1. Background
Polycomb Repressive Complex 2 (PRC2) mediates the deposition of the trimethylation of the lysine 27 of the histone 3 (H3K27me3), a histone modification associated with gene repression in eukaryotes
[1]. PRC2 was first identified in
Drosophila consisting of four core components: Enhancer of zeste (E(z)), a histone methyltransferase unit that catalyses H3K27me3; Extra sex combs (Esc), a WD40 domain protein scaffolding the interactions within the complex; Suppressor of zeste 12 (Su(z)12), a Zinc Finger protein that is essential for binding to nucleosomes; and Nuclear remodeling factor (Nurf55, also called p55), a Trp-Asp (WD) repeat protein involved in nucleosome remodelling
[1][2][1,2]. After discovering PRC2 complexes in
Drosophila as regulators of
Hox genes expression, homologs of PRC2 subunits were identified in plants and other organisms
[3][4][5][3,4,5]. In
Arabidopsis thaliana (
Arabidopsis), there are three E(z) homologs—CURLY LEAF (CLF), SWINGER (SWN) and MEDEA (MEA); three Su(z)12 homologs—EMBRYONIC FLOWER 2 (EMF2), VERNALIZATION 2 (VRN2) and FERTILIZATION-INDEPENDENT SEED 2 (FIS2); a single Esc homolog—FERTILIZATION-INDEPENDENT ENDOSPERM (FIE); and there are five
Arabidopsis homologs of p55 protein—MULTICOPY SUPPRESSOR OF IRA (MSI) 1–5, but MSI1 is the only one demonstrated to be part of the PRC2 complex
[6][7][6,7]. Based on their different subunit compositions, at least three PRC2-like complexes controlling different developmental processes have been described in
Arabidopsis: the EMF, VRN and FIS complexes
[3].
In plants and animals, loss-of-function of core PRC2 subunits results in the abrogation of H3K27me3 levels in PRC2 target genes, which leads to serious developmental defects, highlighting the critical role of PRC2 in development
[7][8][7,8]. In
Arabidopsis chromatin, PRC2 components mimic H3K27me3 localisation
[9]. Genome-wide profiling revealed that 20–25% of
Arabidopsis genes were marked by H3K27me3, and these genes globally display low expression levels
[10][11][12][10,11,12]. Similar percentages of H3K27me3 marked genes were observed in different plant model species (e.g., maize, oilseed rape, rice and
Brachypodium distachyon)
[13][14][15][16][13,14,15,16]. These data further demonstrate the importance of PRC2 activity in regulating the expression of key developmental genes in crops and thereby governing the major agricultural traits, e.g., flowering. Besides PRC2’s pivotal function in controlling development, its key role in the regulation of stress responses and other essential cellular processes, such as metabolism, is emerging
[17][18][19][17,18,19], although still relatively less understood in both plants and animals. Furthermore, the cells perceive the dynamic environmental signals and translate it into differential chromatin and transcriptional states and this is mediated through histone reader proteins that bind to H3K27me3 and/or that affect local chromatin compaction
[18][20][18,20]. In a quest to identify protein reader complexes, two plant-specific H3K27me3 readers, namely EARLY BOLTING IN SHORT DAYS (EBS) and its homolog SHORT LIFE (SHL), were recently discovered and were proposed to act within the PcG pathway causing gene repression
[21][22][21,22]. However, a key question remains unanswered, how do these proteins coordinate their activities with PRC2 to regulate gene expression?
Epigenetic marks such as H3K27me3 can be stably inherited during somatic cell divisions but can be reset during major developmental phase transitions such as the formation of gametes and embryos
[23]. In plants, unique mechanisms exist for the inheritance of H3K27me3 marks compared to their animal counterparts. For instance, a recent study in
Arabidopsis demonstrated a global reduction in H3K27me3 in the paternal germline (i.e., sperm cells), achieved by the coordinated action of three mechanisms: (1) lack of expression of PRC2 histone methyltransferases encoding units such as
CLF,
MEA and
SWN; (2) active removal by Jumonji-C family methylation erasers (histone demethylases); and (3) the global deposition of a sperm cell specific histone H3 variant, H3.10/HTR10, which is resistant to K27 methylation
[24]. Overall, several mechanisms are being elucidated for the transgenerational memory of H3K27me3 in
Arabidopsis, but there is much more yet to be discovered.
In animals, the catalytic and non-catalytic function of PRC2 can be regulated by interaction with protein partners
[25][26][27][25,26,27]. Similarly, plant PRC2 core components are associated with several other proteins, including PRC1 subunits, transcription factors, chromatin-related proteins, the replication machinery, and proteasomal components leading to the modulation of PRC2 activity and/or resulting in its recruitment to target genes
[28]. Essentially, the physical interaction between PRC2 subunits and other proteins helps
the reus
earchers to understand the intricate network of protein–protein interactions that occur to regulate PRC2-mediated gene repression during plant developmental transitions and in response to environmental signals. This re
searchview highlights the protein interactors of the
Arabidopsis PRC2 core subunits identified so far (
Figure 1). Nevertheless, VRN2 and its related VRN-PRC2 complex play a highly specialised role in vernalization-induced flowering that has already been extensively reviewed
[29][30][31][29,30,31]; hence,
thwe
researchers eexcluded its interactors.
The researchers We discuss PRC2 recruitment strategies on target genes mediated by the cooperation with accessory proteins and its associated gene repression and explore the impact of PRC2 interactions especially on the modulation of PRC2 activities.
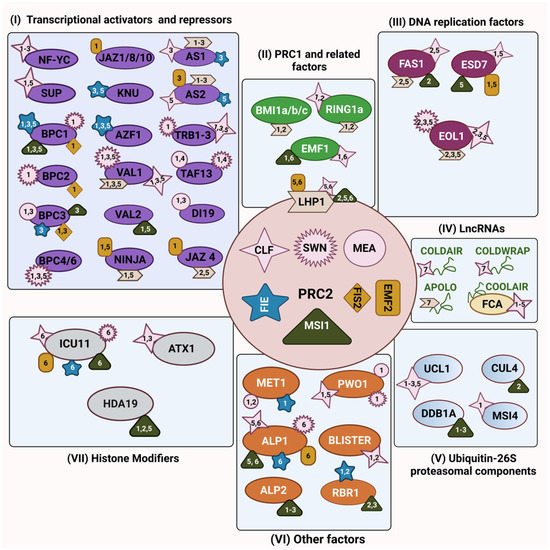
Figure 1. The physical interaction map of the PRC2 complex in Arabidopsis. Each of the PRC2 core components are represented in different shapes enclosed by a circular box in the centre, E(z) homologs are shown in pink colour—CURLY LEAF (CLF) as a four-pointed star, SWINGER (SWN) as a multi-pointed star and MEDEA (MEA) as a circle; Su(z)12 homologs are shown in golden yellow—EMBRYONIC FLOWER 2 (EMF2) as a rectangle, and FERTILIZATION-INDEPENDENT SEED 2 (FIS2) as a rhombus; the ESC homolog—FERTILIZATION-INDEPENDENT ENDOSPERM (FIE)—is represented as a blue five-pointed star; and the p55 protein homolog—MULTICOPY SUPPRESSOR OF IRA 1 (MSI1)—is represented as a dark green triangle. Physical interactors of PRC2 were functionally grouped into six categories: (I) transcriptional activators and repressors (purple); (II) PRC1 and related factors (light green); (III) DNA replication factors (magenta); (IV) long non-coding RNAs (green thread-like structure); (V) ubiquitin-26S proteasomal components (cyan blue); (VI) other factors (orange); and (VII) histone modifiers (grey). Physical interactors from each category may bind to one or more PRC2 components and the numbers (1–6) within the PRC2 component represent different confirmation techniques used for protein–protein interaction studies: 1—yeast two hybrid; 2—pull down assay; 3—biomolecular fluorescence complementation; 4—fluorescence resonance energy transfer; 5—co-immunoprecipitation; 6—co-immunoprecipitation coupled to mass spectrometry; 7—RNA-immunoprecipitation and binding assays. In the figure, LIKE HETEROCHROMATIN PROTEIN 1 (LHP1) is placed at the interface between PRC2 and PRC1. Figure created with BioRender.com (accessed on 21 January 2022).
In the last decade, there has been emerging evidence of the regulation of PRC2 components by their interaction with members of the ubiquitin-26S proteasome, especially with E3 ubiquitin ligases, which facilitate the transfer of ubiquitin to a substrate
[32][33][34][93,94,95]. The ubiquitination of proteins may cause subsequent protein degradation
[35][101], but can also promote changes in the function or activity of the ubiquitinated proteins including chromatin-associated proteins
[36][102]. The ubiquitination of core PRC2 subunits for the control of PRC2 activity and the subsequent protein turnover of PRC2 components will be discussed here.
UPWARD CURLY LEAF1 (UCL1), a plant-specific F-box component of the well-characterised Skp, Cullin, F-box (SCF)-containing E3 ligase complex, physically associates with CLF in the nucleus and subsequently ubiquitinates CLF to target it for degradation via the ubiquitin-26S proteasome pathway. This interaction seems to be quite specific as UCL1 does not interact with MEA
[37][103]. Overexpression of
UCL1 reduces CLF protein levels and alters the expression levels of CLF target genes, suggesting a negative regulation of CLF by UCL1
[37][103]. Moreover, the phenotypes of mutants affected in
UCL1 and
CLF indicate that they may act in the same genetic pathway in which UCL1 may be a negative regulator of CLF
[37][103].
Another multimeric E3 ubiquitin ligase complex contains CULLIN 4 (CUL4), a scaffolding protein, and DAMAGED DNA-BINDING PROTEIN 1 (DDB1), an adaptor protein that associates with the substrate protein and targets it for degradation
[38][104]. In
Arabidopsis, DDB1 physically interacts with MSI1 and CUL4, indicating the possibility for a CUL4–DDB1–MSI1 protein complex
[39][105]. The question was asked as to whether MSI1 could act as a substrate receptor of this E3 ligase complex. The results from two independent studies revealed that MSI1 protein turnover is indeed not under the control of CUL4
[39][40][105,106]. However, when CUL4’s function was compromised, silencing of paternal
MEA was released in the seeds due to the reduction in H3K27me3 levels at this locus and overall
[39][105], pointing to a mediation of CUL4–DDB1 in the activity of the FIS–PRC2 complex. In the
cul4 mutant, there was significant decrease in H3K27me3 levels on
FLC and its downstream target
FT [40][106], further supporting CUL4-DDB1 function in the regulation of PRC2 activities.
CUL4 and DDB1 physically interact with another p55 ortholog, MSI4, and form the CUL4–DDB1–MSI4 complex. MSI4 also interacted with CLF, but not FIE, in Y2H and in planta BiFC assays
[40][106]. Furthermore, loss-of-function mutations of
MSI4 reduce H3K27me3 on
FLC and
FT, resulting in their upregulation and causing a late-flowering phenotype. Therefore, direct regulation of CLF–PRC2 activity by the CUL4–DDB1–MSI4 E3 ubiquitin ligase was plausible
[40][106]. Recently, a plant-specific protein, EMBRYO DEFECTIVE 1579 (EMB1579), implicated in embryo development
[41][107], was demonstrated to recruit and phase condensate CUL4–DDB1–MSI4
[42][108]. In addition, EMB1579 facilitates the physical association of the CUL4–DDB1–MSI4 complex with CLF and contributes to maintaining the proper H3K27me3 levels on
FLC, subsequently controlling flowering
[42][108]. In animals, studies are emerging on how ubiquitination modulates liquid–liquid phase separation of PRCs to mediate large-scale chromatin compaction
[43][109], whereas in plants, there is increasing evidence of the conservation and importance of liquid–liquid phase separation in the organisation of the nuclear space
[44][110]. Understanding the interface among ubiquitination, liquid–liquid phase separation and PRC2 may provide key mechanistic insights into PRC2’s recruitment and dynamics.