In mice and rats, the intronless gene
Dux, encoded by a 4.9 kb retrogene array of more than 28 copies, is located at chromosome 10, whereas the 23
Dux paralog copies in human double homeobox 4 (
DUX4); only 1–12 bases different from mouse
Dux have been identified, in which 14 encoding mRNAs are transcribed in 2-cell-stage embryos as
Dux [
20]. The further study of these 14 mRNAs’ expression regulation will help us to understand the difference between
Dux and its paralog. The intronless orthologs to human
DUX4 are also found in other primates, elephants, hyraxes, and tenrecs. Similar to
Dux, the
DUX4 family also contains a 3.3 kb copies array (D4Z4) located at the distal end of chromosome 4 [
21]. An intron-containing variant, double homeobox B-like (
Duxbl), is only found in rodents, and its pseudogene is found in primates [
21]. The intronless
Dux/
DUX4 was hypothesized to arise from the common intron-containing ancestor in placental mammals through the reverse-transcription and retrotransposition models [
22]. During convergent evolution, environment pressure may further dictate the location of gene insertion. Genomic microsatellite organizations of
Dux and
DUX4 are usually located at the heterochromatinized regions, and their expression is often silenced in most types of cells, including embryonic stem cells [
9,
23]. Epigenetic regulations, such as DNA methylation, are a key determinant for their expression [
23]. Despite the considerable sequence divergence in their DNA-binding domains, DUX and DUX4 shared a more conserved homeodomain 2 (HD2) domain to recognize the 5′-TGA-3′ motif [
24] and activate a subset of genes associated with cleavage-stage embryos [
9]. Interestingly, the swapping of the homeodomain 1 (HD1) and HD2 regions of DUX4 with the corresponding regions from DUX substantially attenuates the activation of Zscan4c and MERVL induced by DUX expression in mouse muscle cells [
25]. Additionally, mouse Dux is myotoxic and shares a partial functional homology with its human paralog DUX4, the aberrant expression of which is linked to facioscapulohumeral muscular dystrophy [
26], which further confirms the conserved roles of Dux/DUX4 in gene regulation.
3. The Regulation of Dux Expression
Dux mRNAs are expressed at the early 2-cell stage in mice during the minor zygotic genome activation (ZGA) stage, whereas it activates several genes during major ZGA, similar to DUX’s homolog DUX4 [
20]. The overexpression of Dux in mESCs resulted in changes in gene expression and endowed these cells with totipotency [
9,
25,
27]. Given its critical role in ZGA and the ESCs to 2CLCs transition, Dux expression must be tightly regulated to ensure the correct developmental progression of cells and tissues. Although transcriptome studies on mouse cells expressing Dux have been widely carried out in vivo and in vitro, the mechanisms for the regulation of
Dux gene expression remain elusive. Thus, we will discuss the recent studies revealing the mechanisms modulating Dux expression, involving transcriptional regulation by transcription factors, epigenetic modifications, signaling pathways and 3D genome conformation.
3.1. Transcriptional Regulation of Dux Expression by Transcription Factors
3.1.1. DPPA2/4
Given that Dux is only expressed in the first or minor wave of ZGA, there must be other factors involved in the precise activation of its expression. By screening epigenetic factors that can increase 2CLCs population in normal ESCs, developmental pluripotency associated 2 (DPPA2) and developmental pluripotency associated 4 (DPPA4) have been identified as upstream factors of Dux that initiate 2C-like transcription [
28]. DPPA2/4 are small putative DNA-binding proteins expressed exclusively in preimplantation embryos and pluripotent cells [
29]. The overexpression of DPPA2/4 can activate an early 2-cell transcriptome, a similar pattern to that seen in mESCs. Additionally, ChIP-seq data reveal that DPPA2/4 can directly bind to the Dux repeats and the promoter region. However, DPPA2 or DPPA4 cannot work alone to activate Dux expression; these two factors must be present in equimolar amounts to transactivate Dux [
30]. Recent studies further revealed that the DPPA2 activity is negatively regulated by a small ubiquitin-like modifier (SUMO) E3 ligasePIAS4, through the SUMOylation of DPPA2, which leads to its degradation (
Figure 1). Either PIAS4 knockout or DPPA2/4 overexpression is sufficient to activate a 2C-like transcriptional program; the expressions of
MERVL and other classic 2C-specific genes, including
Dux, N-acetyltransferase 8 family member 2 (
Cml2),
Zfp352, and zinc finger and SCAN domain containing 4D (
Zscan4d), are then upregulated [
31].
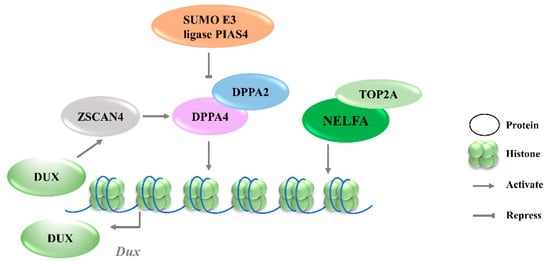
Figure 1. Transcriptional regulation of Dux expression by transcription factors. DPPA2/4 and NELFA transcriptionally activate Dux. Meanwhile, DUX can activate Zscan4 which will upregulate DPPA2/4. PIAS4 will repress DPPA2/4 expression through the SUMOylating of DPPA2. DPPA2/4, developmental pluripotency associated 2/4; PIAS4, protein inhibitor of activated STAT 4; NELFA, negative elongation factor complex member A; ZSCAN4, SCAN domain containing 4; TOP2A, DNA topoisomerase 2a; DUX, double homeobox.
3.1.2. NELFA
Negative elongation factor complex member A (NELFA) was another transcription factor reported to drive the progression to the 2CLCs state by activating Dux [
32]. NELFA is a member of the NELF complex family that regulates RNA polymerase II pausing [
33]. Unlike the enrichment of DPPA2/4 at the Dux locus, ChIP-seq data showed the low enrichment of NELFA at this locus [
34]. However, upon NELFA induction, NELFA located at the Dux locus was responsible for the chromatin opening and the transcriptional activation of Dux. Specifically, the interaction of NELFA with DNA topoisomerase 2a (Top2a) is essential for NELFA to activate Dux; Dux will be silenced in Top2a-deficient cells even when NELFA is overexpressed (
Figure 1) [
32]. The role of NELFA remains controversial, as other studies indicate that NELFA is a direct target of DUX rather than a driver of Dux [
34]. More data from Nelfa knock-out mESCs or embryos will help to clarify the role of NELFA in Dux regulation.
3.1.3. ZSCAN4C
Zinc finger and SCAN domain containing protein 4 C (ZSCAN4C), which shares a similar expression pattern to MERVL in normal mESCs, has also been identified as a 2CLCs marker [
35]. The activation of MERVL by ZSCAN4C is associated with promoting enhancer activity and enhancing histone modification deposition related to gene activation at MERVL LTR loci [
36]. Although Dux activation was observed after the overexpression of Zscan4c, the ChIP-seq data do not show the direct binding of ZSCAN4C in the Dux promoter region, suggesting an indirect transcriptional activation of Dux by ZSCAN4C binding [
36]. However, ZSCAN4 has been demonstrated to facilitate gene expression by inducing global DNA demethylation through silencing the DNA methylation ubiquitin-like components, UHRF1and DNMT1, indicating an additional regulatory layer of Dux by ZSCAN4 (
Figure 2A) [
37]. Moreover, Dux regulators, including DPPA2 and DPPA4, are upregulated by ZSCAN4 overexpression, reflecting that ZSCAN4, DPPA2/4, and DUX may reinforce each other’s expressions and form a positive feedback loop to strengthen 2-cell-like state transition (
Figure 1).
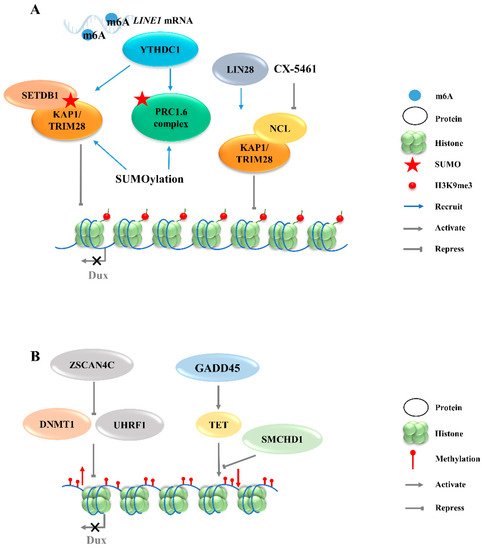
Figure 2. Regulation of Dux expression by epigenetic modifications. (A): Dux expression regulated by histone modifications transcription. The activation of H3K9me3 regulators caused by YTHDC1 and LIN28 will enhance the H3K9me3 levels and silence Dux expression finally. (B): Dux expression regulated by DNA modifications. DNA methylations caused by DNMT1/UHRF1 and TET repress Dux expression. LINE1, long interspersed nuclear element-1; SETDB1, SET domain bifurcated histone lysine methyltransferase 1; KAP1, KRAP-associated transcriptional repressor; TRIM28, tripartite motif-containing protein 28; PRC1.6, polycomb-repressive complexes 1.6; NCL, nucleolin; ZSCAN4C, SCAN domain containing 4C; DNMT1, DNA methyltransferase 1; UHRF1, ubiquitin like with PHD and ring finger domains 1; GADD45, growth arrest and DNA damage 45; TET, ten-eleven translocation; SMCHD1, structural maintenance of chromosomes flexible hinge domain containing 1.
3.2. Regulation of Dux Expression by Epigenetic Modifications
3.2.1. H3K9 Methylation
The totipotent 2CLCs have also been reported to exhibit increased histone modifications in H3K27ac, H3K4me1, and H3K4me3, as compared with ESCs [38].Although these histone modifications are associated with transcriptional activation, no evidence exists to show that Dux expression will be directly regulated by these modifications. The downregulations of chromatin modifiers such as LSD1 and chromatin assembly factor 1 (CAF-1) facilitate MERVL activation [15,39]. Furthermore, MERVL requires lysine (K)-specific demethylase 1A (KDM1A, also as LSD1)—a histone lysine-specific demethylase, a KRAP-associated transcriptional repressor (KAP1), and G9A—a H3K9 histone methyltransferase—for epigenetic repression in normal mESCs [27,40,41]. Likewise, there is no direct evidence showing that the expression of Dux can be regulated by these chromatin modifiers. Recently, LIN28, an RNA-binding protein, was identified as able to repress Dux by an epigenetic program (Figure 2A). H3K9me3 levels were decreased at Dux and its downstream targets, and thus de-repressed Dux expression in Lin28 knockout cells [11]. However, the mechanisms underlying how Lin28 regulates H3K9me3 remain elusive. It is worth noting that Lin28a depletion releases Dux repression by reducing the occupancy of Nucleolin/tripartite motif-containing protein 28 (NCL/TRIM28) in the Dux region. TRIM28 is also known as KAP1, which was demonstrated to repress Dux expression in a long interspersed nuclear element-1 (LINE1)-dependent manner in mESCs [27,42]. LINE1 are Class I transposable elements, which can repress Dux expression by interacting with NCL and KAP1 in mESCs [42]. Mechanistically, after LINE1 RNA is methylated by METTLE3, the m6A-modified LINE1 RNA then works as a scaffold recognized by the YTH domain containing 1 (YTHDC1), which further recruits H3K9me3 regulators, including SET domain bifurcated histone lysine methyltransferase 1 (SETDB1) and KAP1, to the locus of Dux, inhibiting its expression [43,44].Furthermore, polycomb-repressive complexes (PRCs) bind LINE1 RNA and act as an essential partner for Dux gene repression [ 45 ].SUMO modification enhances the H3K9me3 levels on a genome-wide scale, including the Dux locus, and facilitates the recruitment of PRC1.6 and KAP/SETDB1 complexes to the locus to repress Dux gene expression (Figure 2A) [46]. In fact, Ythdc1 depletion results in a global decrease in the SETDB1-mediated H3K9me3 enrichment, which is accompanied by the re-activation of MERVL and Dux [43]. However, Ythdc1-depleted cells still retain the ability to re-activate many retrotransposons upon Dux removal, indicating a parallel regulation pattern between Ythdc1 and Dux with regard to retrotransposon regulation [44]. Due to the sequence differences in LINE1 among species, it is not clear whether LINE1 has similar effects in other mammals, including Homo sapiens [ 47 ]。
3.2.2. Histone Variants
The chromatin assembly factor CAF-1 has been reported to repress MERVL [39], and recent studies revealed its role in establishing the modification of the non-canonical histone variant H3.3, which has been reported to co-enrich with H3K9me3 to silence ERVs in mESCs [48].The knockout of p150, a subunit of the CAF-1 complex, leads to a decrease in the total H3.3 enrichment, accompanied by the upregulation of Dux and MERVL. ChIP-seq data further confirm that H3.3 is enriched at the Dux locus and represses Dux expression [49]. However, the incorporation of H3.3 chaperones HIRA, ATRX or DAXXis not necessary per se for the function of H3.3 in Dux repression, indicating the existence of other H3.3 chaperones that may regulate Dux expression [49].
3.2.3. DNA Methylation
In addition to histone modification, there are additional epigenetic mechanisms that regulate the activity of Dux. Typically occurring at the cytosine in CpG, 5-Methylcytosine (5mC) is a critical modification in the development and differentiation of cell lineages by blocking gene transcription [50]. The structural maintenance of chromosomes flexible hinge domain containing 1 (SMCHD1) cooperates with ten-eleven translocation (TET) proteins to negatively regulate the activities in DNA demethylation (Figure 2B). The removal of SMCHD1 from mESCs induces TET-dependent demethylation, preferentially at SMCHD1 targeting sites, along with the activation of Dux and the Dux pseudogene (Gm4981) [12,、51 ].。_ The受精卵中 siRNA-mediated knockdown of 介导的 Smchd1 in zygotes leads to a continued overexpression of Dux through the 8-cell stage. In addition, the presence of an 敲低导致 Dux 在 8 细胞阶段持续过表达。此外,Dunmethylated state of the Duxx promoter启动子区域在 region in the 2-cell stage indicates that the initial activation of 2 细胞阶段的未甲基化状态的存在表明 Dux DNA is的初始激活是去甲基化依赖性的[ demethylation-dependent [51 , 52 ]. However, the mechanism of re-methylation of the。 Dux然而,在发育后期发生的Dux基因座重新甲基化的机制 locus仍然难以捉摸。还需要进一步的研究来解决 occurring in the later stages of development remains elusive. Further studies are also needed to address whether SMCHD1 can inhibit TET proteins to modulate the DNA demethylation是否可以抑制 TET 蛋白来调节 DNA 去甲基化过程。生长停滞和 process. Growth arrest and DNA damage损伤 45 (GADD45) is another regulator of TET-mediated DNA demethylation.是 TET 介导的 DNA 去甲基化的另一个调节因子。