Human noroviruses (HuNoVs) are the leading causative agents of epidemic and sporadic acute gastroenteritis that affect people of all ages worldwide. They are responsible for over 20% of all the AGE cases annually. Among susceptible individuals, it has been shown that very low inoculum doses are sufficient to generate a full course of infection with high titers of virus shed in feces. Very few dose–response studies have been carried out to determine the median infectious dose of HuNoVs. Here, we evaluated the median infectious dose (ID50) and diarrhea dose (DD50) of the GII.4/2003 variant of HuNoV (Cin-2) in the gnotobiotic pig model of HuNoV infection and disease. Using different mathematical approaches (classical, and contemporary methods), we estimated the ID50 and DD50 to be between 2400–3400 RNA copies, and 21,000–38,000 RNA copies, respectively. Contemporary dose–response models offer greater flexibility and accuracy in estimating ID50. In contrast to classical methods of endpoint estimation, dose–response modelling allows seamless analyses of data that may include inconsistent dilution factors between doses or numbers of subjects per dose group, or small numbers of subjects. Although this investigation is consistent with state-of-the-art ID50 determinations and offers an advancement in clinical data analysis, it is important to underscore that such analyses remain confounded by pathogen aggregation. Regardless, challenging virus strain ID50 determination is crucial for identifying the true infectiousness of HuNoVs and for the accurate evaluation of protective efficacies in pre-clinical studies of therapeutics, vaccines and other prophylactics using this reliable animal model.
- Gnotobiotic pig model,
- human norovirus,
- dose response
- ID50
- DD50
- Approximate Beta-Poisson
1. Introduction
Human noroviruses (HuNoVs) are non-enveloped RNA viruses with a positive-sense single-stranded genome that belong in the Caliciviridae family. They are the most common etiological agents of epidemic and sporadic acute gastroenteritis in people of all age groups [1][2][3] and are responsible for around 125 million cases of foodborne illnesses worldwide each year [4]. HuNoVs are transmitted through a fecal-oral route and epidemics primarily occur most in healthcare facilities, restaurants, and schools among other settings [5]. HuNoV outbreaks primarily occur due to individuals coming in contact with contaminated food and water [6][7][8].
2. Classification
NoVs are single-stranded positive sense RNA viruses that belong to the genus Norovirus within the family Caliciviridae. NoV genome is composed of 3 open reading frames (ORFs). NoV classification is based on the genetic sequence of the entire virus genome or, the sequence of an individual virus gene [9]. Each genogroup is further sub-classified into genotypes based on phylogenetic analysis of the capsid (VP1) and the RNA dependent RNA polymerase (RdRp) protein sequences [10]. NoV is currently categorized into 10 genogroups (G1-GX) which are further classified into 49 genotypes based on VP1 gene [11]. There are 29 different virus strains spread over 3 different genogroups (genogroup I, II, and IV), that are responsible for AGE in humans [9][12][13].
3. Structure of NoV
The NoV capsid is made up of two structural proteins, VP1 (major) and VP2 (minor), coded by ORF 2 and 3 respectively. VP1 is divided into a shell (S) domain and the protruding (P) domain that provide form and stability to the virus [14]. The VP1 has been extensively studied for its antigenic properties and ability to be easily produced using different in vitro expression systems. The VP1 has been observed to self-assemble leading to the formation of virus-like particles (VLPs) that have been the basis of NoV vaccines [15]. Recently, using a feline calicivirus model, the VP2 has been discovered to function as trans-capsid channel that is responsible for the delivery of viral genome upon host cell binding [16].
4. Pathogenesis of NoV
Over 80% of all HuNoV-associated GE around the world is caused by GII.4 NoVs [17]. The absence of a cell culture system has delayed the progress made on the understanding of NoV pathogenesis and immunity and the development of effective vaccines or therapeutics against NoV disease in humans [18]. The site of replication for NoVs has not been well established, but it is assumed that they replicate in the upper intestinal tract. Biopsies collected from the jejunum of volunteers who develop gastrointestinal illness following an oral administration of the Norwalk (GI.1) or Hawaii virus (GII.1) have exhibited histopathologic lesions [19]. Jones et al, reported that a human-derived continuous B-cell line, BJAB, is capable of supporting HuNoV replication [20]. Infectivity of a GII.4 HuNoV in the cell culture system was stimulated by the presence of enteric bacteria or HBGAs, which were proposed to facilitate attachment to the B cells. The same group also showed that murine norovirus (MNV) were capable of replicating in continuous B-cell lines (M12 and WEHI-231). The poor repeatability and low viral titers yielded from using the BJAB cell line gave rise to new challenges that need to be overcome in order to establish a successful cell culture model. This theory of B-cells being permissive for HuNoV infections was tested in human patients with severe combined immunodeficiency (SCID). Two groups, those with intact (but sometimes dysfunctional) B cells (n = 10) and those without (n = 8), and all were tested for the presence of NoV RNA in stool. The viral RNA titers were similar between the two groups, leading to the conclusion that NoV infections do occur in the absence of B-cells [21]. In a study involving severe combined immunodeficiency (SCID) Gn pigs that were deficient in B cells (and in T cells and NK cells by RAG2/IL2RG gene knockout), after an oral inoculation with HuNoV, fecal virus shedding was observed for a longer duration and the virus-tissue distribution was wider compared to the wildtype Gn pigs [22]. This could either mean that B-cells are not the primary targets of HuNoV infection or that HuNoVs are capable of infecting a broader range of cell-types similar to the findings in RVs. The results of this study were further supported by data recently published by another group that showed no difference between rates and severity of NoV infection among SCID and non-SCID Gn pigs indicating that the innate immune mechanism might be crucial for NoV clearance [23].
Successful NoV cell-culture has been achieved using a stem-cell derived human enteroid system. Bile acid and the presence of HBGA receptors on the cell surfaces were identified to be key determinants of virus entry and propagation [24]. Enteroids derived from all three segments of the small intestine (duodenum, jejunum, and ileum) were permissive for NoV replication up to 96 hours post inoculation. Immunocytochemistry analysis on intestinal biopsies collected from a transplant patient who was suffering from severe GII.4 GE revealed that epithelial cells are the primary site of replication for NoV [25]. In another study, duodenal biopsies taken from an individual suffering from common variable immune deficiency who was chronically infected with HuNoV also showed viral antigens (capsid proteins and RdRp) within enterocytes [26].
5. Correlates of protection against NoV infections
Current knowledge on immune responses to HuNoV comes predominantly from experimental human infection models and clinical studies with VLP vaccines. Similar to some RV species as mentioned earlier, HBGAs are host susceptibility determinants for NoV infections. HBGA found on the surface of intestinal epithelial cells of humans with the secretor phenotype [27] have been determined as key regulators of susceptibility to HuNoV binding and entry in humans [28][29]. Furthermore, the ABO blood group antigens have also been linked to susceptibility to Norwalk (HuNoV GI.1 prototype) infection, with AB, and B blood group antigen individuals showing a lower incidence of infection and illness with HuNoV [28]. Understanding the binding capacity of HuNoV VLPs to blood group antigens allowed for the development of Hemagglutination Inhibition Assay [30]. Hemagglutination inhibition (HAI) titers have since been used as a correlate of protection by determining the antibody titers that can effectively block the binding of blood group antigens with HuNoV[31].
Antibodies found in serum that block NoV-HBGA binding (HBGA blocking antibodies) have been suggested as a direct correlate of protection from clinical illness (vomiting and diarrhea) following GII.4 HuNoV vaccination or challenge. In a HuNoV human volunteer challenge study, pre-challenge serum titer levels (>1:500) of functional antibodies that block binding of HuNoV VLPs to HBGAs correlated with lower risk of illness [32]. High NoV-specific serum, and salivary IgA and NoV-specific memory IgG cells have been identified as correlates of protection against NoV AGE [31][33].
Interferon-γ (IFN-γ) secreted by CD4+ and CD8+ T cells in response to NoV infections have been correlated with protection in humans [34], mice[35][36], and Gn pig models [37][38]. The host factors associated with HuNoV infection are poorly understood due to their host-specificity and the limited access to laboratory models to study its pathogenesis. Incremental advancements have been made by studying different aspects of NoV infection using different animal models and collating the results together to form a better understanding of NoV infection in humans [39].
6. Animal models for NoV research
Currently, there is no animal model that can directly recapitulate the full range of NoV disease symptoms that are observed in humans, but animals in which evidence for infection has been reported following challenge with HuNoV include Gn piglets and calves [40][41][42][43], monkeys and chimpanzees [44], , and immunodeficient mice [45]. MNV has been used as a surrogate to study the pathogenesis, immunology, and replication of HuNoV infections, and a large amount of data has been generated [46]. However, the limitations of this model are obvious due to the difference between the two viruses in clinical manifestations (without diarrhea/vomiting), host receptors (sialic acid versus HBGAs), infected cell types (dendritic/macrophages versus digestive epithelial cells), and pathogenesis [47].
An ideal animal model of HuNoV infection and disease would be one that can replicate the biological and clinical features associated with human disease, including the route of infection, the infective dose, the disease progression and pathogenesis, and correlates of protection [48]. Among the different models used in norovirus (NoV) research [44][40][41][45], gnotobiotic (Gn) pigs provide a suitable solution for studying HuNoV infection and disease, as well as for evaluating immunogenicity and protective the efficacy of novel vaccine candidates. This is attributed to the close similarity in physiology, immune development, virus binding patterns, and histo-blood group antigen (HBGA) phenotypes between pigs and humans [40]. Since their introduction into research in the early 1960s, Gn pigs have been extensively utilized for the study of enteric viruses and bacteria [49][50]. A Gn pig infection model for two different HuNoV GII.4 variants (GII.4/HS66 and GII.4/2006b) has been established for evaluating HuNoV vaccines [43][38][51][52], displaying similar levels of infectivity compared to HuNoV GI.1/Norwalk and GII.4/2003 Cin-1 in human challenge studies and vaccine trials [53][54][55].
7. Vaccine Development
There are currently no licensed vaccines available for HuNoVs, but several virus-like particle (VLP) vaccine candidates are currently undergoing various stages of clinical trials and have thus far shown some promise in both immunogenicity, and protective efficacy studies, although the correlates of protection to norovirus infections are yet to be fully determined [56][53][54][57]. Treatment for NoV infections is centered over palliative care. This is particularly difficult in developing countries where access to healthcare is limited, and in some locations, clean water is scarce. Due to these reasons, NoV vaccines are urgently needed and they need to be safe, affordable, and efficacious, with a reasonable shelf-life. Vaccines developed to confer protection against NoVs need to account for the genetically and antigenically diverse nature of NoVs [54]. VLPs have been most widely explored as a candidate for NoV vaccines. They are readily produced using various expression systems [58][59][60] and self-assemble to a confirmation that is structurally and antigenically similar to an infectious NoV particle.
VLPs based on GI.1 NoVs were initially developed as a monovalent vaccine candidate and evaluated for its safety, immunogenicity and efficacy in a proof-of-principle study [55]. They were administered intranasally together with monophosphoryl lipoprotein (MPL) and chitosan adjuvants. Study participants who received the vaccine had a reduced incidence rate (37%) of GI.1-associated GE compared to participants who received the placebo vaccine (69%). Infection characterized by the presence of viral RNA detected by RT-PCR was observed among 61% of vaccine recipients compared to 82% who received the placebo.Protection conferred by the vaccine was synonymous with the presence of Norwalk virus-specific serum IgA and IgG levels, along with the induction of IgA ASCs and memory B cells [61][62]. It was soon identified that a bivalent intramuscular (IM) vaccine (GI.1 and GII.4) adjuvanted with aluminum hydroxide elicited a stronger immune response compared to the intranasal administration [61][57][54][53]. This bivalent IM VLP vaccine is currently under Phase III clinical evaluations and is leading the race for licensure for a NoV vaccine.
Another vaccine that is under clinical evaluations is the adenovirus-vectored G1.1 VP1 vaccine which recently concluded its Phase I clinical trials. This vaccine is orally administered in the form of a tablet and is comprised of a non-replicating adenovirus vector expressing the NoV GI.1 VP1 gene adjuvanted with a double-stranded RNA. The vaccine generated a 2-fold increase in blocking titers among 78% of the vaccinated individuals and generated strong systemic and mucosal IgA and IgG responses [63]
P particles [38], VLPs [40]and lactic acid bacteria-based vaccine candidates [64] have been evaluated for their immunogenicity and protective efficacy in Gn pig model of HuNoV infection and diarrhea. Several other candidate vaccines that are still under development include: a RV-VP6 intermediate capsid expressing the NoV GI.3 and GII.4 VLPs as a trivalent vaccine [65]; a bivalent GII.4 NoV and enterovirus 71 VLP-based vaccine [66]; and a Newcastle disease virus vector expressing a GII.4 VP1 gene [67].
8. HuNoV dose-response studies
HuNoVs are known to be highly infectious. A few challenge studies have evaluated the infection potential [68][69][70] and calculated the median infectious dose (ID50) of various norovirus strains. In humans, the ID50 of Norwalk virus was identified to be between 18 and 2800 genomic equivalents [71][72]. Similarly, the ID50 of a GII.4/2006b variant was identified to be ≤2.74 × 103 RNA copies in newborns (4–5 days of age) and 6.43 × 104 RNA copies in older (33–34 days of age) Gn pigs [43]. Dose–response data are critically important for the standardization of the animal challenge model used in the pre-clinical evaluation of vaccine efficacy and anti-viral agents. While ID50 is of primary interest to clinicians, such studies are also widely used to interpret pathogen occurrence and exposure data and translate them to health outcomes. For example, quantitative microbial risk assessments (QMRAs) for pathogen infections in humans are regularly used to develop and improve industrial and regulatory policy in the water and food safety sectors [73][74]. Utilization of dose–response approaches (experimental design and analysis) concurrently enables wider application and impact from experimentally derived data.
ID50 values are often estimated using classical methods, such as the Reed–Muench, Dragstedt–Behrens, and Spearman–Karber methods or more recently by logistic regression. In recent decades, several mechanistic models have been developed to describe plausible phenomena that are inherent to experimentally derived dose–response data [71][75]. Critically, such models provide a basis for inference about the probability of infection at any dose level (i.e., not just ID50), though low-dose extrapolation is an ever-present concern for applications, such as drinking water risk assessments in which mean doses are often less than one pathogen [76]. Contemporary dose–response models may offer greater flexibility and accuracy (i.e., managing inconsistent dilution factors between doses or numbers of subjects per dose group or small numbers of subjects) in estimating ID50.
9. Virus Inoculum
The GII.4 HuNoV challenge pool (103,041) used in this study, identified henceforth as Cin-2, was the unfiltered 10% suspension of stool samples collected from a volunteer who was challenged with HuNoV GII.4/2003 (Hu/GII.4/Cin-1/2003/US), a 2002 Farmington Hills-like variant (GenBank number JQ965810), as part of a vaccine study conducted by Xi Jiang’s laboratory at Cincinnati Children’s Hospital Medical Center. The volunteer developed a 3-day illness characterized by diarrhea, vomiting, nausea, abdominal cramps, and fever [54][77]. The HuNoV concentration of the challenge pool was determined by RT-qPCR to be approximately 2 × 106 viral genomic RNA copies/mL of stool. The challenge pool was stored at −80 °C in individual 1 mL aliquots until the day of Gn pig inoculation.
10. Assessment of Infection Status in Gn Pigs
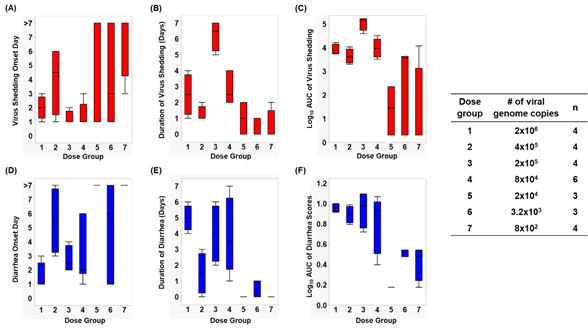
Infection caused by HuNoV Cin-2 was defined by the presence of viral RNA quantified using RT-qPCR carried out on samples isolated from rectal swabs collected from PID 1 to 7. The percentage of affected animals, mean days to onset, duration, peak titer and area under the curve (AUC) of virus shed in feces of pigs in each of the seven dose groups are summarized in Table 1. An increase in inoculation dose was positively associated with a shorter incubation period (Figure 1A), which coincided with the observed increased duration (Figure 1B) as well as increased overall virus shedding in feces measured by AUC (Figure 1C). All pigs belonging to dose groups 1 to 4 shed detectable titers of virus in their feces, while 67% of pigs in dose groups 5 and 6 and 25% of the pigs in dose group 7 shed viruses. It is important to note that pigs in dose group 3 shed a significantly higher (>1 × 106 RNA copies) amount of viruses in their feces (p < 0.0001) and shed viruses for a significantly longer duration (>3 days) than pigs in the other groups (p < 0.0001) (Figure 1A,B).
Figure 1. Box and whisker plots showing (A) virus shedding onset day, (B) duration of virus shedding, (C) log10 AUC of virus shedding, (D) diarrhea onset day, (E) duration of diarrhea, and (F) log10 cumulative diarrhea scores, among each dose group. The maximum and minimum values are denoted by the whisker and the boundaries of each box represent the quartiles with the mean indicated by a black line.
11. Assessment of Diarrhea Status in Gn Pigs
After inoculation, pigs were monitored daily for diarrhea status. The percentage of pigs with diarrhea, the mean duration, the AUC and mean onset day of diarrhea from Cin-2 inoculated pigs are summarized in Table 1. A faster diarrhea onset (Figure 1D), a longer duration of diarrhea (Figure 1E) and a higher cumulative diarrhea score (Figure 1F) were observed in Gn pigs that were inoculated with higher doses of Cin-2. Pigs in dose group 3 that had the highest virus titers shed in their feces also experienced diarrhea for a longer duration of time than the pigs inoculated with other doses. Pigs belonging to dose group 1 experienced diarrhea for the longest duration of time (5 out of 7 days) and had the quickest onset (within 1.5 days after inoculation), indicating severe clinical onset, mimicking that in humans [40][52], while HuNoV-associated gastroenteritis typically lasts for 3–5 days after infection among susceptible populations [78]. Pigs in dose group 6 had a delayed onset to diarrhea, together with lower cumulative fecal consistency scores, demonstrating a milder diarrhea burden. No diarrhea was observed among pigs in dose groups 5 and 7.
12. Determination of ID50 and DD50 Using Various Dose–Response Models
(Table 1 and see details in doi:10.3390/v12090955).
|
Table 1. ID50 and DD50 calculations of Cin-2 |
|||||
|
Method |
Log10ID50 |
ID50 |
Log10DD50 |
DD50 |
|
|
Reed-Muench |
|||||
|
Hand calculation “skrmdb” R script |
3.40 |
2.51x103 |
4.58 |
3.80x104 |
|
|
Dragstedt-Behrens “skrmdb” R script |
3.39 |
2.45x103 |
4.58 |
3.80x104 |
|
|
Spearman-Karber Hand calculation “skrmdb” R script online calculator |
3.52 |
3.31x103 |
4.49 |
3.09x104 |
|
|
Logistic Regression online calculator |
3.40 |
2.51x103 |
|
4.34 |
2.18x104 |
|
Exponential a R script |
3.76 |
5.75x103 |
|
4.76 |
5.75x104 |
|
Approximate Beta-Poisson a |
|
|
|
|
|
|
R script |
3.41 |
2.57x103 |
|
4.33 |
2.13x104 |
|
a. Exponential and Beta-Poisson were determined based on R script used by Weir et al. 2017 [48]. |
|||||
References
- Daniel C. Payne; Jan Vinjé; Peter G. Szilagyi; Kathryn M. Edwards; Mary Allen Staat; Geoffrey A. Weinberg; Caroline B. Hall; James Chappell; David I. Bernstein; Aaron T. Curns; et al.Mary WikswoS. Hannah ShirleyAron J. HallBenjamin LopmanUmesh D. Parashar Norovirus and Medically Attended Gastroenteritis in U.S. Children. New England Journal of Medicine 2013, 368, 1121-1130, 10.1056/nejmsa1206589.
- Kimberly Pringle; Benjamin Lopman; Everardo Vega; Jan Vinjé; Umesh D Parashar; Aron J Hall; Noroviruses: epidemiology, immunity and prospects for prevention. Future Microbiology 2014, 10, 53-67, 10.2217/fmb.14.102.
- Saba Rouhani; Pablo Peñataro Yori; Maribel Paredes Olortegui; Mery Siguas Salas; Dixner Rengifo Trigoso; Dinesh Mondal; Ladaporn Bodhidatta; James Platts-Mills; Amidou Samie; Furqan Kabir; et al.Aldo Am LimaSudhir BabjiCarl J. MasonAdil KalamPascal Obong BessongTahmeed AhmedEstomih MdumaZulfiqar A. BhuttaIla LimaRakhi RamdassDennis LangAjila GeorgeAnita K. M. ZaidiGagandeep KangEric HouptMargaret N. KosekCesar Banda ChavezJulian Torres FloresAngel Orbe VasquezSilvia Rengifo PinedoAngel Mendez AcostaImran AhmedDidar AlamAsad AliShahida QureshiMuneera RasheedSajid SoofiAli TurabAisha K YousafzaiAnuradha BoseM. Steffi JenniferSushil JohnShiny KakiBeena KoshyJayaPrakash MuliyilMohan Venkata RaghavaAnup RamachandranSrujan L. SharmaRahul J. ThomasWilliam PanRamya AmbikapathiDanny CarreonVivek CharuLeyfou DaboViyada DoanJhanelle GrahamChristel HoestStacey KnoblerBenjamin McCormickMonica McGrathMark MillerArchana MohaleGaurvika NayyarStephanie PsakiZeba RasmussenStephanie A. RichardJessica C SeidmanVivian WangRebecca BlankMichael GottliebKaren H. TountasCaroline AmourBuliga Mujaga SwemaLadislaus YarrotRosemary NshamaA. M. Shamsir AhmedFahmida TofailRashidul HaqueIqbal HossainMunirul IslamMustafa MahfuzRam Krishna ChandyoPrakash Sunder ShresthaRita ShresthaManjeswori UlakRobert BlackLaura CaulfieldWilliam CheckleyPing ChenGwenyth LeeLaura E. Murray-KolbBarbara SchaeferLaura PendergastCláudia AbreuA. HavtHilda CostaAlessandra Di MouraJose Quirino FilhoÁlvaro LeiteNoélia LimaBruna MacielMilena MoraesFrancisco MotaReinaldo B. OriáJosiane QuetzAlberto SoaresCrystal L PatilCloupas MahopoAngelina MaphulaCebisa NesamvuniEmanuel NyathiLeah BarrettJean GratzRichard GuerrantWilliam PetriRebecca ScharfBinob ShresthaSanjaya Kumar ShresthaTor A. StrandErling Svensen Norovirus Infection and Acquired Immunity in 8 Countries: Results From the MAL-ED Study. Clinical Infectious Diseases 2016, 62, 1210-1217, 10.1093/cid/ciw072.
- Martyn Kirk; Sara Monteiro Pires; Robert E. Black; Marisa Caipo; John A. Crump; Brecht Devleesschauwer; Dörte Döpfer; Aamir Fazil; Christa L. Fischer-Walker; Tine Hald; et al.Aron J. HallKaren H. KeddyRobin J. LakeClaudio F. LanataPaul TorgersonArie H. HavelaarFrederick J. Angulo World Health Organization Estimates of the Global and Regional Disease Burden of 22 Foodborne Bacterial, Protozoal, and Viral Diseases, 2010: A Data Synthesis. PLOS Medicine 2015, 12, e1001921, 10.1371/journal.pmed.1001921.
- Sharia M Ahmed; Aron J. Hall; Anne E Robinson; Linda Verhoef; Prasanna S Premkumar; Umesh D Parashar; Marion Koopmans; Ben Lopman; Global prevalence of norovirus in cases of gastroenteritis: a systematic review and meta-analysis. The Lancet Infectious Diseases 2014, 14, 725-730, 10.1016/s1473-3099(14)70767-4.
- CDC (2011). "Updated norovirus outbreak management and disease prevention guidelines." MMWR Recomm Rep 60(Rr-3): 1-18
- Julia Brennan; Steffany J. Cavallo; Katie Garman; Kailey Lewis; D.J. Irving; Christina Moore; Linda Thomas; Jeffrey Hill; Raquel Villegas; Joe F. Norman; et al.John R. DunnWilliam SchaffnerTimothy F. Jones Notes from the Field: Multiple Modes of Transmission During a Thanksgiving Day Norovirus Outbreak — Tennessee, 2017. MMWR. Morbidity and Mortality Weekly Report 2018, 67, 1300-1301, 10.15585/mmwr.mm6746a4.
- Rebecca J. Free; Bryan F. Buss; Samir Koirala; Monica Ulses; Anna Carlson; Brianna Loeck; Tom Safranek; Successive Norovirus Outbreaks at an Event Center - Nebraska, October-November, 2017.. MMWR. Morbidity and Mortality Weekly Report 2019, 68, 627-630, 10.15585/mmwr.mm6828a2.
- Annelies Kroneman; Everardo Vega; Harry Vennema; Jan Vinjé; Peter A. White; G. S. Hansman; Kim Green; Vito Martella; Kazuhiko Katayama; Marion Koopmans; et al. Proposal for a unified norovirus nomenclature and genotyping. Archives of Virology 2013, 158, 2059-2068, 10.1007/s00705-013-1708-5.
- Miranda De Graaf; Janko Van Beek; Marion Koopmans; Human norovirus transmission and evolution in a changing world. Nature Reviews Microbiology 2016, 14, 421-433, 10.1038/nrmicro.2016.48.
- Preeti Chhabra; Miranda De Graaf; Gabriel I. Parra; Martin C. W. Chan; Kim Y. Green; Vito Martella; Qiuhong Wang; Peter A. White; Kazuhiko Katayama; Harry Vennema; et al.Marion P. G. KoopmansJan Vinjé Updated classification of norovirus genogroups and genotypes. Journal of General Virology 2019, 100, 1393-1406, 10.1099/jgv.0.001318.
- Jan Vinjé; Advances in Laboratory Methods for Detection and Typing of Norovirus. Journal of Clinical Microbiology 2014, 53, 373-381, 10.1128/jcm.01535-14.
- Nicolas W. Cortes-Penfield; Sasirekha Ramani; Mary K. Estes; Robert L Atmar; Prospects and Challenges in the Development of a Norovirus Vaccine. Clinical Therapeutics 2017, 39, 1537-1549, 10.1016/j.clinthera.2017.07.002.
- Ming Tan; Xi Jiang; The P Domain of Norovirus Capsid Protein Forms a Subviral Particle That Binds to Histo-Blood Group Antigen Receptors. Journal of Virology 2005, 79, 14017-14030, 10.1128/jvi.79.22.14017-14030.2005.
- Ming Tan; Xi Jiang; Tan; Norovirus Capsid Protein-Derived Nanoparticles and Polymers as Versatile Platforms for Antigen Presentation and Vaccine Development. Pharmaceutics 2019, 11, 472, 10.3390/pharmaceutics11090472.
- Michaela Conley; Marion McElwee; Liyana Azmi; Mads Gabrielsen; Olwyn Byron; Ian Goodfellow; David Bhella; Calicivirus VP2 forms a portal-like assembly following receptor engagement. Nature 2019, 565, 377-381, 10.1038/s41586-018-0852-1.
- Mark S Riddle; Richard I. Walker; Status of vaccine research and development for norovirus. Vaccine 2016, 34, 2895-2899, 10.1016/j.vaccine.2016.03.077.
- Erwin Duizer; Kellogg J. Schwab; Frederick H. Neill; Robert L. Atmar; Marion P. G. Koopmans; Mary K. Estes; Laboratory efforts to cultivate noroviruses. Journal of General Virology 2003, 85, 79-87, 10.1099/vir.0.19478-0.
- Umesh C. Karandikar; Sue E. Crawford; Nadim J. Ajami; Kosuke Murakami; Baijun Kou; Khalil Ettayebi; Genovefa Papanicolaou; Ubonvan Jongwutiwes; Miguel-Angel Perales; Jinru Shia; et al.David MercerMilton J. FinegoldJan VinjéRobert L. AtmarMary K. Estes Detection of human norovirus in intestinal biopsies from immunocompromised transplant patients. Journal of General Virology 2016, 97, 2291-2300, 10.1099/jgv.0.000545.
- Melissa K. Jones; Makiko Watanabe; Shu Zhu; Christina L. Graves; Lisa R. Keyes; Katrina R. Grau; Mariam B. Gonzalez-Hernandez; Nicole M. Iovine; Christiane E. Wobus; Jan Vinjé; et al.Scott A. TibbettsShannon M. WalletStephanie M. Karst Enteric bacteria promote human and mouse norovirus infection of B cells. Science 2014, 346, 755-759, 10.1126/science.1257147.
- Julianne R Brown; Kimberly Gilmour; Judith Breuer; Norovirus infections occur in B cell deficient patients. Clinical Infectious Diseases 2016, 62, 1136-1138, 10.1093/cid/ciw060.
- Shaohua Lei; Junghyun Ryu; Ke Wen; Erica Twitchell; Tammy Bui; Ashwin Ramesh; Mariah Weiss; Guohua Li; Helen Samuel; Sherrie Clark-Deener; et al.Xi JiangKiho LeeLijuan Yuan Increased and prolonged human norovirus infection in RAG2/IL2RG deficient gnotobiotic pigs with severe combined immunodeficiency. Scientific Reports 2016, 6, 25222, 10.1038/srep25222.
- Thavamathi Annamalai; Zhongyan Lu; Kwonil Jung; Stephanie N. Langel; Christopher K. Tuggle; Jack C.M. Dekkers; Emily H. Waide; Sukumar Kandasamy; Linda J. Saif; Infectivity of GII.4 human norovirus does not differ between T-B-NK+ severe combined immunodeficiency (SCID) and non-SCID gnotobiotic pigs, implicating the role of NK cells in mediation of human norovirus infection. Virus Research 2019, 267, 21-25, 10.1016/j.virusres.2019.05.002.
- Khalil Ettayebi; Sue E. Crawford; Kosuke Murakami; James R. Broughman; Umesh Karandikar; Victoria R. Tenge; Frederick H. Neill; Sarah E. Blutt; Xi-Lei Zeng; Lin Qu; et al.Baijun KouAntone R. OpekunDouglas G. BurrinDavid Y. GrahamSasirekha RamaniRobert L. AtmarMary K. Estes Replication of human noroviruses in stem cell-derived human enteroids. Science 2016, 353, 1387-1393, 10.1126/science.aaf5211.
- Kim Y. Green; Stuart S. Kaufman; Bianca M. Nagata; Natthawan Chaimongkol; Daniel Y. Kim; Eric A. Levenson; Christine M. Tin; Allison Behrle Yardley; Jordan A. Johnson; Ana Beatriz F. Barletta; et al.Khalid M. KhanNada A. YazigiSukanya SubramanianSangeetha R. MoturiThomas M. FishbeinIan N. MooreStanislav V. Sosnovtsev Human norovirus targets enteroendocrine epithelial cells in the small intestine. Nature Communications 2020, 11, 2759, 10.1038/s41467-020-16491-3.
- Mary K. Estes; Khalil Ettayebi; Victoria R. Tenge; Kosuke Murakami; Umesh Karandikar; Shih-Ching Lin; B. Vijayalakshmi Ayyar; Nicolás W Cortés-Penfield; Kei Haga; Frederick H. Neill; et al.Antone R OpekunJames R. BroughmanXi-Lei ZengSarah E. BluttSue E. CrawfordSasirekha RamaniDavid Y GrahamRobert L Atmar Human Norovirus Cultivation in Nontransformed Stem Cell-Derived Human Intestinal Enteroid Cultures: Success and Challenges. Viruses 2019, 11, 638, 10.3390/v11070638.
- Severine Marionneau; Nathalie Ruvoën; Beatrice Le Moullac-Vaidye; Monique Clement; Anne Cailleau-Thomas; Guillermo Ruiz-Palacois; Pengwei Huang; Xi Jiang; Jacques Le Pendu; Norwalk virus binds to histo-blood group antigens present on gastroduodenal epithelial cells of secretor individuals.. Gastroenterology 2002, 122, 1967-77, 10.1053/gast.2002.33661.
- Lisa Lindesmith; Christine Moe; Séverine Marionneau; Nathalie Ruvoen; Xi Jiang; Lauren Lindblad; Paul Stewart; Jacques LePendu; Ralph S. Baric; Human susceptibility and resistance to Norwalk virus infection. Nature Medicine 2003, 9, 548-553, 10.1038/nm860.
- Anne M. Hutson; Fabrice Airaud; Jacques LePendu; Mary K. Estes; Robert L. Atmar; Norwalk virus infection associates with secretor status genotyped from sera. Journal of Medical Virology 2005, 77, 116-120, 10.1002/jmv.20423.
- Rita Czako; Robert L Atmar; Antone R. Opekun; Mark A. Gilger; David Y. Graham; Mary K. Estes; Serum Hemagglutination Inhibition Activity Correlates with Protection from Gastroenteritis in Persons Infected with Norwalk Virus. Clinical and Vaccine Immunology 2011, 19, 284-287, 10.1128/cvi.05592-11.
- Sasirekha Ramani; Mary K. Estes; Robert L Atmar; Correlates of Protection against Norovirus Infection and Disease—Where Are We Now, Where Do We Go?. PLOS Pathogens 2016, 12, e1005334, 10.1371/journal.ppat.1005334.
- Amanda Reeck; Owen Kavanagh; Mary K. Estes; Antone R. Opekun; Mark A. Gilger; David Y. Graham; Robert L. Atmar; Serological Correlate of Protection against Norovirus‐Induced Gastroenteritis. The Journal of Infectious Diseases 2010, 202, 1212-1218, 10.1086/656364.
- Robert L Atmar; David I. Bernstein; G. Marshall Lyon; John J. Treanor; Mohamed S. Al-Ibrahim; David Y. Graham; Jan Vinjé; Xi Jiang; Nicole Gregoricus; Robert W. Frenck; et al.Christine L. MoeWilbur H. ChenJennifer FerreiraJill BarrettAntone R. OpekunMary K. EstesAstrid BorkowskiFrank BaehnerRobert GoodwinAnthony EdmondsPaul M. Mendelman Serological Correlates of Protection against a GII.4 Norovirus. Clinical and Vaccine Immunology 2015, 22, 923-929, 10.1128/cvi.00196-15.
- Lisa Lindesmith; Christine Moe; Jacques LePendu; Jeffrey A. Frelinger; John Treanor; Ralph S. Baric; Cellular and Humoral Immunity following Snow Mountain Virus Challenge. Journal of Virology 2005, 79, 2900-2909, 10.1128/jvi.79.5.2900-2909.2005.
- Shu Zhu; Doron Regev; Makiko Watanabe; Danielle Hickman; Nissin Moussatché; Desyreé Murta Jesus; Shannon M. Kahan; Sawsan Napthine; Ian Brierley; Robert N. Hunter; et al.Divya DevabhaktuniMelissa K. JonesStephanie M. Karst Identification of Immune and Viral Correlates of Norovirus Protective Immunity through Comparative Study of Intra-Cluster Norovirus Strains. PLOS Pathogens 2013, 9, e1003592, 10.1371/journal.ppat.1003592.
- Marisela R. Rodriguez; Kristen Monte; Larissa B. Thackray; Deborah J. Lenschow; ISG15 Functions as an Interferon-Mediated Antiviral Effector Early in the Murine Norovirus Life Cycle. Journal of Virology 2014, 88, 9277-9286, 10.1128/jvi.01422-14.
- Shaohua Lei; Ashwin Ramesh; Erica Twitchell; Ke Wen; Tammy Bui; Mariah Weiss; Xingdong Yang; Jacob Kocher; Guohua Li; Ernawati Giri-Rachman; et al.Nguyen Van TrangXi JiangElizabeth P. RyanLijuan Yuan High Protective Efficacy of Probiotics and Rice Bran against Human Norovirus Infection and Diarrhea in Gnotobiotic Pigs. Frontiers in Microbiology 2016, 7, 1-12, 10.3389/fmicb.2016.01699.
- Jacob Kocher; Tammy Bui; Ernawati Giri-Rachman; K. Wen; Guohua Li; Xingdong Yang; Fangning Liu; Ming Tan; Ming Xia; Weiming Zhong; et al.Xi JiangLijuan Yuan Intranasal P Particle Vaccine Provided Partial Cross-Variant Protection against Human GII.4 Norovirus Diarrhea in Gnotobiotic Pigs. Journal of Virology 2014, 88, 9728-9743, 10.1128/jvi.01249-14.
- Christiane E. Wobus; J.B. Cunha; M.D. Elftman; A.O. Kolawole; Animal Models of Norovirus Infection. Congenital and Other Related Infectious Diseases of the Newborn 2015, 1, 397-422, 10.1016/b978-0-12-802241-2.00019-5.
- S. Cheetham; Menira B. L. D. Souza; R. McGregor; T. Meulia; Q. Wang; L. J. Saif; Binding Patterns of Human Norovirus-Like Particles to Buccal and Intestinal Tissues of Gnotobiotic Pigs in Relation to A/H Histo-Blood Group Antigen Expression. Journal of Virology 2007, 81, 3535-3544, 10.1128/jvi.01306-06.
- Menira B. L. D. Souza; M. S. P. Azevedo; K. Jung; S. Cheetham; L. J. Saif; Pathogenesis and Immune Responses in Gnotobiotic Calves after Infection with the Genogroup II.4-HS66 Strain of Human Norovirus. Journal of Virology 2007, 82, 1777-1786, 10.1128/jvi.01347-07.
- Kwonil Jung; Qiuhong Wang; Yunjeong Kim; Kelly Scheuer; Zhenwen Zhang; Quan Shen; Kyeong-Ok Chang; Linda J. Saif; The Effects of Simvastatin or Interferon-α on Infectivity of Human Norovirus Using a Gnotobiotic Pig Model for the Study of Antivirals. PLOS ONE 2012, 7, e41619, 10.1371/journal.pone.0041619.
- Tammy Bui; Jacob Kocher; Yanru Li; K. Wen; Guohua Li; Fangning Liu; Xingdong Yang; Tanya Leroith; Ming Tan; Ming Xia; et al.Weiming ZhongX. JiangLijuan Yuan Median infectious dose of human norovirus GII.4 in gnotobiotic pigs is decreased by simvastatin treatment and increased by age. Journal of General Virology 2013, 94, 2005-2016, 10.1099/vir.0.054080-0.
- Karin Bok; Gabriel I. Parra; Tanaji Mitra; Eugenio Abente; Charlene K. Shaver; Denali Boon; Ronald Engle; Claro Yu; Albert Z. Kapikian; Stanislav V. Sosnovtsev; et al.Robert H. PurcellKim Y. Green Chimpanzees as an animal model for human norovirus infection and vaccine development. Proceedings of the National Academy of Sciences 2010, 108, 325-330, 10.1073/pnas.1014577107.
- Stefan Taube; Abimbola O. Kolawole; Marina Höhne; John E. Wilkinson; Scott A. Handley; Jeffrey W. Perry; Larissa B. Thackray; Ramesh Akkina; Christiane E. Wobus; A Mouse Model for Human Norovirus. mBio 2013, 4, e00450-13, 10.1128/mbio.00450-13.
- Christiane E. Wobus; Stephanie M Karst; Larissa B Thackray; Kyeong-Ok Chang; Stanislav V Sosnovtsev; Gael Belliot; Anne Krug; Jason M MacKenzie; Kim Y Green; Herbert W. Virgin; et al. Replication of Norovirus in Cell Culture Reveals a Tropism for Dendritic Cells and Macrophages. PLOS Biology 2004, 2, e432, 10.1371/journal.pbio.0020432.
- Ming Tan; Xi Jiang; Norovirus Gastroenteritis, Carbohydrate Receptors, and Animal Models. PLOS Pathogens 2010, 6, e1000983, 10.1371/journal.ppat.1000983.
- Volker Gerdts; Heather L Wilson; F. Meurens; Sylvia Van Drunen Littel - Van Den Hurk; Don Wilson; Stewart Walker; Colette Wheler; Hugh Townsend; Andrew A. Potter; Large Animal Models for Vaccine Development and Testing. ILAR Journal 2015, 56, 53-62, 10.1093/ilar/ilv009.
- Kyle V. Todd; Ralph A. Tripp; Human Norovirus: Experimental Models of Infection. Viruses 2019, 11, 151, 10.3390/v11020151.
- Lijuan Yuan; Annelise Geyer; Douglas C. Hodgins; Zhiqian Fan; Yuan Qian; Kyeong-Ok Chang; Sue E. Crawford; Viviana Parreño; Lucy A. Ward; Mary K. Estes; et al.Margaret E. ConnerLinda J. Saif Intranasal Administration of 2/6-Rotavirus-Like Particles with Mutant Escherichia coli Heat-Labile Toxin (LT-R192G) Induces Antibody-Secreting Cell Responses but Not Protective Immunity in Gnotobiotic Pigs. Journal of Virology 2000, 74, 8843-8853, 10.1128/jvi.74.19.8843-8853.2000.
- Menira B. L. D. Souza; Veronica Costantini; Marli. S.P. Azevedo; Linda J. Saif; A human norovirus-like particle vaccine adjuvanted with ISCOM or mLT induces cytokine and antibody responses and protection to the homologous GII.4 human norovirus in a gnotobiotic pig disease model. Vaccine 2007, 25, 8448-8459, 10.1016/j.vaccine.2007.09.040.
- Sonia Cheetham; Menira B. L. D. Souza; Tea Meulia; Sheila Grimes; Myung Guk Han; Linda J. Saif; Pathogenesis of a Genogroup II Human Norovirus in Gnotobiotic Pigs. Journal of Virology 2006, 80, 10372-10381, 10.1128/jvi.00809-06.
- Aarthi Sundararajan; Mark Sangster; Sharon Frey; Robert L Atmar; Wilbur Chen; Jennifer Ferreira; Robert Bargatze; Paul M. Mendelman; John J. Treanor; David J. Topham; et al. Robust mucosal-homing antibody-secreting B cell responses induced by intramuscular administration of adjuvanted bivalent human norovirus-like particle vaccine. Vaccine 2014, 33, 568-576, 10.1016/j.vaccine.2014.09.073.
- David I. Bernstein; Robert L. Atmar; G. Marshall Lyon; John J. Treanor; Wilbur H. Chen; X. Jiang; Jan Vinjé; Nicole Gregoricus; Robert W. Frenck; Christine L. Moe; et al.Mohamed S. Al-IbrahimJill BarrettJennifer FerreiraMary K. EstesDavid Y. GrahamRobert GoodwinAstrid BorkowskiRalf ClemensPaul M. Mendelman Norovirus Vaccine Against Experimental Human GII.4 Virus Illness: A Challenge Study in Healthy Adults. The Journal of Infectious Diseases 2014, 211, 870-878, 10.1093/infdis/jiu497.
- Robert L Atmar; David I. Bernstein; Clayton D. Harro; Mohamed S. Al-Ibrahim; Wilbur H. Chen; Jennifer Ferreira; Mary K. Estes; David Y. Graham; Antone R. Opekun; Charles Richardson; et al.Paul M. Mendelman Norovirus Vaccine against Experimental Human Norwalk Virus Illness. New England Journal of Medicine 2011, 365, 2178-2187, 10.1056/nejmoa1101245.
- Robert L Atmar; Frank Baehner; Jakob P Cramer; Eric Lloyd; James Sherwood; Astrid Borkowski; Paul M Mendelman; NOR-201 Study Group; Mohamed S Al-Ibrahim; David L Bernstein; et al.Donald M BrandonLaurence ChuMatthew G DavisRobert J EpsteinSharon E FreyJeffrey B RosenJohn J Treanor Antibody persistence to two Virus-Like Particle norovirus vaccine candidate formulations in healthy adults: one-year follow-up with memory probe vaccination.. The Journal of Infectious Diseases 2019, 220, 603-614, 10.1093/infdis/jiz170.
- J. J. Treanor; R. L. Atmar; S. E. Frey; R. Gormley; W. H. Chen; J. Ferreira; R. Goodwin; A. Borkowski; R. Clemens; P. M. Mendelman; et al.Bao G. VuChristopher S. StachWilmara Salgado-PabónDaniel J. DiekemaSue E. GardnerP. M. Schlievert A Novel Intramuscular Bivalent Norovirus Virus-Like Particle Vaccine Candidate--Reactogenicity, Safety, and Immunogenicity in a Phase 1 Trial in Healthy Adults. The Journal of Infectious Diseases 2014, 210, 1763-1771, 10.1093/infdis/jiu337.
- X Jiang; M Wang; D Y Graham; M K Estes; Expression, self-assembly, and antigenicity of the Norwalk virus capsid protein.. Journal of Virology 1991, 66, 6527-6532, 10.1128/jvi.66.11.6527-6532.1992.
- Kirsi Tamminen; Leena Huhti; Tiia Koho; Suvi Lappalainen; Vesa P. Hytönen; Timo Vesikari; Vesna Blazevic; A comparison of immunogenicity of norovirus GII-4 virus-like particles and P-particles. Immunology 2011, 135, 89-99, 10.1111/j.1365-2567.2011.03516.x.
- Ming Tan; Xi Jiang; Norovirus P particle: a subviral nanoparticle for vaccine development against norovirus, rotavirus and influenza virus. Nanomedicine 2012, 7, 889-897, 10.2217/nnm.12.62.
- Samer S. El‐Kamary; Marcela F Pasetti; Paul M. Mendelman; Sharon E. Frey; David I. Bernstein; John J. Treanor; Jennifer Ferreira; Wilbur H. Chen; Richard Sublett; Charles Richardson; et al.Robert F. BargatzeMarcelo B. SzteinCarol O. Tacket Adjuvanted Intranasal Norwalk Virus-Like Particle Vaccine Elicits Antibodies and Antibody-Secreting Cells That Express Homing Receptors for Mucosal and Peripheral Lymphoid Tissues. The Journal of Infectious Diseases 2010, 202, 1649-1658, 10.1086/657087.
- Karina Ramírez; Rezwanul Wahid; Charles Richardson; Robert F. Bargatze; Samer S. El-Kamary; Marcelo B. Sztein; Marcela F Pasetti; Intranasal vaccination with an adjuvanted Norwalk virus-like particle vaccine elicits antigen-specific B memory responses in human adult volunteers. Clinical Immunology 2012, 144, 98-108, 10.1016/j.clim.2012.05.006.
- Leesun Kim; David Liebowitz; Karen Lin; Kassandra Kasparek; Marcela F Pasetti; Shaily J. Garg; Keith Gottlieb; George Trager; Sean N. Tucker; Safety and immunogenicity of an oral tablet norovirus vaccine, a phase I randomized, placebo-controlled trial.. JCI Insight 2018, 3, 1, 10.1172/jci.insight.121077.
- Kelsey Craig; Xianjun Dai; Anzhong Li; Mijia Lu; Miaoge Xue; Lucia Rosas; Thomas Z. Gao; Andrew J. Niehaus; Ryan N. Jennings; Jiànróng Lǐ; et al. A Lactic Acid Bacteria (LAB)-Based Vaccine Candidate for Human Norovirus.. Viruses 2019, 11, 213, 10.3390/v11030213.
- Maria Malm; Kirsi Tamminen; Suvi Lappalainen; Timo Vesikari; Vesna Blazevic; Rotavirus Recombinant VP6 Nanotubes Act as an Immunomodulator and Delivery Vehicle for Norovirus Virus-Like Particles. Journal of Immunology Research 2015, 2016, 1-13, 10.1155/2016/9171632.
- Xiaoli Wang; Zhiqiang Ku; Wenlong Dai; Tan Chen; Xiaohua Ye; Chao Zhang; Yingyi Zhang; Qingwei Liu; Xia Jin; Zhong Huang; et al. A bivalent virus-like particle based vaccine induces a balanced antibody response against both enterovirus 71 and norovirus in mice. Vaccine 2015, 33, 5779-5785, 10.1016/j.vaccine.2015.09.043.
- Shin-Hee Kim; Shun Chen; X. Jiang; Kim Y. Green; Siba K. Samal; Newcastle Disease Virus Vector Producing Human Norovirus-Like Particles Induces Serum, Cellular, and Mucosal Immune Responses in Mice. Journal of Virology 2014, 88, 9718-9727, 10.1128/jvi.01570-14.
- Françoise S. Le Guyader; Frederick H. Neill; Eric Dubois; Fabienne Bon; Fabienne Loisy; Evelyne Kohli; Monique Pommepuy; Robert L. Atmar; A semiquantitative approach to estimate Norwalk-like virus contamination of oysters implicated in an outbreak. International Journal of Food Microbiology 2003, 87, 107-112, 10.1016/s0168-1605(03)00058-8.
- Françoise S. Le Guyader; Joanna Krol; Katia Ambert-Balay; Nathalie Ruvoen-Clouet; Benedicte Desaubliaux; Sylvain Parnaudeau; Jean-Claude Le Saux; Agnès Ponge; Pierre Pothier; Robert L. Atmar; et al.Jacques Le Pendu Comprehensive Analysis of a Norovirus-Associated Gastroenteritis Outbreak, from the Environment to the Consumer. Journal of Clinical Microbiology 2010, 48, 915-920, 10.1128/jcm.01664-09.
- J. A. Lowther; Nicole E. Gustar; Rachel E. Hartnell; David N. Lees; Comparison of Norovirus RNA Levels in Outbreak-Related Oysters with Background Environmental Levels. Journal of Food Protection 2012, 75, 389-393, 10.4315/0362-028x.jfp-11-360.
- Peter Teunis; Christine L. Moe; Pengbo Liu; Sara E. Miller; Lisa Lindesmith; Ralph S. Baric; Jacques Le Pendu; Rebecca L. Calderon; Norwalk virus: How infectious is it?. Journal of Medical Virology 2008, 80, 1468-1476, 10.1002/jmv.21237.
- Robert L Atmar; Antone R. Opekun; Mark A. Gilger; Mary K. Estes; Sue E. Crawford; Frederick H. Neill; Sasirekha Ramani; Heather Hill; Jennifer Ferreira; David Y. Graham; et al. Determination of the 50% human infectious dose for Norwalk virus.. The Journal of Infectious Diseases 2013, 209, 1016-22, 10.1093/infdis/jit620.
- Haas, C.N.; Rose, J.B.; Gerba, C.P. Quantitative Microbial Risk Assessment; Wiley: Hoboken, NJ, USA, 2014
- World Health Organization. Quantitative Microbial Risk Assessment: Application for Water Safety Management; World Health Organization: Geneva, Switzerland, 2016
- Philip J. Schmidt; Norovirus Dose-Response: Are Currently Available Data Informative Enough to Determine How Susceptible Humans Are to Infection from a Single Virus?. Risk Analysis 2014, 35, 1364-1383, 10.1111/risa.12323.
- Michael J. Messner; Philip Berger; Sharon P. Nappier; Fractional Poisson-A Simple Dose-Response Model for Human Norovirus. Risk Analysis 2014, 34, 1820-1829, 10.1111/risa.12207.
- Robert Frenck; David I. Bernstein; Ming Xia; Pengwei Huang; Weiming Zhong; Susan Parker; Michelle Dickey; Monica M. McNeal; Xi Jiang; Predicting Susceptibility to Norovirus GII.4 by Use of a Challenge Model Involving Humans. The Journal of Infectious Diseases 2012, 206, 1386-1393, 10.1093/infdis/jis514.
- Green, K. Caliciviridae: The Noroviruses. In Fields Virology, 6th ed.; Knipe, D.M., Howley, P.M., Eds.; Wolters Kluwer Health/Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2013; Volume 1, pp. 582–608
- Green, K. Caliciviridae: The Noroviruses. In Fields Virology, 6th ed.; Knipe, D.M., Howley, P.M., Eds.; Wolters Kluwer Health/Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2013; Volume 1, pp. 582–608