Autophagy is a widely conserved process in eukaryotes that is involved in a series of physiological and pathological events, including development, immunity, neurodegenerative disease, and tumorigenesis. It is regulated by nutrient deprivation, energy stress, and other unfavorable conditions through multiple pathways. In general, autophagy is synergistically governed at the RNA and protein levels. The upstream transcription factors trigger or inhibit the expression of autophagy- or lysosome-related genes to facilitate or reduce autophagy. Moreover, a significant number of non-coding RNAs (microRNA, circRNA, and lncRNA) are reported to participate in autophagy regulation. Finally, post-transcriptional modifications, such as RNA methylation, play a key role in controlling autophagy occurrence.
- autophagy
- regulatory mechanisms
- transcription
- ncRNA
- RNA methylation
1. Introduction
2. Regulation of Autophagy by Transcriptional and Post-Transcriptional Modifications
2.1. Transcription Factors Regulate Autophagy at RNA Level
| Transcription Factor | Function |
|---|---|
| Leucine zipper transcription factors (MiT/TFE) | MiT/TFE recognize promoters of lysosomal and Atg genes and represent transcriptional controllers of lysosomal biogenesis and autophagy [4][13]. |
| Nuclear receptors PPARα and FXR |
PPARα and FXR oppositely control the expression of Atg7, Beclin1, Bnip3 Atg7, Beclin1, Bnip3, and LC3 and autophagic vesicle formation [6]. LC3 and autophagic vesicle formation [6]. |
| Small heterodimer partner (SHP) |
SHP decreases mRNA levels of Atg genes and inhibits autophagy [21]. |
| Transcription factors FOXO/FOXA | Activation of FOXO/FOXA induces the expression of multiple Atg genes and lysosomal genes [17][32]. |
| CCAAT/enhancer binding protein beta (C/EBPβ) | C/EBPβ targets key Atg genes and induces the expression of Atg genes [23][33]. |
| Activating transcription factor 4(ATF4) |
ATF4 is involved in the cellular stress response and autophagosome formation [5][34]. |
| Nuclear factor-kappa B (NF-κB) |
NF-κB activates the expression of Atg genes and induces autophagy [28][35]. |
| Zinc-finger-family DNA-binding protein, ZKSCAN3 | ZKSCAN3 decreases mRNA levels of Atg genes and inhibits autophagy [16]. |
| Tumor suppressor p53 | In the nucleus, P53 transactivates Atg genes and induces autophagy by inhibiting mTOR; in the cytoplasm, P53 suppresses autophagy [36][37]. |
| Signal transducer and activator of transcription (STAT) | STAT3 phosphorylation upregulates BNIP3 expression; STAT1 suppresses the expression of Atg genes [38][39]. |
| Transcription factor E2F | Activation of E2F1 upregulates the expression of Atg genes [24]. |
| TGA9 (TGACG motif-binding protein 9) | TGA9 activates autophagy by upregulating the expression of Atg genes [31]. |
| E93 | Knockdown of E93 reduces the expression of several Atg genes in B. mori [40]. |
| EcR-USP | 20E-EcR-USP upregulates the transcription of Atg genes to induce autophagy [11]. |
2.2. Regulation of Autophagy by Non-Coding RNAs
| Non-Coding RNAs | Target Genes | Species | Impact on Autophagy |
|---|---|---|---|
| miR30b | Atg12, Beclin-1 | Helicobacter pylori |
↓[58] |
| miR-17 | Ulk1 | Mouse | ↓[59] |
| miR-30a | Beclin1, Atg12, Atg5 | Mouse | ↓[44][60] |
| miR-188-3p | Atg7 | Mouse | ↓[61] |
| miR-93, miR106b, miR142-3p | ULK1, ATG16L | Human | ↓[44][62][63] |
| miR-101 | ATG4D, LC3 | Human | ↓[43][44] |
| miR-155 | ATG3 | Human | ↓[64] |
| miR-214-3p | ATG5, ATG12 | Human | ↓[65] |
| miR-216b | BECLIN1 | Human | ↓[66] |
| miR-103a-3p | ATG5 | Human | ↓[45] |
| miR-183, miR-204 | LC3B1/LC3-II | Human | ↓[44][67] |
| miR-83, miR-29 | atg-4.2 / ATG4D, ATG9a | Caenorhabditis elegans/Human | ↓[8][46] |
| miR-34 | Atg9a/ATG9a | Caenorhabditis elegans/Human | ↓[47] |
| miR-4459 | ATG13 | Human | ↓[68] |
| miR-23b | ATG12 | Human | ↓[69] |
| miR-19a | BECLIN1, LC3 | Human | ↓[70] |
| miR-376b | ATG4C, BECLIN1 | Human | ↓[44] |
| miR-15a, miR-16 | Rictor (mTORC1) | Human | ↑[71] |
| circNF1-419 | Dynamin-1 | Mouse | ↑[50] |
| circHIPK2 | ATG5, BECLIN1-1 | Human | ↑[72] |
| circPABPN1 | ATG16l1 | Human | ↓[51] |
| lncRNA APF | Atg7 | Mouse | ↑[61] |
| lncRNA NEAT1, lncRNA XIST | Atg9a | Mouse | ↑[8][53] |
| lncRNA HAGLROS | PI3K-AKT-NF-κB | Human | ↑[73] |
| lncRNA TGFB2-OT1 | ATG3, ATG7, ATG13 | Human | ↑[74] |
| lncRNA CA7-4 | AMPK | Human | ↑[75] |
| lncRNA GBCDRlnc1 | BECLIN1, ATG5, ATG12 | Human | ↑[54][55] |
| lncRNA MALAT1 | Beclin1, LC3 | Mouse | ↓[76] |
| lncRNA LINC00470 | BECLIN1, ATG3, ATG7 | Human | ↓[77] |
| lncRNA CTA | Unknown | Human | ↓[78] |
| lncRNA HOTAIR | BECLIN1, LC3, ATG3, ATG7 | Human | ↓[57] |
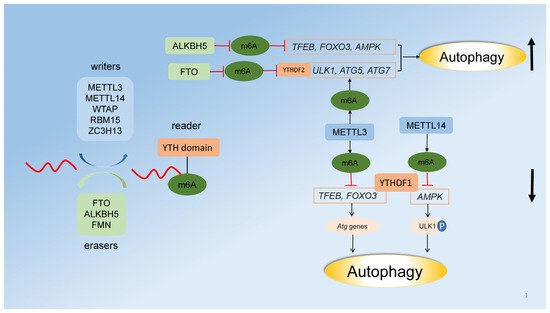
2.3. Regulation of Autophagy by RNA Methylation
References
- Levine, B.; Kroemer, G. Autophagy in the Pathogenesis of Disease. Cell 2008, 132, 27–42.
- Fullgrabe, J.; Klionsky, D.J.; Joseph, B. The return of the nucleus: Transcriptional and epigenetic control of autophagy. Nat. Rev. Mol. Cell Bio. 2014, 15, 65–74.
- Cardenal-Muñoz, E.; Arafah, S.; López-Jiménez, A.T.; Kicka, S.; Falaise, A.; Bach, F.; Schaad, O.; King, J.S.; Hagedorn, M.; Soldati, T. Mycobacterium marinum antagonistically induces an autophagic response while repressing the autophagic flux in a TORC1- and ESX-1-dependent manner. PLoS Pathog. 2017, 13, e1006344.
- Dai, Y.; Li, K.; Wu, W.; Wu, K.; Yi, H.; Li, W.; Xiao, Y.; Zhong, Y.; Cao, Y.; Tian, L. Steroid hormone 20-hydroxyecdysone induces the transcription and complex assembly of V-ATPases to facilitate autophagy in Bombyx mori. Insect Biochem. Molec. 2020, 116, 103255.
- Luhr, M.; Torgersen, M.L.; Szalai, P.; Hashim, A.; Brech, A.; Staerk, J.; Engedal, N. The kinase PERK and the transcription factor ATF4 play distinct and essential roles in autophagy resulting from tunicamycin-induced ER stress. J. Biol. Chem. 2019, 294, 8197–8217.
- Lee, J.M.; Wagner, M.; Xiao, R.; Kim, K.H.; Feng, D.; Lazar, M.A.; Moore, D.D. Nutrient-sensing nuclear receptors coordinate autophagy. Nature 2014, 516, 112–115.
- Rodriguez-Muela, N.; Germain, F.; Marino, G.; Fitze, P.S.; Boya, P. Autophagy promotes survival of retinal ganglion cells after optic nerve axotomy in mice. Cell Death Differ. 2012, 19, 162–169.
- Kong, Y.; Huang, T.; Zhang, H.; Zhang, Q.; Ren, J.; Guo, X.; Fan, H.; Liu, L. The lncRNA NEAT1/miR-29b/Atg9a axis regulates IGFBPrP1-induced autophagy and activation of mouse hepatic stellate cells. Life Sci. 2019, 237, 116902.
- Feng, Y.; Klionsky, D.J. Autophagic membrane delivery through ATG9. Cell Res. 2017, 27, 161–162.
- Conte, A.; Paladino, S.; Bianco, G.; Fasano, D.; Gerlini, R.; Tornincasa, M.; Renna, M.; Fusco, A.; Tramontano, D.; Pierantoni, G.M. High mobility group A1 protein modulates autophagy in cancer cells. Cell Death Differ. 2017, 24, 1948–1962.
- Tian, L.; Ma, L.; Guo, E.; Deng, X.; Ma, S.; Xia, Q.; Cao, Y.; Li, S. 20-Hydroxyecdysone upregulates Atg genes to induce autophagy in the Bombyx fat body. Autophagy 2013, 9, 1172–1187.
- Wu, W.; Luo, M.; Li, K.; Dai, Y.; Yi, H.; Zhong, Y.; Cao, Y.; Tettamanti, G.; Tian, L. Cholesterol derivatives induce dephosphorylation of the histone deacetylases Rpd3/HDAC1 to upregulate autophagy. Autophagy 2021, 17, 512–528.
- Annunziata, I.; van de Vlekkert, D.; Wolf, E.; Finkelstein, D.; Neale, G.; Machado, E.; Mosca, R.; Campos, Y.; Tillman, H.; Roussel, M.F.; et al. MYC competes with MiT/TFE in regulating lysosomal biogenesis and autophagy through an epigenetic rheostat. Nat. Commun. 2019, 10, 1–18.
- Pastore, N.; Vainshtein, A.; Herz, N.J.; Huynh, T.; Brunetti, L.; Klisch, T.J.; Mutarelli, M.; Annunziata, P.; Kinouchi, K.; Brunetti-Pierri, N.; et al. Nutrient-sensitive transcription factors TFEB and TFE3 couple autophagy and metabolism to the peripheral clock. EMBO J. 2019, 38, e101347.
- Settembre, C.; De Cegli, R.; Mansueto, G.; Saha, P.K.; Vetrini, F.; Visvikis, O.; Huynh, T.; Carissimo, A.; Palmer, D.; Klisch, T.J.; et al. TFEB controls cellular lipid metabolism through a starvation-induced autoregulatory loop. Nat. Cell Biol. 2013, 15, 647–658.
- Chauhan, S.; Goodwin, J.G.; Chauhan, S.; Manyam, G.; Wang, J.; Kamat, A.M.; Boyd, D.D. ZKSCAN3 Is a Master Transcriptional Repressor of Autophagy. Mol. Cell 2013, 50, 16–28.
- Zhao, J.; Brault, J.J.; Schild, A.; Cao, P.; Sandri, M.; Schiaffino, S.; Lecker, S.H.; Goldberg, A.L. FoxO3 coordinately activates protein degradation by the autophagic/lysosomal and proteasomal pathways in atrophying muscle cells. Cell Metab. 2007, 6, 472–483.
- Milan, G.; Romanello, V.; Pescatore, F.; Armani, A.; Paik, J.; Frasson, L.; Seydel, A.; Zhao, J.; Abraham, R.; Goldberg, A.L.; et al. Regulation of autophagy and the ubiquitin–proteasome system by the FOXO transcriptional network during muscle atrophy. Nat. Commun. 2015, 6, 6670.
- Zhang, N.; Zhao, Y. Other Molecular Mechanisms Regulating Autophagy. In Advances in Experimental Medicine and Biology; Qin, Z.H., Ed.; Science Press: Beijing, China, 2019; Volume 1206, pp. 260–270.
- Shin, H.R.; Kim, H.; Oh, S.; Lee, J.; Kee, M.; Ko, H.; Kweon, M.; Won, K.; Baek, S.H. AMPK–SKP2–CARM1 signalling cascade in transcriptional regulation of autophagy. Nature 2016, 534, 553–557.
- Kim, D.; Kwon, S.; Byun, S.; Xiao, Z.; Park, S.; Wu, S.; Chiang, C.; Kemper, B.; Kemper, J.K. Critical role of RanBP2-mediated SUMOylation of Small Heterodimer Partner in maintaining bile acid homeostasis. Nat. Commun. 2016, 7, 12179.
- Byun, S.; Kim, Y.C.; Zhang, Y.; Kong, B.; Guo, G.; Sadoshima, J.; Ma, J.; Kemper, B.; Kemper, J.K. A postprandial FGF19-SHP-LSD1 regulatory axis mediates epigenetic repression of hepatic autophagy. EMBO J. 2017, 36, 1755–1769.
- Ahmed, M.; Lai, T.H.; Hwang, J.S.; Zada, S.; Pham, T.M.; Kim, D.R. Transcriptional Regulation of Autophagy Genes via Stage-Specific Activation of CEBPB and PPARG during Adipogenesis: A Systematic Study Using Public Gene Expression and Transcription Factor Binding Datasets. Cells 2019, 8, 1321.
- Polager, S.; Ofir, M.; Ginsberg, D. E2F1 regulates autophagy and the transcription of autophagy genes. Oncogene 2008, 27, 4860–4864.
- Mu, N.; Lei, Y.; Wang, Y.; Wang, Y.; Duan, Q.; Ma, G.; Liu, X.; Su, L. Inhibition of SIRT1/2 upregulates HSPA5 acetylation and induces pro-survival autophagy via ATF4-DDIT4-mTORC1 axis in human lung cancer cells. Apoptosis 2019, 24, 798–811.
- Liu, H.; Wang, J.; Li, S. E93 predominantly transduces 20-hydroxyecdysone signaling to induce autophagy and caspase activity in Drosophila fat body. Insect Biochem. Molec. 2014, 45, 30–39.
- Demontis, F.; Perrimon, N. FOXO/4E-BP signaling in Drosophila muscles regulates organism-wide proteostasis during aging. Cell 2010, 143, 813–825.
- Liu, Y.; Gordesky-Gold, B.; Leney-Greene, M.; Weinbren, N.L.; Tudor, M.; Cherry, S. Inflammation-Induced, STING-Dependent Autophagy Restricts Zika Virus Infection in the Drosophila Brain. Cell Host Microbe 2018, 24, 57–68.
- Tettamanti, G.; Casartelli, M. Cell death during complete metamorphosis. Philos. Trans. R. Soc. Lond. B 2019, 374, 20190065.
- Li, X.; Qian, X.; Lu, Z. Local histone acetylation by ACSS2 promotes gene transcription for lysosomal biogenesis and autophagy. Autophagy 2017, 13, 1790–1791.
- Wang, P.; Nolan, T.M.; Yin, Y.; Bassham, D.C. Identification of transcription factors that regulate ATG8 expression and autophagy in Arabidopsis. Autophagy 2020, 16, 123–139.
- Lapierre, L.R.; Kumsta, C.; Sandri, M.; Ballabio, A.; Hansen, M. Transcriptional and epigenetic regulation of autophagy in aging. Autophagy 2015, 11, 867–880.
- Guo, L.; Huang, J.; Liu, Y.; Li, X.; Zhou, S.; Qian, S.; Liu, Y.; Zhu, H.; Huang, H.; Dang, Y.; et al. Transactivation of Atg4b by C/EBPβ Promotes Autophagy To Facilitate Adipogenesis. Mol. Cell Biol. 2013, 33, 3180–3190.
- Fawcett, T.W.; Martindale, J.L.; Guyton, K.Z.; Hai, T.; Holbrook, N.J. Complexes containing activating transcription factor (ATF)/cAMP-responsive-element-binding protein (CREB) interact with the CCAAT/enhancer-binding protein (C/EBP)-ATF composite site to regulate Gadd153 expression during the stress response. Biochem. J. 1999, 339, 135–141.
- Copetti, T.; Bertoli, C.; Dalla, E.; Demarchi, F.; Schneider, C. p65/RelA ModulatesBECN1 Transcription and Autophagy. Mol. Cell Biol. 2009, 29, 2594–2608.
- Feng, Z.; Zhang, H.; Levine, A.J.; Jin, S. The coordinate regulation of the p53 and mTOR pathways in cells. Proc. Natl. Acad. Sci. USA 2005, 102, 8204–8209.
- Tasdemir, E.; Maiuri, M.C.; Galluzzi, L.; Vitale, I.; Djavaheri-Mergny, M.; D’Amelio, M.; Criollo, A.; Morselli, E.; Zhu, C.; Harper, F.; et al. Regulation of autophagy by cytoplasmic p53. Nat. Cell Biol. 2008, 10, 676–687.
- You, L.; Wang, Z.; Li, H.; Shou, J.; Jing, Z.; Xie, J.; Sui, X.; Pan, H.; Han, W. The role of STAT3 in autophagy. Autophagy 2015, 11, 729–739.
- McCormick, J.; Suleman, N.; Scarabelli, T.M.; Knight, R.A.; Latchman, D.S.; Stephanou, A. STAT1 deficiency in the heart protects against myocardial infarction by enhancing autophagy. J. Cell Mol. Med. 2012, 16, 386–393.
- Liu, X.; Dai, F.; Guo, E.; Li, K.; Ma, L.; Tian, L.; Cao, Y.; Zhang, G.; Palli, S.R.; Li, S. 20-Hydroxyecdysone (20E) Primary Response Gene E93 Modulates 20E Signaling to Promote Bombyx Larval-Pupal Metamorphosis. J. Biol. Chem. 2015, 290, 27370–27383.
- Xu, Z.; Yan, Y.; Qian, L.; Gong, Z. Long non-coding RNAs act as regulators of cell autophagy in diseases (Review). Oncol. Rep. 2017, 37, 1359–1366.
- Zhao, Y.; Wang, Z.; Zhang, W.; Zhang, L. Non-coding RNAs regulate autophagy process via influencing the expression of associated protein. Prog. Biophys. Mol. Biol. 2020, 151, 32–39.
- Frankel, L.B.; Wen, J.; Lees, M.; Hoyer-Hansen, M.; Farkas, T.; Krogh, A.; Jaattela, M.; Lund, A.H. microRNA-101 is a potent inhibitor of autophagy. EMBO J. 2011, 30, 4628–4641.
- Fu, L.; Wen, X.; Bao, J.; Liu, B. MicroRNA-modulated autophagic signaling networks in cancer. Int. J. Biochem. Cell Biol. 2012, 44, 733–736.
- Zhang, C.; Wang, H.; Qi, Y.; Kan, Y.; Ge, Z. Effects of miR-103a-3p on the autophagy and apoptosis of cardiomyocytes by regulating Atg5. Int. J. Mol. Med. 2019, 43, 1951–1960.
- Zhou, Y.; Wang, X.; Song, M.; He, Z.; Cui, G.; Peng, G.; Dieterich, C.; Antebi, A.; Jing, N.; Shen, Y. A secreted microRNA disrupts autophagy in distinct tissues of Caenorhabditis elegans upon ageing. Nat. Commun. 2019, 10, 4827.
- Yang, J.; Chen, D.; He, Y.; Meléndez, A.; Feng, Z.; Hong, Q.; Bai, X.; Li, Q.; Cai, G.; Wang, J.; et al. MiR-34 modulates Caenorhabditis elegans lifespan via repressing the autophagy gene atg9. Age 2013, 35, 11–22.
- Wei, D.M.; Jiang, M.T.; Lin, P.; Yang, H.; Dang, Y.W.; Yu, Q.; Liao, D.Y.; Luo, D.Z.; Chen, G. Potential ceRNA networks involved in autophagy suppression of pancreatic cancer caused by chloroquine diphosphate: A study based on differentially expressed circRNAs, lncRNAs, miRNAs and mRNAs. Int. J. Oncol. 2019, 54, 600–626.
- Memczak, S.; Jens, M.; Elefsinioti, A.; Torti, F.; Krueger, J.; Rybak, A.; Maier, L.; Mackowiak, S.D.; Gregersen, L.H.; Munschauer, M.; et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 2013, 495, 333–338.
- Diling, C.; Yinrui, G.; Longkai, Q.; Xiaocui, T.; Yadi, L.; Xin, Y.; Guoyan, H.; Ou, S.; Tianqiao, Y.; Dongdong, W.; et al. Circular RNA NF1-419 enhances autophagy to ameliorate senile dementia by binding Dynamin-1 and Adaptor protein 2 B1 in AD-like mice. Aging 2019, 11, 12002–12031.
- Li, X.X.; Xiao, L.; Chung, H.K.; Ma, X.X.; Liu, X.; Song, J.L.; Jin, C.Z.; Rao, J.N.; Gorospe, M.; Wang, J.Y. Interaction between HuR and circPABPN1 Modulates Autophagy in the Intestinal Epithelium by Altering ATG16L1 Translation. Mol. Cell Biol. 2020, 40.
- Palanisamy, K.; Tsai, T.H.; Yu, T.M.; Sun, K.T.; Yu, S.H.; Lin, F.Y.; Wang, I.K.; Li, C.Y. RNA-binding protein, human antigen R regulates hypoxia-induced autophagy by targeting ATG7/ATG16L1 expressions and autophagosome formation. J. Cell Physiol. 2019, 234, 7448–7458.
- Xie, Z.Y.; Wang, F.F.; Xiao, Z.H.; Liu, S.F.; Lai, Y.L.; Tang, S.L. Long noncoding RNA XIST enhances ethanol-induced hepatic stellate cells autophagy and activation via miR-29b/HMGB1 axis. Iubmb Life 2019, 71, 1962–1972.
- Cai, Q.; Wang, S.; Jin, L.; Weng, M.; Zhou, D.; Wang, J.; Tang, Z.; Quan, Z. Long non-coding RNA GBCDRlnc1 induces chemoresistance of gallbladder cancer cells by activating autophagy. Mol. Cancer 2019, 18, 1–16.
- Qian, X.; Li, X.; Cai, Q.; Zhang, C.; Yu, Q.; Jiang, Y.; Lee, J.; Hawke, D.; Wang, Y.; Xia, Y.; et al. Phosphoglycerate Kinase 1 Phosphorylates Beclin1 to Induce Autophagy. Mol. Cell 2017, 65, 917–931.
- Xu, S.; Wang, P.; Zhang, J.; Wu, H.; Sui, S.; Zhang, J.; Wang, Q.; Qiao, K.; Yang, W.; Xu, H.; et al. Ai-lncRNA EGOT enhancing autophagy sensitizes paclitaxel cytotoxicity via upregulation of ITPR1 expression by RNA-RNA and RNA-protein interactions in human cancer. Mol. Cancer 2019, 18, 1–18.
- Wang, X.; Liu, W.; Wang, P.; Li, S. RNA interference of long noncodingRNA HOTAIR suppresses autophagy and promotes apoptosis and sensitivity to cisplatin in oral squamous cell carcinoma. J. Oral. Pathol. Med. 2018, 47, 930–937.
- Tang, B.; Li, N.; Gu, J.; Zhuang, Y.; Li, Q.; Wang, H.; Fang, Y.; Yu, B.; Zhang, J.; Xie, Q.; et al. Compromised autophagy byMIR30B benefits the intracellular survival ofHelicobacter pylori. Autophagy 2014, 8, 1045–1057.
- Wu, H.; Wang, F.; Hu, S.; Yin, C.; Li, X.; Zhao, S.; Wang, J.; Yan, X. MiR-20a and miR-106b negatively regulate autophagy induced by leucine deprivation via suppression of ULK1 expression in C2C12 myoblasts. Cell Signal. 2012, 24, 2179–2186.
- Hu, J.; Meng, Y.; Zhang, Z.; Yan, Q.; Jiang, X.; Lv, Z.; Hu, L. MARCH5 RNA promotes autophagy, migration, and invasion of ovarian cancer cells. Autophagy 2017, 13, 333–344.
- Wang, K.; Liu, C.; Zhou, L.; Wang, J.; Wang, M.; Zhao, B.; Zhao, W.; Xu, S.; Fan, L.; Zhang, X.; et al. APF lncRNA regulates autophagy and myocardial infarction by targeting miR-188-3p. Nat. Commun. 2015, 6, 6779.
- Li, W.; Yang, Y.; Ba, Z.; Li, S.; Chen, H.; Hou, X.; Ma, L.; He, P.; Jiang, L.; Li, L.; et al. MicroRNA-93 Regulates Hypoxia-Induced Autophagy by Targeting ULK1. Oxid. Med. Cell Longev. 2017, 2017, 2709053.
- Zhai, Z.; Wu, F.; Dong, F.; Chuang, A.Y.; Messer, J.S.; Boone, D.L.; Kwon, J.H. Human autophagy geneATG16L1 is post-transcriptionally regulated byMIR142-3p. Autophagy 2014, 10, 468–479.
- Etna, M.P.; Sinigaglia, A.; Grassi, A.; Giacomini, E.; Romagnoli, A.; Pardini, M.; Severa, M.; Cruciani, M.; Rizzo, F.; Anastasiadou, E.; et al. Mycobacterium tuberculosis-induced miR-155 subverts autophagy by targeting ATG3 in human dendritic cells. PLoS Pathog. 2018, 14, e1006790.
- Wang, J.; Wang, W.N.; Xu, S.B.; Wu, H.; Dai, B.; Jian, D.D.; Yang, M.; Wu, Y.T.; Feng, Q.; Zhu, J.H.; et al. MicroRNA-214-3p: A link between autophagy and endothelial cell dysfunction in atherosclerosis. Acta Physiol. 2018, 222, e12973.
- Chen, L.; Han, X.; Hu, Z.; Chen, L. The PVT1/miR-216b/Beclin-1 regulates cisplatin sensitivity of NSCLC cells via modulating autophagy and apoptosis. Cancer Chemoth. Pharm. 2019, 83, 921–931.
- Jian, X.; Xiao-yan, Z.; Bin, H.; Yu-feng, Z.; Bo, K.; Zhi-nong, W.; Xin, N. MiR-204 regulate cardiomyocyte autophagy induced by hypoxia-reoxygenation through LC3-II. Int. J. Cardiol. 2011, 148, 110–112.
- Ge, D.; Han, L.; Huang, S.; Peng, N.; Wang, P.; Jiang, Z.; Zhao, J.; Su, L.; Zhang, S.; Zhang, Y.; et al. Identification of a novel MTOR activator and discovery of a competing endogenous RNA regulating autophagy in vascular endothelial cells. Autophagy 2014, 10, 957–971.
- Pan, B.; Feng, B.; Chen, Y.; Huang, G.; Wang, R.; Chen, L.; Song, H. MiR-200b regulates autophagy associated with chemoresistance in human lung adenocarcinoma. Oncotarget 2015, 6, 32805–32820.
- Yang, L.; Wang, H.; Shen, Q.; Feng, L.; Jin, H. Long non-coding RNAs involved in autophagy regulation. Cell Death Dis. 2017, 8, e3073.
- Huang, N.; Wu, J.; Qiu, W.; Lyu, Q.; He, J.; Xie, W.; Xu, N.; Zhang, Y. MiR-15a and miR-16 induce autophagy and enhance chemosensitivity of Camptothecin. Cancer Biol. Ther. 2015, 16, 941–948.
- Huang, R.; Zhang, Y.; Han, B.; Bai, Y.; Zhou, R.; Gan, G.; Chao, J.; Hu, G.; Yao, H. Circular RNA HIPK2 regulates astrocyte activation via cooperation of autophagy and ER stress by targeting MIR124-2HG. Autophagy 2017, 13, 1722–1741.
- Liu, M.; Han, T.; Shi, S.; Chen, E. Long noncoding RNA HAGLROS regulates cell apoptosis and autophagy in lipopolysaccharides-induced WI-38 cells via modulating miR-100/NF-κB axis. Biochem. Biophys. Res. Commun. 2018, 500, 589–596.
- Huang, S.; Lu, W.; Ge, D.; Meng, N.; Li, Y.; Su, L.; Zhang, S.; Zhang, Y.; Zhao, B.; Miao, J. A new microRNA signal pathway regulated by long noncoding RNA TGFB2-OT1 in autophagy and inflammation of vascular endothelial cells. Autophagy 2015, 11, 2172–2183.
- Zhao, X.; Su, L.; He, X.; Zhao, B.; Miao, J. Long noncoding RNA CA7-4 promotes autophagy and apoptosis via sponging MIR877-3P and MIR5680 in high glucose-induced vascular endothelial cells. Autophagy 2020, 16, 70–85.
- Hu, H.; Wu, J.; Yu, X.; Zhou, J.; Yu, H.; Ma, L. Long non-coding RNA MALAT1 enhances the apoptosis of cardiomyocytes through autophagy inhibition by regulating TSC2-mTOR signaling. Biol. Res. 2019, 52.
- Liu, C.; Fu, H.; Liu, X.; Lei, Q.; Zhang, Y.; She, X.; Liu, Q.; Liu, Q.; Sun, Y.; Li, G.; et al. LINC00470 Coordinates the Epigenetic Regulation of ELFN2 to Distract GBM Cell Autophagy. Mol. Ther. 2018, 26, 2267–2281.
- Wang, Z.; Liu, Z.; Wu, S. Long non-coding RNA CTA sensitizes osteosarcoma cells to doxorubicin through inhibition of autophagy. Oncotarget 2017, 8, 31465–31477.
- Chen, J.; Wang, C.; Fei, W.; Fang, X.; Hu, X. Epitranscriptomic m6A modification in the stem cell field and its effects on cell death and survival. Am. J. Cancer Res. 2019, 9, 752–764.
- Yang, Y.; Hsu, P.J.; Chen, Y.S.; Yang, Y.G. Dynamic transcriptomic m6A decoration: Writers, erasers, readers and functions in RNA metabolism. Cell Res. 2018, 28, 616–624.
- Liu, S.; Li, Q.; Li, G.; Zhang, Q.; Zhuo, L.; Han, X.; Zhang, M.; Chen, X.; Pan, T.; Yan, L.; et al. The mechanism of m6A methyltransferase METTL3-mediated autophagy in reversing gefitinib resistance in NSCLC cells by β-elemene. Cell Death Dis. 2020, 11, 1–14.
- Lin, Z.; Niu, Y.; Wan, A.; Chen, D.; Liang, H.; Chen, X.; Sun, L.; Zhan, S.; Chen, L.; Cheng, C.; et al. RNA m6A methylation regulates sorafenib resistance in liver cancer through FOXO3-mediated autophagy. EMBO J. 2020, 39, e103181.
- Song, H.; Feng, X.; Zhang, H.; Luo, Y.; Huang, J.; Lin, M.; Jin, J.; Ding, X.; Wu, S.; Huang, H.; et al. METTL3 and ALKBH5 oppositely regulate m6A modification of TFEB mRNA, which dictates the fate of hypoxia/reoxygenation-treated cardiomyocytes. Autophagy 2019, 15, 1419–1437.
- Jin, S.; Zhang, X.; Miao, Y.; Liang, P.; Zhu, K.; She, Y.; Wu, Y.; Liu, D.; Huang, J.; Ren, J.; et al. m6A RNA modification controls autophagy through upregulating ULK1 protein abundance. Cell Res. 2018, 28, 955–957.
- Wang, C.; Lin, T.; Ho, M.; Yeh, J.; Tsai, M.; Hung, K.; Hsieh, I.; Wen, M. Regulation of autophagy in leukocytes through RNA N6-adenosine methylation in chronic kidney disease patients. Biochem. Biophys. Res. Commun. 2020, 527, 953–959.
- Wang, X.; Wu, R.; Liu, Y.; Zhao, Y.; Bi, Z.; Yao, Y.; Liu, Q.; Shi, H.; Wang, F.; Wang, Y. m6A mRNA methylation controls autophagy and adipogenesis by targeting Atg5 and Atg7. Autophagy 2020, 16, 1221–1235.
- Yang, S.; Wei, J.; Cui, Y.; Park, G.; Shah, P.; Deng, Y.; Aplin, A.E.; Lu, Z.; Hwang, S.; He, C.; et al. m6A mRNA demethylase FTO regulates melanoma tumorigenicity and response to anti-PD-1 blockade. Nat. Commun. 2019, 10, 2782.
- Chen, Y.; Wang, J.; Xu, D.; Xiang, Z.; Ding, J.; Yang, X.; Li, D.; Han, X. m6A mRNA methylation regulates testosterone synthesis through modulating autophagy in Leydig cells. Autophagy 2021, 17, 457–475.
