Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is a comparison between Version 1 by José M. Izquierdo and Version 5 by Jessie Wu.
T-cell intracellular antigen 1 (TIA1) is an RNA-binding protein that is expressed in many tissues and in the vast majority of species, although it was first discovered as a component of human cytotoxic T lymphocytes. TIA1 has a dual localization in the nucleus and cytoplasm, where it plays an important role as a regulator of gene-expression flux. As a multifunctional master modulator, TIA1 controls biological processes relevant to the physiological functioning of the organism and the development and/or progression of several human pathologies.
- TIA1
- gene expression
- biological processes
- pathophysiology
1. T-Cell Intracellular Antigen 1 : Gene, Isoforms and Protein Structure
T-cell intracellular antigen 1 (TIA1) is a member of the RNA-binding protein (RBP) family. It is now 30 years since the first report on the cloning and characterization of TIA1 was published by the laboratory of Paul Anderson and colleagues (Figure 1), who identified it as a component of cytotoxic T lymphocyte granules involved in DNA fragmentation [1]. Since its discovery, many research groups have provided insights into the role of this multifunctional master regulator in human molecular and cellular biology, physiology and pathology, which is reflected by a steadily increasing number of publications in the PubMed database (Figure 1).
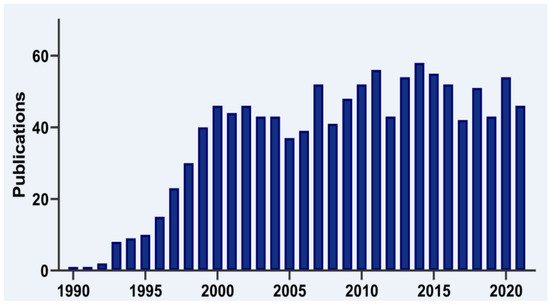
Figure 1. Timeline of TIA1 publications. Histogram representation of the timeline of literature, searching for TIA1/TIA-1 terms in the PubMed/MEDLINE database.
The human TIA1 gene is located on chromosomal region 2p13 and consists of 13 exons that, by alternative splicing, can give rise to two major isoforms. At the structural level, TIA1 is similar to other classical RBPs, consisting of three RNA-recognition motifs (RRMs) and a low-complexity glutamine and asparagine (Q/N)-rich carboxyl-terminal domain, which favors protein-protein interactions [2] (Figure 2). In addition, two ribonucleoprotein-forming consensus peptide sequences are conserved within each RRM: RNP1 (8 amino acids), and RNP2 (6 amino acids) [1][2][1,2] (Figure 2). Of its 13 exons, exons 1–4 encode RRM1, 5–8 RRM2 and 9–11 RRM3, and exons 12 and 13 encode the C-terminal domain and the 3′-untranslated region (3′-UTR) of its mature messenger RNA (mRNA). Exon 5, which encodes 11 amino acids at the beginning of RRM2, is subject to alternative splicing, which gives rise to TIA1a (43 kDa, including the exon) and TIA1b (40 kDa, omitting the exon) isoforms [1][2][1,2] (Figure 2). In the absence of an RNA to bind to, the three RRMs of TIA1 behave as independent structural modules connected by peptide bridges, with almost no intramolecular interaction. Binding to RNA triggers the compaction of the protein domains to form a more rigid RNA-protein complex [3][4][3,4].
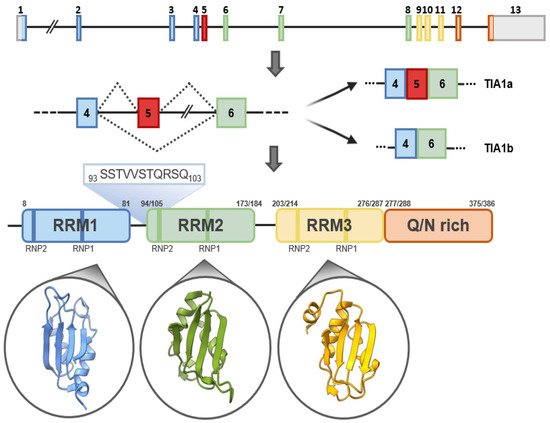
Figure 2. TIA1: gene, isoforms and protein structural details. Schematic representation of human TIA1 gene. Organization of exons and introns of the main isoforms, a and b, generated by alternative splicing, as well as of functional domains of TIA1. TIA1 contains three RNA-recognition motifs (RRMs) and an auxiliary domain rich in asparagine and glutamine (Q/N-rich domain). The amino-acid sequences that differentiate TIA1 a and b isoforms are shown in green. The secondary-tertiary structures of each of the three RRMs are highlighted in the corresponding circles.
The Anderson laboratory later identified TIA1-related/like protein (TIAR/TIAL1), which shares 85% amino-acid sequence similarity with TIA1 in the amino-terminal region but only 51% in the C-terminal region, the expression of which also promotes DNA fragmentation [5][6][5,6].
2. Regulation of Gene Expression
Gene expression in eukaryotic cells is an orderly process comprising: DNA replication, transcription; post-transcriptional processes, such as processing/splicing of pre-mRNAs; and transport, stability and translation of mature mRNAs. Additionally, post-translational regulatory mechanisms modulate the function, stability and fate of proteins. By virtue of its properties of binding to cellular RNA/DNA, TIA1 participates in the regulation of many of these processes. In the presence of RNA, it binds to small fragments of 3–11 uracil-rich ribonucleotides [7][21], but it is also known to have capacity to bind to equivalent DNA sequences through the RRM2 [8][26] .2.1. Transcriptional Rates
TIA1 was first implicated in transcriptional regulation in 2007, when it was described that the C-terminal domain of RNA polymerase II interacted with components of the spliceosome processing factor snRNP U1 and other proteins, including TIA1 [9][27] (Figure 3). These interactions facilitated simultaneous transcription and processing/splicing of cellular pre-mRNAs. Once TIA1 proteins are bound to DNA, RNA sequester them by affinity, facilitating dynamic post-transcriptional regulation [9][27]. The mechanism of action is speculated to involve the slowing of RNA polymerase II [8][26], as well as the end of transcription and associated 3′ end processing [10][28]. Genes such as COL2A1 (procollagen, type II) [11][24], IGFBP-3 (insulin-like growth-factor binding protein-3) [12][29] or testicular PACAP (pituitary adenylate cyclase-activating polypeptide) [13][30] seem to be preferentially regulated by this pathway. A comprehensive analysis of the transcriptome of HeLa cells with transient reduction in TIA protein expression revealed changes in a large number of mRNAs associated with cellular processes such as inflammation, cell signaling, suppression of immune response, angiogenesis, apoptosis, metabolism and cell proliferation [14][31]. In this context, the participation of these proteins in the direct modulation of PTGS2 (prostaglandin-endoperoxide synthase 2) and IL-6 (interleukin 6) gene transcription was demonstrated [14][31]. Similar results were obtained in the study of the transcriptome of mouse neuronal tissues (spinal cord and cerebellum) in the absence of TIA1 expression [15][32].
2.2. Post-Transcriptional Control
One of the nuclear functions of TIA1 is to regulate the alternative splicing (and also constitutive splicing) of some pre-mRNAs to favor the inclusion [16][17][18][19][19,33,34,35] or selective exclusion [20][36] of exons (Figure 3). For splicing to occur, the spliceosome must first be assembled, which begins with the recognition of the 5′ intronic splicing site (5′ss) by the nuclear ribonucleoprotein U1 (snRNP U1). It is in this context that TIA1 exerts its function. To promote exon inclusion, TIA1 binds to uridine-rich RNA sequences located near both constitutive and alternative 5′ss with weakly conserved consensus sequences for U1 snRNA hybridization and U1 snRNP anchoring and subsequently binds to the U1-C protein of the U1 snRNP [21][16][17][18][19][20][22][23][24][13,19,33,34,35,36,37,38,39]. In this process, RRM2 binds to the uridine-rich regions near the 5′ss of the intron, favored by RRM3, and the Q/N-rich domain binds to the N-terminal region of the U1-C subunit of the U1 snRNP by an RRM1-favored process [23][24][38,39]. Its role as a splicing repressor is linked to competition for uridine-rich sequences within the polypyrimidine-rich regions located at positions near the 3′ss of introns to interfere with auxiliary-factor (perhaps U2AF65) binding by preventing U2 snRNP recruitment [19][20][35,36].
The cellular relevance of TIA1 in the regulation of alternative and constitutive splicing was confirmed through the establishment of a binding map of these regulators to RNA in HeLa cells using in vivo irradiation of cells with ultraviolet light and subsequent immunoprecipitation of RNA-protein complexes (termed iCLIP) [16][19]. Results from this seminal study have been corroborated by recent data in other cell lines using PARCLIP and eCLIP approaches [18][19][34,35]. These approaches demonstrate that TIA1 preferentially binds to ARE sequences located in introns, in 5′ and especially 3′ UTR positions––preferably UUUUA or AUUUU tandem pentamers––of mRNAs, as well as to non-coding RNAs. From these studies, it is estimated that TIA1 modulates gene expression of 2–5% of the human genome.
2.3. Regulation of Translation
TIA1 can modulate cellular translation by limiting the availability of the ribosomal machinery and the translational efficiency of specific cellular mRNAs, either under conditions of stress, to safeguard cell viability, or under conditions of homeostasis [16][25][26][27][28][29][19,40,41,42,43,44] (Figure 3).
In the absence of stress, 48S translation initiation complexes, which include the ternary translation-initiator complex eIF2-GTP-tRNAMet, the small subunit of the ribosome, other translation initiation factors (eIFs) and mRNA, are formed in the cell. When translation is initiated, eIF5 causes hydrolysis of eIF2-associated GTP. The 60S subunit of the ribosome then binds, displacing some eIFs, and the 80S ribosome is formed. However, under stress, the α-subunit of eIF2 is phosphorylated by a family of kinases such that eIF2B cannot exchange GDP for GTP, eIF2-GTP-tRNAMet levels decrease and translation pre-initiator complexes cannot form properly. At this point, TIA1 associates with the complex formed by eIF4E, eIF4G, eIF4A, eIF4B, eIF3 and the small subunit of ribosome in an anomalous 48S complex. These inactive translation pre-initiator complexes associated with mRNAs accumulate in the cytoplasm and, through the aggregation properties of the Q/N-rich domain of TIA1 and PABP (Poly(A) Binding Protein) proteins, bind together, forming large foci of mRNAs and proteins termed stress granules (SGs) [30][31][32][33][45,46,47,48]. The formation of SGs favors cell survival in stressful situations, such as during viral infection, fasting or limited amino-acid availability, and oxidative or osmotic stress. In these adverse conditions, the cell enters a state of biological quiescence, inhibiting general translation, which allows energy to be conserved to repair the damage produced by the stressful insult. For example, cells respond to changes in nutrient conditions by regulating the expression of the so-called 5′TOP (5′-Terminal OligoPyrimidine tract) mRNAs, which encode factors for protein biosynthesis, including many ribosomal protein-coding mRNAs. These mRNAs contain an oligopyrimidine (UC)-rich sequence at the 5′-UTR end. It has been suggested that TIA1, through GCN2 activation, binds to these sequences and facilitates mRNA arrest in the SGs under fasting conditions. Thus, reprogramming of the translational machinery and protein expression ensues, favoring cell survival [34][35][49,50], although these claims are controversial [36][37][51,52]. On the other hand, there are other biological processes for which the formation of SGs is relevant from a physiological perspective––for example, in the differentiation of naïve CD4+ T lymphocytes to cytokine-secreting effector cells. After priming of naïve lymphocytes, the requirements for differentiation and rapid clonal expansion are so high that they trigger nutrient deficiencies and induce stress. Under these conditions, eIF2α is phosphorylated, and cytokine mRNAs accumulate in SGs. Stimulation of lymphocytes leads to dephosphorylation of eIF2α, the disappearance of SGs and the reactivation of cytokine synthesis [38][53].
SGs are dynamic structures, and most of their components are in a constant flux of assembly and disassembly [39][40][41][54,55,56]. SGs are formed approximately 15 min after the onset of the stressful stimulus, and their formation is reversible, disappearing within a few hours (between 2 and 6 h) of the cessation of the stimulus, provided that it is not lethal [30][31][39][45,46,54]. Microscopic observation of SGs shows that their size is variable and cell-type-dependent, ranging from 150 to 200 nm in stressed rat hippocampal cells [30][45]; 0.1–0.2 µm in COS cells [42][57]; or 1–2 µm in HeLa or HEK293 cells [41][56]. There is some controversy regarding the involvement of TIA1 and/or TIAR in SG formation. It has been suggested that both TIAR and TIA1 are needed [42][57], yet other authors have shown that the presence of one of the two proteins is sufficient for SG generation [27][42]. In fact, the results of high-resolution fluorescence microscopy, together with fluorescence recovery after photobleaching experiments, suggest that SGs have two different layers with different components, dynamics and functions: a stable inner structure or core surrounded by a less dense layer or shell. Accordingly, the components of the structure that make up the core would be less dynamic, whereas the components of the shell would be more dynamic. This hypothesis allows individuals to propose two models to explain the SG assembly process. The first model suggests that SG formation begins with the assembly of the riboproteomic core, followed by rapid growth or assembly of the envelope. The first small SGs would fuse to generate larger SGs through liquid-liquid phase-separation (LLPS) processes. Alternatively, the second model proposes that it is the LLPS states promoted by the formation of translationally repressed riboproteomic complexes that form first, subsequently promoting the separation of droplets that constitute cores in the context of elevated protein concentrations [43][44][58,59]. TIA1 would be one of the constituents of the nuclei of SGs [45][60].
There are many proteins (i.e., RBPs, non-RNA-binding proteins and translation-initiation factors) that have been identified as components of SGs, in addition to RNAs (coding and non-coding). These include the HuR protein (Hu antigen R/Embryonic lethal, abnormal vision, Drosophila-like 1) [45][60] and some proteins involved in RNA stability/degradation, such as TDP-43 (TAR DNA-binding protein) [46][47][61,62], hSMG-1 (phosphatidylinositol 3-kinase-related kinase) [48][63] and TTP (TrisTetraProlin) [45][60]. The presence of these proteins alongside TIA1 implies a model of equilibrium between antagonistic factors capable of promoting the degradation or stability of mRNAs located in SGs. Finally, the role of G3BP1 and 2 (GTPase-activating protein 1 and 2) in the formation of some SGs is noteworthy. This protein does not interact with TIA1 per se but contains a self-aggregation domain and is functionally similar to TIA1. An immunoprecipitation study of RNA-protein complexes identified several hundred TIA1-associated mRNAs under heat-shock stress conditions. The study revealed the binding of TIA1 to uridine (U)-rich motifs at 5′ ends and to adenosine-uridine (AU) at 3′ ends of the 3′-UTR region of mRNAs, such as ACTB (actin, beta), CALM2 (calmodulin 2) or CASP7 (caspase 7), among others [27][42]. Similarly, analysis of TIAR-associated mRNAs under UV stress conditions, without apparent phosphorylation of eIF2α, demonstrated the repression of translation of some translation initiation factors, such as EIF4A, eIF4E and eIF1B, as well as the transcription factor C-MYC, which would favor the inhibition of translation once the stress is relieved, causing the inhibition of global translation of the cell [49][64]. The involvement of TIA1 in the regulation of mitochondrial cytochrome c mRNA translation [27][28][42,43] of the transcription factor HIF-1α (hypoxia inducible factor 1, alpha subunit) [50][65] and of the molecular chaperone Hsp70 (heat-shock protein 70) [51][66] has also been demonstrated. These results have been confirmed by iCLIP/PARCLIP/eCLIP analysis of TIA1 in other cell models [16][18][19][19,34,35]. It appears, however, that selective translational activation of some mRNAs excluded from SGs occurs under stress conditions [30][41][52][45,56,67].
TIA1 has the ability to globally and/or specifically inhibit the translation of cellular mRNAs in stress-independent conditions by binding to specific sequences located in the 5′- and/or 3′-UTR region of mRNAs [16][19]. The first evidence of this phenomenon was documented in activated macrophages in which deficiency of TIA1 led to an increase in the levels of the pro-inflammatory protein TNFα (tumor necrosis factor α) without affecting the relative levels of its mRNA [25][53][54][40,68,69]. This occurred through the binding of TIA1 to AU-rich sequences in the 3′UTR region of TNFα mRNA [53][55][68,70], a process that has also been described for other cellular mRNAs [16][19][26][19,35,41].
2.4. Stability of mRNAs
mRNA turnover is the process by which an mRNA is degraded before or after translation. This can occur at the 5′ end of the mRNA through decapping by the activity of DCP1 and DCP2 (dipeptidyl carboxypeptidase 1 and 2) and the exonuclease XRN1 (5′-3′ exoribonuclease 1) [56][71] or at the 3′ end, mediated by the exosome, resulting in deadenylation of the poly(A) tail and recruitment of exonucleases [57][72]. The mRNA regions involved are the ARE sequences located in the 3′-UTR that favor the binding of TIA1, AUF1 (AU-rich element RNA-binding protein 1), KRSP and TTP, which recruit proteins involved in the degradation process. Binding of TIA1 to these regions favors both mRNA deadenylation and stimulation of 5′cap removal, in a cell-specific and stress-independent manner [19][35]. By contrast, proteins such as HuR stabilize mRNA, likely because of their inability to recruit the exosome to ARE sequences [58][73] (Figure 3). mRNA stability can also be regulated through the binding of microRNAs (miRNAs), which can trigger degradation. miRNAs are small fragments of non-coding RNA, of 19–24 nucleotides, that regulate gene expression by pairing with sequences typically located in the 3′ UTR regions of mRNAs. The interaction of miRNA with RNA stimulates the recruitment of the RISC complex (RNA-induced silencing complex) and mRNA cleavage. Several studies suggest that 20–30% of gene expression is controlled by miRNAs [59][74]. A large-scale expression-platform study demonstrated that transient knockdown of TIA1 and TIAR in HeLa cells resulted in the overexpression of 29 miRNAs [60][75]. This result was interpreted as a strategy to counteract the differential expression and cellular phenotypes associated with the downregulation of TIA proteins [60][75]. Moreover, in cellular microvesicles abundantly expressing TIA1 and representing a mechanism of cellular communication in stem cells, 365 miRNAs were identified, the target mRNAs of which are related to organism development, survival and differentiation, as well as regulation of the immune response [61][76]. This evidence suggests that TIA1 can activate or repress the transcription of miRNAs through a mechanism that is currently unknown and can interact with them to modulate gene expression. Finally, it is worth noting that non-coding RNAs have been identified, both small RNAs and long, non-coding RNAs, that repress TIA1-associated expression through direct interaction with its mRNA, impacting its translation and/or stability (e.g., miR-19a [62][77], miR-487a [63][78], mivaRNAI-138 [64][79]) or the direct sequestration of TIA1 and other RNAs through a molecular sponge mechanism (e.g., FLJ11812) [65][80], NORAD [66][81], MALAT1 [67][82]).
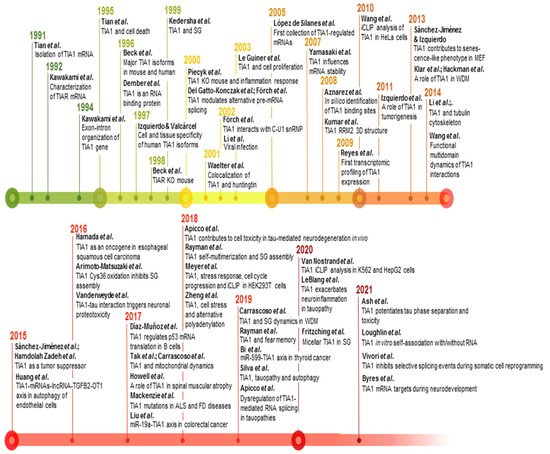

Figure 3. Timeline and milestones of TIA1 research. Milestones in the study of TIA1/TIA-1 found in PubMed/MEDLINE database. The references to build this figure are the following: [1][2][3][4][5][6][21][16][68][7][11][14][17][18][19][20][22][23][25][27][29][31][58][62][69][70][71][72][73][74][75][76][77][78][79][80][81][82][83][84][85][86][87][88][89][90][91][92][93][94][95][96][97][98][99][100][1,2,3,4,5,6,13,19,20,21,24,31,33,34,35,36,37,38,40,42,44,46,73,77,83,84,85,86,87,88,89,90,91,92,93,94,95,96,97,98,99,100,101,102,103,104,105,106,107,108,109,110,111,112,113,114].
