosomal fertility factors kl-2 and kl-3 of Drosophila melanogaster encode dynein heavy chain polypeptides. Proc. Natl. Acad. Sci. USA 2000, 97, 13239–13244.
- Goldstein, L.S.B.; Hardy, R.W.; Lindsley, D.L. Structural genes on the Y chromosome of Drosophila melanogaster. Proc. Natl. Acad. Sci. USA 1982, 79, 7405–7409.
- Gepner, J.; Hays, T.S. A fertility region on the Y chromosome of Drosophila melanogaster encodes a dynein microtubule motor. Proc. Natl. Acad. Sci. USA 1993, 90, 11132–11136.
- Rasmusson, K.; Serr, M.; Gepner, J.; Gibbons, I.; Hays, T.S. A family of dynein genes in Drosophila melanogaster. Mol. Biol. Cell 1994, 5, 45–55.
- Ishikawa, T. Axoneme Structure from Motile Cilia. Cold Spring Harb. Perspect. Biol. 2017, 9, a028076.
- Yu, Z.; Ren, M.; Wang, Z.; Zhang, B.; Rong, Y.S.; Jiao, R.; Gao, G. Highly efficient genome modifications mediated by CRISPR/Cas9 in Drosophila. Genetics 2013, 195, 289–291.
- Hafezi, Y.S.S.; Tarrash, S.R.; Wolfner, M.F.; Clark, A.G. Dissecting fertility functions of Drosophila Y chromosome genes with CRISPR. Genetics 2020, 214, 977–990.
- Mahadevaraju, S.; Fear, J.M.; Akeju, M.; Galletta, B.J.; Pinheiro, M.M.L.S.; Avelino, C.C.; Cabral-de-Mello, D.C.; Conlon, K.; Dell’Orso, S.; Demere, Z.; et al. Dynamic sex chromosome expression in Drosophila male germ cells. Nat. Commun. 2021, 12, 892.
- Fingerhut, J.M.; Moran, J.V.; Yamashita, Y.M. Satellite DNA-containing gigantic introns in a unique gene expression program during Drosophila spermatogenesis. PLoS Genet. 2019, 15, e1008028.
- Fingerhut, J.M.; Yamashita, Y.M. mRNA localization mediates maturation of cytoplasmic cilia in Drosophila spermatogenesis. J. Cell Biol. 2020, 219, e202003084.
- Bonaccorsi, S.; Pisano, C.; Puoti, F.; Gatti, M. Y chromosome loops in Drosophila melanogaster. Genetics 1988, 120, 1015–1034.
- Bonaccorsi, S.; Gatti, M.; Pisano, C.; Lohe, A. Transcription of a satellite DNA on two Y chromosome loops of Drosophila melanogaster. Chromosoma 1990, 99, 260–266.
- Piergentili, R.; Mencarelli, C. Drosophila melanogaster kl-3 and kl-5 Y-loops harbor triple-stranded nucleic acids. J. Cell Sci. 2008, 121, 1605–1612.
- Redhouse, J.L.; Mozziconacci, J.; White, R.A. Co-transcriptional architecture in a Y loop in Drosophila melanogaster. Chromosoma 2011, 120, 399–407.
- Liao, S.E.; Ai, Y.; Fukunaga, R. An RNA-binding protein Blanks plays important roles in defining small RNA and mRNA profiles in Drosophila testes. Heliyon 2018, 4, e00706.
- Zhu, L.; Fukunaga, R. RNA-binding protein Maca is crucial for gigantic male fertility factor gene expression, spermatogenesis, and male fertility, in Drosophila. PLoS Genet. 2021, 17, e1009655.
- Hundertmark, T.; Kreutz, S.; Merle, N.; Nist, A.; Lamp, B.; Stiewe, T.; Brehm, A.; Renkawitz-Pohl, R.; Rathke, C. Drosophila melanogaster tPlus3a and tPlus3b ensure full male fertility by regulating transcription of Y-chromosomal, seminal fluid, and heat shock genes. PLoS ONE 2019, 14, e0213177.
- Cheng, M.H.; Maines, J.Z.; Wasserman, S.A. Biphasic subcellular localization of the DAZL-related protein boule in Drosophila spermatogenesis. Dev. Biol. 1998, 204, 567–576.
- Heatwole, V.M.; Haynes, S.R. Association of RB97D, an RRM protein required for male fertility, with a Y chromosome lampbrush loop in Drosophila spermatocytes. Chromosoma 1996, 105, 285–292.
- Ceprani, F.; Raffa, G.D.; Petrucci, R.; Piergentili, R. Autosomal mutations affecting Y chromosome loops in Drosophila melanogaster. BMC Genet. 2008, 9, 32.
- Hackstein, J.H.; Leoncini, O.; Beck, H.; Peelen, G.; Hennig, W. Genetic fine structure of the Y chromosome of Drosophila hydei. Genetics 1982, 101, 257–277.
- Vogt, P.; Hennig, W. Molecular structure of the lampbrush loops nooses of the Y chromosome of Drosophila hydei: I. The Y chromosome-specific repetitive DNA sequence family ay1 is dispersed in the loop DNA. Chromosoma 1986, 94, 449–458.
- Vogt, P.; Hennig, W. Molecular structure of the lampbrush loops nooses of the Y chromosome of Drosophila hydei: II. DNA sequences with homologies to multiple gemonic locations are major constituents of the loop. Chromosoma 1986, 94, 459–467.
- Wlaschek, M.; Awgulewitsch, A.; Bunemann, H. Structure and function of Y chromosomal DNA. I. Sequence organization and localization of four families of repetitive DNA on the Y chromosome of Drosophila hydei. Chromosoma 1988, 96, 145–158.
- Piergentili, R. Evolutionary conservation of lampbrush-like loops in drosophilids. BMC Cell Biol. 2007, 8, 35.
- Huijser, P.; Beckers, L.; Top, B.; Hermans, M.; Sinke, R.; Hennig, W. Poly·poly is highly transcribed in the testes of Drosophila hydei. Chromosoma 1990, 100, 48–55.
- Trapitz, P.; Glatzer, K.H.; Bunemann, H. Towards a physical map of the fertility genes on the heterochromatic Y chromosome of Drosophila hydei: Families of repetitive sequences transcribed on the lampbrush loops Nooses and Threads are organized in extended clusters of several hundred kilobases. Mol. Gen. Genet. 1992, 235, 221–234.
- Hochstenbach, R.; Brand, R.; Hennig, W. Transcription of repetitive DNA sequences in the lampbrush loop pair Nooses formed by sterile alleles of fertility gene Q on the Y chromosome of Drosophila hydei. Mol. Gen. Genet. 1994, 244, 653–660.
- Shermoen, A.W.; O’Farrell, P.H. Progression of the cell cycle through mitosis leads to abortion of nascent transcripts. Cell 1991, 67, 303–310.
- Koenig, M.; Beggs, A.H.; Moyer, M.; Scherpf, S.; Heindrich, K.; Bettecken, T.; Meng, G.; Müller, C.R.; Lindlöf, M.; Kaariainen, H.; et al. The molecular basis for Duchenne versus Becker muscular dystrophy: Correlation of severity with type of deletion. Am. J. Hum. Genet. 1989, 45, 498–506.
- Pozzoli, U.; Sironi, M.; Cagliani, R.; Comi, G.P.; Bardoni, A.; Bresolin, N. Comparative analysis of the human dystrophin and utrophin gene structures. Genetics 2002, 160, 793–798.
- Le Rumeur, E. Dystrophin and the two related genetic diseases, Duchenne and Becker muscular dystrophies. Bosn. J. Basic Med. Sci. 2015, 15, 14–20.
- Gao, Q.Q.; McNally, E.M. The Dystrophin Complex: Structure, Function, and Implications for Therapy. Compr. Physiol. 2015, 5, 1223–1239.
- Verhaart, I.E.C.; Aartsma-Rus, A. Therapeutic developments for Duchenne muscular dystrophy. Nat. Rev. Neurol. 2019, 15, 373–386.
- Pastinen, T. Genome-wide allele-specific analysis: Insights into regulatory variation. Nat. Rev. Genet. 2010, 11, 533–538.
- Gaur, U.; Li, K.; Mei, S.; Liu, G. Research progress in allele-specific expression and its regulatory mechanisms. J. Appl. Genet. 2013, 54, 271–283.
- McStay, B. Nucleolar dominance: A model for rRNA gene silencing. Genes Dev. 2006, 20, 1207–1214.
- Bayes, J.J.; Malik, H.S. Altered heterochromatin binding by a hybrid sterility protein in Drosophila sibling species. Science 2009, 326, 1538–1541.
- Araripe, L.O.; Tao, Y.; Lemos, B. Interspecific Y chromosome variation is sufficient to rescue hybrid male sterility and is influenced by the grandparental origin of the chromosomes. Heredity 2016, 116, 516–522.
- Durica, D.S.; Krider, H.M. Studies on the ribosomal RNA cistrons in interspecific Drosophila hybrids: I. Nucleolar dominance. Dev. Biol. 1977, 59, 62–74.
- Lewis, M.S.; Cheverud, J.M.; Pikaard, C.S. Evidence for nucleolus organizer regions as the units of regulation in nucleolar dominance in Arabidopsis thaliana interecotype hybrids. Genetics 2004, 167, 931–939.
- Greil, F.; Ahmad, K. Nucleolar dominance of the Y chromosome in Drosophila melanogaster. Genetics 2012, 191, 1119–1128.
- Zhou, J.; Sackton, T.B.; Martinsen, L.; Lemos, B.; Eickbush, T.H.; Hartl, D.L. Y chromosome mediates ribosomal DNA silencing and modulates the chromatin state in Drosophila. Proc. Natl. Acad. Sci. USA 2012, 109, 9941–9946.
- Preuss, S.; Pikaard, C.S. rRNA gene silencing and nucleolar dominance: Insights into a chromosome-scale epigenetic on/off switch. Biochim. Biophys. Acta. 2007, 1769, 383–392.
- Tucker, S.; Vitins, A.; Pikaard, C.S. Nucleolar dominance and ribosomal RNA gene silencing. Curr. Opin. Cell Biol. 2010, 22, 351–356.
- Moss, T.; Stefanovsky, V.Y. At the center of eukaryotic life. Cell 2002, 109, 545–548.
- Long, E.O.; Dawid, I.B. Repeated genes in eukaryotes. Annu. Rev. Biochem. 1980, 49, 727–764.
- Sinclair, D.A.; Guarente, L. Extrachromosomal rDNA circles—A cause of aging in yeast. Cell 1997, 91, 1033–1042.
- Helmrich, A.; Ballarino, M.; Nudler, E.; Tora, L. Transcription-replication encounters, consequences and genomic instability. Nat. Struct. Mol. Biol. 2013, 20, 412–418.
- Lyckegaard, E.M.; Clark, A.G. Evolution of ribosomal RNA gene copy number on the sex chromosomes of Drosophila melanogaster. Mol. Biol. Evol. 1991, 8, 458–474.
- Gibbons, J.G.; Branco, A.T.; Godinho, S.A.; Yu, S.; Lemos, B. Concerted copy number variation balances ribosomal DNA dosage in human and mouse genomes. Proc. Natl. Acad. Sci. USA 2015, 112, 2485–2490.
- Ganley, A.R.; Kobayashi, T. Ribosomal DNA and cellular senescence: New evidence supporting the connection between rDNA and aging. FEMS Yeast Res. 2014, 14, 49–59.
- Kobayashi, T. How does genome instability affect lifespan?: Roles of rDNA and telomeres. Genes Cells 2011, 16, 617–624.
- Eickbush, T.H.; Burke, W.D.; Eickbush, D.G.; Lathe, W.C., III. Evolution of R1 and R2 in the rDNA units of the genus Drosophila. Genetica 1997, 100, 49–61.
- Hawley, R.S.; Marcus, C.H. Recombinational controls of rDNA redundancy in Drosophila. Annu. Rev. Genet. 1989, 23, 87–120.
- Peng, J.C.; Karpen, G.H. H3K9 methylation and RNA interference regulate nucleolar organization and repeated DNA stability. Nat. Cell Biol. 2007, 9, 25–35.
- Warsinger-Pepe, N.; Li, D.; Yamashita, Y.M. Regulation of Nucleolar Dominance in Drosophila melanogaster. Genetics 2020, 214, 991–1004.
- Lu, K.L.; Nelson, J.O.; Watase, G.J.; Warsinger-Pepe, N.; Yamashita, Y.M. Transgenerational dynamics of rDNA copy number in Drosophila male germline stem cells. eLife 2018, 7, e32421.
- McKee, B.D.; Karpen, G.H. Drosophila ribosomal RNA genes function as an X-Y pairing site during male meiosis. Cell 1990, 61, 61–72.
- Watase, G.J.; Yamashita, Y.M. Non-random sister chromatid segregation mediates rDNA copy number maintenance in Drosophila. bioRxiv 2022.
- Yadlapalli, S.; Yamashita, Y.M. DNA asymmetry in stem cells—Immortal or mortal? J. Cell Sci. 2013, 126, 4069–4076.
- Roussel, P.; André, C.; Comai, L.; Hernandez-Verdun, D. The rDNA transcription machinery is assembled during mitosis in active NORs and absent in inactive NORs. J. Cell Biol. 1996, 133, 235–246.
- Héliot, L.; Mongelard, F.; Klein, C.; O’Donohue, M.F.; Chassery, J.M.; Robert-Nicoud, M.; Usson, Y. Nonrandom distribution of metaphase AgNOR staining patterns on human acrocentric chromosomes. J. Histochem. Cytochem. 2000, 48, 13–20.
- Hori, Y.; Shimamoto, A.; Kobayashi, T. The human ribosomal DNA array is composed of highly homogenized tandem clusters. Genome Res. 2021, 31, 1971–1982.
- Le Thomas, A.; Stuwe, E.; Li, S.; Du, J.; Marinov, G.; Rozhkov, N.; Chen, Y.C.; Luo, Y.; Sachidanandam, R.; Toth, K.F.; et al. Transgenerationally inherited piRNAs trigger piRNA biogenesis by changing the chromatin of piRNA clusters and inducing precursor processing. Genes Dev. 2014, 28, 1667–1680.
- Ozata, D.M.; Gainetdinov, I.; Zoch, A.; O’Carroll, D.; Zamore, P.D. PIWI-interacting RNAs: Small RNAs with big functions. Nat. Rev. Genet. 2019, 20, 89–108.
- Ramat, A.; Simonelig, M. Functions of PIWI Proteins in Gene Regulation: New Arrows Added to the piRNA Quiver. Trends Genet. 2021, 37, 188–200.
- Aravin, A.A.; Hannon, G.J.; Brennecke, J. The Piwi-piRNA pathway provides an adaptive defense in the transposon arms race. Science 2007, 318, 761–764.
- Malone, C.D.; Brennecke, J.; Dus, M.; Stark, A.; McCombie, W.R.; Sachidanandam, R.; Hannon, G.J. Specialized piRNA pathways act in germline and somatic tissues of the Drosophila ovary. Cell 2009, 137, 522–535.
- Gainetdinov, I.; Colpan, C.; Arif, A.; Cecchini, K.; Zamore, P.D. A Single Mechanism of Biogenesis, Initiated and Directed by PIWI Proteins, Explains piRNA Production in Most Animals. Mol. Cell 2018, 71, 775–790.e5.
- Brennecke, J.; Aravin, A.A.; Stark, A.; Dus, M.; Kellis, M.; Sachidanandam, R.; Hannon, G.J. Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell 2007, 128, 1089–1103.
- Mohn, F.; Sienski, G.; Handler, D.; Brennecke, J. The Rhino-Deadlock-Cutoff Complex Licenses Noncanonical Transcription of Dual-Strand piRNA Clusters in Drosophila. Cell 2014, 157, 1364–1379.
- Chen, Y.A.; Stuwe, E.; Luo, Y.; Ninova, M.; Le Thomas, A.; Rozhavskaya, E.; Li, S.; Vempati, S.; Laver, J.D.; Patel, D.J.; et al. Cutoff Suppresses RNA Polymerase II Termination to Ensure Expression of piRNA Precursors. Mol. Cell. 2016, 63, 97–109.
- Zhang, Z.; Wang, J.; Schultz, N.; Zhang, F.; Parhad, S.S.; Tu, S.; Vreven, T.; Zamore, P.D.; Weng, Z.; Theurkauf, W.E. The HP1 homolog rhino anchors a nuclear complex that suppresses piRNA precursor splicing. Cell 2014, 157, 1353–1363.
- Wang, K.; Wang, T.; Gao, X.Q.; Chen, X.Z.; Wang, F.; Zhou, L.Y. Emerging functions of piwi-interacting RNAs in diseases. J. Cell. Mol. Med. 2021, 25, 4893–4901.
- Huang, S.; Yoshitake, K.; Asakawa, S. A Review of Discovery Profiling of PIWI-Interacting RNAs and Their Diverse Functions in Metazoans. Int. J. Mol. Sci. 2021, 22, 11166.
- Kotov, A.A.; Adashev, V.E.; Godneeva, B.K.; Ninova, M.; Shatskikh, A.S.; Bazylev, S.S.; Aravin, A.A.; Olenina, L.V. piRNA silencing contributes to interspecies hybrid sterility and reproductive isolation in Drosophila melanogaster. Nucleic Acids Res. 2019, 47, 4255–4271.
- Aravin, A.A.; Naumova, N.M.; Tulin, A.V.; Vagin, V.V.; Rozovsky, Y.M.; Gvozdev, V.A. Double-stranded RNA-mediated silencing of genomic tandem repeats and transposable elements in the D. melanogaster germline. Curr. Biol. 2001, 11, 1017–1027.
- Vagin, V.V.; Sigova, A.; Li, C.; Seitz, H.; Gvozdev, V.; Zamore, P.D. A distinct small RNA pathway silences selfish genetic elements in the germline. Science 2006, 313, 320–324.
- Balakireva, M.D.; Shevelyov, Y.Y.; Nurminsky, D.I.; Livak, K.J.; Gvozdev, V.A. Structural organization and diversification of Y-linked sequences comprising Su(Ste) genes in Drosophila melanogaster. Nucleic Acids Res. 1992, 20, 3731–3736.
- Adashev, V.E.; Kotov, A.A.; Bazylev, S.S.; Shatskikh, A.S.; Aravin, A.A.; Olenina, L.V. Stellate Genes and the piRNA Pathway in Speciation and Reproductive Isolation of Drosophila melanogaster. Front. Genet. 2021, 11, 610665.
- Hurst, L.D. Is Stellate a relict meiotic driver? Genetics 1992, 130, 229–230.
- Aravin, A.A. Pachytene piRNAs as beneficial regulators or a defense system gone rogue. Nat. Genet. 2020, 52, 644–645.
- Chen, P.; Luo, Y.; Aravin, A.A. RDC complex executes a dynamic piRNA program during Drosophila spermatogenesis to safeguard male fertility. PLoS Genet. 2021, 17, e1009591.
- Olenkina, O.M.; Egorova, K.S.; Kibanov, M.V.; Gervaziev, Y.V.; Gvozdev, V.A.; Olenina, L.V. Promoter contribution to the testis-specific expression of Stellate gene family in Drosophila melanogaster. Gene 2012, 499, 143–153.
- Ishizu, H.; Iwasaki, Y.W.; Hirakata, S.; Ozaki, H.; Iwasaki, W.; Siomi, H.; Siomi, M.C. Somatic Primary piRNA Biogenesis Driven by cis-Acting RNA Elements and trans-Acting Yb. Cell Rep. 2015, 12, 429–440.
- Reddy, H.M.; Bhattacharya, R.; Tiwari, S.; Mishra, K.; Annapurna, P.; Jehan, Z.; Praveena, N.M.; Alex, J.L.; Dhople, V.M.; Singh, L.; et al. Y chromosomal noncoding RNAs regulate autosomal gene expression via piRNAs in mouse testis. BMC Biol. 2021, 19, 198.
- Kiuchi, T.; Koga, H.; Kawamoto, M.; Shoji, K.; Sakai, H.; Arai, Y.; Ishihara, G.; Kawaoka, S.; Sugano, S.; Shimada, T.; et al. A single female-specific piRNA is the primary determiner of sex in the silkworm. Nature 2014, 509, 633–636.
- Ellison, C.; Bachtrog, D. Recurrent gene co-amplification on Drosophila X and Y chromosomes. PLoS Genet. 2019, 15, e1008251.
- Sarkar, A.; Maji, R.K.; Saha, S.; Ghosh, Z. piRNAQuest: Searching the piRNAome for silencers. BMC Genom. 2014, 15, 555.
- Özata, D.M.; Yu, T.; Mou, H.; Gainetdinov, I.; Colpan, C.; Cecchini, K.; Kaymaz, Y.; Wu, P.H.; Fan, K.; Kucukural, A.; et al. Evolutionarily conserved pachytene piRNA loci are highly divergent among modern humans. Nat. Ecol. Evol. 2020, 4, 156–168.
- Ha, H.; Song, J.; Wang, S.; Kapusta, A.; Feschotte, C.; Chen, K.C.; Xing, J. A comprehensive analysis of piRNAs from adult human testis and their relationship with genes and mobile elements. BMC Genom. 2014, 15, 545.
- Chalvet, F.; di Franco, C.; Terrinoni, A.; Pelisson, A.; Junakovic, N.; Bucheton, A. Potentially active copies of the gypsy retroelement are confined to the Y chromosome of some strains of Drosophila melanogaster possibly as the result of the female-specific effect of the flamenco gene. J. Mol. Evol. 1998, 46, 437–441.
- Gvozdev, V.A.; Kogan, G.L.; Usakin, L.A. The Y chromosome as a target for acquired and amplified genetic material in evolution. Bioessays 2005, 27, 1256–1262.
- Chang, C.H.; Chavan, A.; Palladino, J.; Wei, X.; Martins, N.M.C.; Santinello, B.; Chen, C.C.; Erceg, J.; Beliveau, B.J.; Wu, C.T.; et al. Islands of retroelements are major components of Drosophila centromeres. PLoS Biol. 2019, 17, e3000241.
- Berloco, M.; Fanti, L.; Sheen, F.; Levis, R.W.; Pimpinelli, S. Heterochromatic distribution of HeT-A- and TART-like sequences in several Drosophila species. Cytogenet. Genome Res. 2005, 110, 124–133.
- Cacchione, S.; Cenci, G.; Raffa, G.D. Silence at the End: How Drosophila Regulates Expression and Transposition of Telomeric Retroelements. J. Mol. Biol. 2020, 432, 4305–4321.
- Lawlor, M.A.; Cao, W.; Ellison, C.E. A transposon expression burst accompanies the activation of Y-chromosome fertility genes during Drosophila spermatogenesis. Nat. Commun. 2021, 12, 6854.
- Cox, D.N.; Chao, A.; Lin, H. piwi encodes a nucleoplasmic factor whose activity modulates the number and division rate of germline stem cells. Development 2000, 127, 503–514.
- Kibanov, M.V.; Kotov, A.A.; Olenina, L.V. Multicolor fluorescence imaging of whole-mount Drosophila testes for studying spermatogenesis. Anal. Biochem. 2013, 436, 55–64.
