Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is a comparison between Version 1 by Hailing Yang and Version 2 by Rita Xu.
Laccase (p-diphenol, EC1.10.3.2) is a copper-containing glycoprotein oxidase. Plant laccase is a ubiquitous multifunctional protein that encodes a large gene family.
- Laccase
- tandem duplication
- Populus trichocarpa
- gene family
- functional divergence
1. Introduction
Oryza sativa L. [1], Brachypodium distachyon (L.) Beauv. [2], and Arabidopsis thaliana (L.) Heynh. [3] have 30, 29, and 17 laccase genes, respectively. The functions of some plant laccases have been revealed through genetics and molecular biology strategies. For example, Arabidopsis laccases AtLAC4, AtLAC11, and AtLAC17 play a key role in the polymerization of lignin [4]. AtLAC15 is involved in the oxidative polymerization of flavonoids [5]. O. sativa laccases Os12g15680 and Os01g63180 participate in herbicide degradation [6]. Zea mays Linn. laccase ZMLAC1 responds to salt stress in maize roots [7]. The cotton laccase GaLAC1 enhances the resistance of Arabidopsis to phenolic allelochemicals [8].
Comparative plant genomics studies confirmed that plant gene families are largely conserved [9]. However, lineage-specific fluctuations in gene family size often occur [9][10][11][9,10,11]. This kind of fluctuation is produced by gene duplication and is specifically used to improve plant adaptability. Plants have a higher gene duplication rate than most eukaryotes [12]. Most of these plant gene copies are derived from whole-genome replication and/or tandem duplication. Earlier studies showed that many repeated genes are retained during the evolutionary process, and that these genes have functional divergence [13]. The effect of duplication mechanism on gene retention has been studied in different species [14][15][14,15]. It is a key issue that needs to be discussed in depth in the study of the evolution of gene families.
The whole-genome sequence of P. trichocarpa has been reported in 2006 as the first woody plant genome data [16]. P. trichocarpa has experienced three genome duplication events, the most recent of which occurred approximately 60–65 million years ago. The annotation results of the P. trichocarpa genome identified about 45,555 protein-coding gene loci. About 4839 tandem-duplicated genes are present in P. trichocarpa and account for 10.6% of the total number of genes [16]. Whether the genes produced by tandem or segment duplication have functional redundancy is of concern. The laccases PtLAC3 (Poptri.010G193100) and PtLAC2 (Poptri.008G064000) of P. trichocarpa are a pair of genes produced by segmental duplication. After inhibition of the expression of PtLAC3, the lignin content and composition are not significantly changed in P. trichocarpa. The growth and development of P. trichocarpa are not affected, but the integrity of the xylem fiber cell wall is affected [17]. Anthony C. Bryan has shown that reducing the expression of PdLAC2 (the ortholog of PtLAC2) can cause changes in the syringyl/guaiacyl ratio of Populus deltoides. PdLAC2 is considered to be involved in the oxidation of phenolics and the binding of flavonoids that interact with cell wall lignin [18]. These findings indicate that the genes produced by segmental duplication have functional differences. However, the current research on the function of tandem-duplicated genes in plants is rare. Therefore, studies of gene function are necessary to reveal the functional differentiation of tandem duplicates.
2. Identification of the LAC Gene Family in the Populus Genome
Previous studies identified 53 PtLAC genes in the P. trichocarpa genome [18]. In this study, re-identification and manual re-annotation were performed to confirm the PtLAC genes of P. trichocarpa further. In total, 58 putative laccase protein-coding gene loci were tentatively identified through BLAST from Phytozome using Populus trichocarpa v3.1 release. Fifty-three full-length genes were numbered PtLAC1–PtLAC53 (Table 1), and these genes all contain typical laccase copper oxidase-conserved domains. The molecular weight of PtLAC was 555–582 KD, and the pI was between 6.46 and 9.91. Five fragments with incomplete domains were considered to be pseudogenes, named PtLAC1-fr–PtLAC5-fr (*indicated in Table 1). Frameshift mutations or premature stop codons in these sequences resulted in incomplete protein domains. After the annotation information release of Populus trichocarpa v4.1, authorswe have compared the PtLAC information of these two versions. In v4.1, PtLAC43, PtLAC44, and PtLAC52 are not annotated. Except for these three genes, the IDs of the remaining laccase genes have not changed. In the phylogenetic tree, PtLAC43 and PtLAC40 are 100% clustered, PtLAC44 and PtLAC41 are 100% clustered, PtLAC52 and PtLAC53 are 100% clustered; these three genes do not affect our classification results and authorswe have not continued to study the functions of these genes. In this article, we still use the v3.1 version.
Table 1. Information of Laccase gene family in Populus trichocarpa.
| Gene | Gene Model | Amino Acid | Isoelectronic Point (pI) | Molecular Weight | Gene | Gene Model | Amino Acid | Isoelectronic Point (pI) | Molecular Weight |
|---|---|---|---|---|---|---|---|---|---|
| PtLAC1 | Potri.001G054600 | 581 | 9.29 | 64.262KD | PtLAC30 | Potri.011G071100 | 555 | 8.72 | 60.919KD |
| PtLAC2 | Potri.001G184300 | 581 | 9.3 | 64.015KD | PtLAC31 | Potri.011G120200 | 562 | 6.67 | 63.079KD |
| PtLAC3 | Potri.001G206200 | 564 | 9.19 | 63.345KD | PtLAC32 | Potri.011G120300 | 581 | 9.03 | 63.935KD |
| PtLAC4 | Potri.001G248700 | 557 | 9.05 | 61.145KD | PtLAC33 | Potri.012G048900 |
ATR), Glycine max (Gm), O. sativa (Os), Physcomitrella patens (Pp), Vitis vinifera (Vv), Herba Selaginellae moellendorfii (SM), and Populus trichocarpa (Pt) under the model of WAG + I + G and bootstrap = 1000. Subfamilies I–VII are marked on the outermost of the tree. The branch length was shown next to the branches. (b) Hypothetical evolutionary histories of laccases in terrestrial plants.
4. Positive Selection Analysis of Plant Laccases
The branch-site model allowed ω ratios to simultaneously vary between sites and branches in order to detect positive selection which affecting a few sites along particular lineages [20][33]. Table 2 lists the parameter estimates for branches under positive selection. When Subfamily I was set as the foreground branch, 2∆lnL between Null and Alternative models was 6.624184 (p < 0.05), and found two possible positive selection sites (163 L 334 F, p > 0.95) (Table 2). In addition, a possible positive selection site was found (26 E, p > 0.95) when Subfamily II or Subfamily III were set as the foreground branch, and the p-value between the null and alternative models was p < 0.05. The 2∆lnL value between the null and alternative models was 5.42867 and 5.278286, respectively. When Subfamily IV was set as the foreground branch, the value of 2∆lnL between Null and Alternative models was 13.565518 (p < 0.01). In total, eight sites (55 C, 93 T, 95 L, 170 M, 227 G, 236 S, 242 A, 324 F, p > 0.95) were found positively selected at a level of 95%. When subfamily V is the foreground branch, the value of 2∆lnL between the null and alternative models was 6.82632 (p < 0.01). Five sites were found (2 V, 33 H, 50 R, 179 I, 252 R), and the posterior probability according to the BEB method was significantly greater than the critical value of 0.95 (p > 0.95). When Subfamily VI and VII were used as foreground branches, no potential positive selection sites were found, and the value of 2∆lnL between the null and alternative models was 3.31621 (p > 0.05) and 0.055894 (p > 0.05), respectively.
Table 2.
Selective pressure analyses of LACCASE genes in terrestrial lants by branch-site model.
| Foreground Branches |
Model | np | LNL | 2Δ (LNL) | Estimates of Parameters | Model Compared | LRT | p | -Value | Positive Sites | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Subfamily I | Model A | 420 | −105,416.831390 | 6.62418 | Site class | 0 | 1 | 2a | 2b | Model A vs.Model A null | p | < 0.05 | 163 L 334 F ( | p | > 0.95) | ||||||
| f | 0.83718 | 0.09491 | 0.06099 | 0.00691 | |||||||||||||||||
| ω0 | 0.12738 | 1.00000 | 0.12738 | 1.00000 | |||||||||||||||||
| 581 | |||||||||||||||||||||
| ω1 | 9.18 | 64.027KD | |||||||||||||||||||
| 0.12738 | 1.00000 | 18.62163 | 18.62163 | PtLAC5 | Potri.001G341600 | 580 | 8.91 | ||||||||||||||
| Model A null | 419 | −105,420.143482 | 64.163KD | PtLAC34 | 1 | Potri.013G152700 | 579 | 9.35 | 64.456KD | ||||||||||||
| Not Allowed | PtLAC6 | Potri.001G401100 | 581 | 9.19 | |||||||||||||||||
| Subfamily II | Model A | 64.264KD | PtLAC35 | 412 | −103,339.374083 | 5.42867 | Potri.014G100600 | 576 | 9.29 | 64.491KD | |||||||||||
| Site class | 0 | 1 | 2a | 2b | Model A vs.Model A null | p | < 0.05 | 26 E ( | p | > 0.95) | PtLAC7 | Potri.001G401300 | 581 | 9.2 | 64.197KD | PtLAC36 | Potri.015G040400 | 572 | 7.26 | 63.456KD | |
| PtLAC8 | Potri.004G156400 | 573 | 6.46 | 64.341KD | PtLAC37 | Potri.015G040600 | 579 | 8.52 | 64.38KD | ||||||||||||
| PtLAC9 | Potri.005G200500 | 564 | 8.94 | 62.557KD | PtLAC38 | Potri.015G040700 | 579 | 8.96 | 64.499KD | ||||||||||||
| PtLAC10 | Potri.005G200600 | 566 | 7.04 | 63.558KD | PtLAC39 | Potri.015G040800 | 559 | 9.91 | 62.010KD | ||||||||||||
| PtLAC11 | Potri.005G200700 | 566 | 7.62 | 63.558KD | PtLAC40 | Potri.016G106000 | −105,420.468381568 | 6.82 | 61.924KD | ||||||||||||
| 5.27829 | Site class | 0 | 1 | 2a | 2b | Model A vs.Model A null | p | < 0.05 | 26 E ( | p | > 0.95) | PtLAC12 | Potri.006G087100 | 565 | 6.5 | 63.026KD | PtLAC41 | Potri.016G106100 | 569 | 8.49 | 62.01KD |
| f | 0.85571 | 0.09694 | 0.04254 | 0.00482 | PtLAC13 | Potri.006G087500 | 580 | 9.19 | 63.79KD | PtLAC42 | |||||||||||
| Potri.016G106300 | 562 | 8.31 | 61.538KD | ||||||||||||||||||
| ω0 | 0.12741 | 1.00000 | 0.12741 | 1.00000 | PtLAC14 | Potri.006G094100 | 579 | 9.84 | 64.094KD | PtLAC43 | Potri.016G107500 | 568 | 6.82 | 61.958KD | |||||||
| PtLAC15 | |||||||||||||||||||||
| ω1 | 0.12741 | 1.00000 | 43.09093 | 43.09093 | Potri.006G096900 | 562 | 8.3 | 61.508KD | |||||||||||||
| Model A null | 419 | −105,423.107524 | 1 | PtLAC44 | Not Allowed | ||||||||||||||||
| Subfamily IV | Model A | 420 | −105,405.397615 | 13.5655 | Site class | 0 | 1 | 2a | 2b | Model A vs.Model A null | p | < 0.01 | 55 C 93 T 95 L 170 M 227 G 236 S 242 A 324 F ( | p | > 0.95) | ||||||
| f | 0.80518 | PtLAC19 | Potri.008G064000 | 562 | 7.26 | 62.356KD | PtLAC48 | Potri.019G088600 | 567 | 7.29 | 62.781KD | ||||||||||
| PtLAC20 | Potri.008G073700 | 556 | 7.27 | 60.948KD | PtLAC49 | Potri.019G088700 | 567 | 7.68 | 62.77KD | ||||||||||||
| PtLAC21 | Potri.008G073800 | 574 | 9.23 | 63.889KD | PtLAC50 | Potri.019G088800 | 566 | 6.49 | 62.735KD | ||||||||||||
| PtLAC22 | Potri.009G034500 | 582 | 6.62 | 63.906KD | PtLAC51 | ||||||||||||||||
| f | 0.85422 | 0.09827 | 0.04261 | 0.00490 | |||||||||||||||||
| ω0 | 0.12861 | 1.00000 | 0.12861 | 1.00000 | |||||||||||||||||
| ω1 | 0.12861 | 1.00000 | 49.21745 | 49.21745 | |||||||||||||||||
| Model A null | 411 | −103,342.088418 | 1 | Not Allowed | |||||||||||||||||
| Subfamily III | Model A | 420 | Potri.016G107900 | 569 | 8.05 | 62.029KD | |||||||||||||||
| PtLAC16 | Potri.006G097000 | 558 | 9.52 | 61.311KD | PtLAC45 | Potri.016G112000 | 557 | 9.3 | 60.916KD | ||||||||||||
| PtLAC17 | Potri.006G097100 | 559 | 9.56 | 61.285KD | PtLAC46 | Potri.016G112100 | 0.09121564 | 0.093079.53 | 0.0105462.027KD | ||||||||||||
| PtLAC18 | Potri.007G023300 | 559 | 9.56 | 61.426KD | PtLAC47 | Potri.019G088500 | 567 | ||||||||||||||
| 7.01 | 62.76KD | ω0 | 0.1271 | 1.00000 | 0.12713 | 1.00000 | |||||||||||||||
| ω1 | 0.12713 | 1.00000 | 7.23646 | 7.23646 | |||||||||||||||||
| Model A null | 419 | −105,412.180374 | 1 | Not Allowed | |||||||||||||||||
| Subfamily V | Model A | 420 | −105,411.671727 | 6.82632 | Site class | 0 | 1 | 2a | 2b | Model A vs.Model A null | p | < 0.01 | 2 V 33 H 50 R 179 I 252 R ( | p | > 0.95) | Potri.019G088900 | |||||
| f | 567 | 6.78 | 62.919KD | ||||||||||||||||||
| 0.73746 | 0.08362 | 0.16070 | 0.01822 | PtLAC23 | Potri.009G042500 | 576 | 9.35 | 63.523KD | PtLAC52 | Potri.019G121700 | |||||||||||
| 576 | 8.23 | 63.9KD | |||||||||||||||||||
| ω0 | 0.12711 | 1.00000 | PtLAC24 | Potri.009G102700 | 556 | 9.03 | 60.74KD | PtLAC53 | Potri.019G124300 | 576 | 8.23 | 63.888KD | |||||||||
| 0.12711 | 1.00000 | ||||||||||||||||||||
| ω1 | 0.12711 | 1.00000 | 9.12388 | 9.12388 | PtLAC25 | Potri.009G156600 | |||||||||||||||
| Model A null | 561 | 8.91 | 62.735KD | * PtLAC1-fr | Potri.001G243200 | 259 | |||||||||||||||
| 419 | −105,415.084887 | 1 | Not Allowed | PtLAC26 | Potri.009G156800 | 576 | 9.2 | 63.512KD | * PtLAC2-fr | Potri.001G401000 | 215 | ||||||||||
| Subfamily VI | Model A | 420 | −105,420.856612 | 3.31621 | Site class | 0 | 1 | 2a | 2b | Model A vs.Model A null | p | > 0.05 | Not Found | PtLAC27 | Potri.010G183500 | 576 | 9.31 | 63.481KD | * PtLAC3-fr | Potri.016G083300 | |
| 147 | |||||||||||||||||||||
| f | 0.85708 | 0.09723 | 0.04103 | 0.00466 | PtLAC28 | Potri.010G183600 | 582 | 7.63 | 64.164KD | * PtLAC4-fr | Potri.016G083400 | 189 | |||||||||
| PtLAC29 | Potri.010G193100 | 575 | 8.75 | 63.672KD | * PtLAC5-fr | Potri.T096500 | 183 | ||||||||||||||
* It is considered as a fragment of pseudogene in this paper.
3. Evolutionary Analysis of the Laccase Gene Family in Terrestrial Plants
Eight terrestrial plants with completed whole-genome sequencing were used to represent different evolutionary stages to study the evolutionary course of laccase. Laccases were excavated in the genomes of these species for comprehensive phylogenetic analysis. The examined plant species include one moss (P. patens), one fern (Selaginellae moellendorfii), one gymnosperm (P. taeda), one archaic angiosperm (A. trichopoda), one monocot (O. sativa), and three dicots (A. thaliana, V. vinifera, and G. max). Four laccase genes were present in the bryophyte P. patens. No laccase gene was identified in the whole genomes of green algae (Ostreococcus lucimarinus) and Micromonas sp. Therefore, laccase might have originated from the plant’s landing process. The numbers of laccase genes in S. moellendorfii, P. taeda, and A. trichopoda were 11, 11, and 13, respectively. In total, 29, 17, 35 and 52 laccases were present in the monocot O. sativa, dicot A. thaliana, V. vinifera, and G. max, respectively. The gene family size of laccase in species indicated that this gene family began to expand during the evolution of angiosperms.
The cladogram based on the phylogenetic tree is shown in Figure 1a. In terms of the classification criteria of Arabidopsis laccase and location on the phylogenetic tree, the 221 land plant laccases were clearly divided into seven branches of Subfamilies I–VII (support rate > 65%) [19][47]. Subfamilies I–V contained gymnosperms and angiosperms. Subfamily VI is a unique branch of angiosperms. Subfamily VII was a basal branch and contained ferns and angiosperms. Gymnosperm laccases might have been lost in Subfamilies II, VI, and VII. According to the ML (Maximum likelihood) tree, the ancestors of vascular plants had at least two laccase genes. The fern laccase ancestor might have lost one copy. This copy produced two duplicates before it was lost. One of the duplicates was lost in the subsequent evolutionary process. Two of the five laccase ancestors of seed plants experienced duplication events. Eventually, eight ancient laccase genes were present. Five of the eight laccase gene ancestors might be lost in gymnosperms. Gymnosperms only had three ancient laccase genes (Figure 1b).
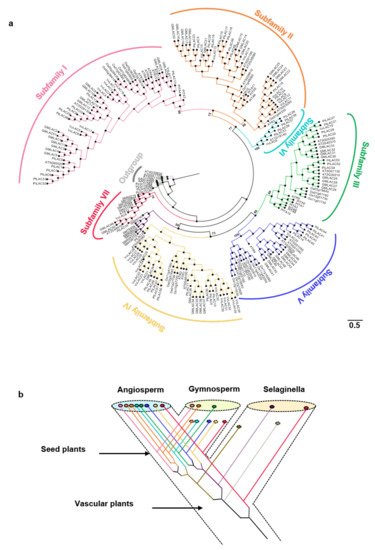

Figure 1. (a) Phylogenetic tree for the PtLACs and other plant laccases. The phylogenetic relationship was constructed using the maximum likelihood procedure on the basis of the amino acids of the Arabidopsis (AT), Amborella trichopoda (
| ω0 | |||||||||||||
| 0.12743 | |||||||||||||
| 1.00000 | |||||||||||||
| 0.12743 | |||||||||||||
| 1.00000 | |||||||||||||
| ω1 | |||||||||||||
| 0.12743 | |||||||||||||
| 1.00000 | 4.04102 | 4.04102 | |||||||||||
| Model A null | 419 | −105,422.514717 | 1 | Not Allowed | |||||||||
| Subfamily VII | Model A | 420 | −105,423.378874 | 0.05589 | Site class | 0 | 1 | 2a | 2b | Model A vs.Model A null | p | > 0.05 | Not Found |
| f | 0.00193 | 0.00022 | 0.89619 | 0.10167 | |||||||||
| ω0 | 0.12745 | 1.00000 | 0.12745 | 1.00000 | |||||||||
| ω1 | 0.12745 | 1.00000 | 5.99537 | 5.99537 | |||||||||
| Model A null | 419 | −105,423.406821 | 1 | Not Allowed |
Note: 2∆(LNL): Log likelihood difference between models using the c2-test.
5. Formation Mechanism Analysis of the Laccase Gene Family in P. trichocarpa
The salicoid genome duplication events occurred at least three times. The most recent one occurred between 60 and 65 Mya [16]. The chromosome rearrangement of poplar after the most recent whole-genome duplication (WGD) event is shown in Figure 2. Collinear regions between two chromosomes were shaded grey and connected with lines. These regions were used to represent homologous chromosomal fragments produced by segment-duplicated fragments. In total, 53 full-length PtLACs were located on 15 chromosomes of P. trichocarpa. The distribution of PtLACs on chromosomes was uneven. No laccase or laccase fragment was observed on chromosomes 2, 3, 17, and 18. PtLACs produced by tandem duplication formed 11 tightly connected clusters (clusters 1–11), and these gene clusters were distributed in certain regions of chromosomes 10, 11, 15, 16, and 19 with 2–5 cluster members. The gene cluster was generated by the tandem duplication of its core gene, which could be determined from the sequence similarity between cluster members. Clusters are shown in the box of Figure 2. PtLAC4/23, PtLAC14/40, and PtLAC34/53 were located in homologous chromosomal segments, which indicated that they were produced by WGD. Gene clusters 1/7, 3/10, and 4/6 were located in homologous chromosomal fragments, and these three pairs of gene clusters were also generated by WGD. Cluster 8 in the homologous chromosome segment did not correspond to the laccase gene cluster, but PtLAC33. Cluster 9 did not correspond to the laccase gene cluster instead of PtLAC14. This result indicated that the duplicated gene copies tended to be lost after genome-wide duplication events. P. trichocarpa produced 56.6% (30 of 53) of laccase members through tandem duplication. Therefore, tandem duplication was considered to be the main expansion formation of the laccase gene family.
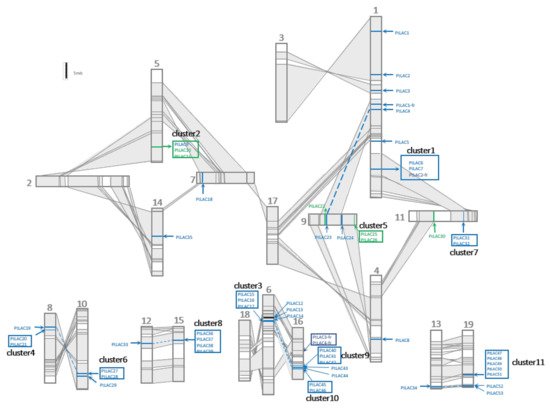

Figure 2. Genomic localization of P. trichocarpa laccase gene family. Schematic view of chromosome reorganization by the most recent whole-genome duplication in Populus. Regions that were assumed to correspond to homologous genome blocks were shaded grey and connected by lines. Paralogous LAC genes and clusters were indicated by dashed lines within grey-shaded trapezoids. The genes for functional research in this paper are marked in green.
6. Gene Structure Analysis and Motif Detection of PtLACs
The conserved motif of PtLACs and the details of the motif were analyzed using the MEME website to characterize the protein sequence (Figure 3). The results showed that PtLAC genes with close phylogenetic relationships contained similar exon/intron patterns, including the number of exons, exon length, intron phases, and splicing patterns. The length of the amino acid motif of PtLACs was 15–50, and the number of motifs was 9–11. The height of the amino acid residue letter at each position represented the degree of conservation. Results showed that P. trichocarpa laccase had four conserved Cu2+ binding motifs HXHG, HXH, HXXHXH, and HCHXXXHXXXXM/L. PtLAC18, PtLAC33, PtLAC36, and PtLAC37 had 11 motifs, whereas PtLAC39 had nine motifs. In addition, the members of cluster 8 contained highly conserved motifs, which had one more motif 8 than the members in other clusters. Since exon loss/gain and sequence polymorphisms were identified in the PtLAC genes, there is likely functional diversity in the gene family as well.
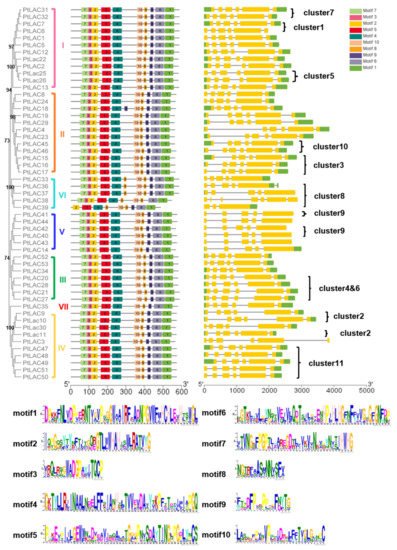

Figure 3. Motif distribution and gene structure of PtLACs. The multiple alignment of 53 full-length PtLAC amino acid sequences was performed by MAFFT v7. Each PtLACs subfamily is marked. MEME identification of conserved motifs in PtLAC proteins. Each motif is represented by a different color block. The sequence logos of 10 conserved motifs were named 1–10. Yellow rectangles represented exons and horizontal lines represented introns. The scale estimates the lengths of exons and introns. The scale bar represents 100 amino acid residues for protein sequences and 1.0 kb for gene structure.
The structural features of each of the PtLAC genes in subfamilies are shown in Figure 3. Structural analyses of all of the PtLAC genes in P. trichocarpa revealed that the number of exons varied from three to seven. The structural analysis of 53 PtLAC genes revealed that genes from the same subfamily, particularly Subfamilies I–IV, contained similar gene structures. The members of Subfamily V contained five exons. The members of Subfamily VI had 3–7 exons. There were no intronless genes. The exon/intron pattern belonging to the same subfamily was more similar than those from different subfamilies (Figure 3). This suggested that genes with similar functions may be clustered in the same group.
