The circadian clock is a fundamental biological timing mechanism that generates nearly 24 h rhythms of physiology and behaviors, including sleep/wake cycles, hormone secretion, and metabolism. Evolutionarily, the endogenous clock is thought to confer living organisms, including humans, with survival benefits by adapting internal rhythms to the day and night cycles of the local environment. Mirroring the evolutionary fitness bestowed by the circadian clock, daily mismatches between the internal body clock and environmental cycles, such as irregular work (e.g., night shift work) and life schedules (e.g., jet lag, mistimed eating), have been recognized to increase the risk of cardiac, metabolic, and neurological diseases. Moreover, increasing numbers of studies with cellular and animal models have detected the presence of functional circadian oscillators at multiple levels, ranging from individual neurons and fibroblasts to brain and peripheral organs. These oscillators are tightly coupled to timely modulate cellular and bodily responses to physiological and metabolic cues.
- circadian clock
- circadian disruption
- SCN
- brain clocks
- peripheral clocks
- redox metabolism
1. Introduction
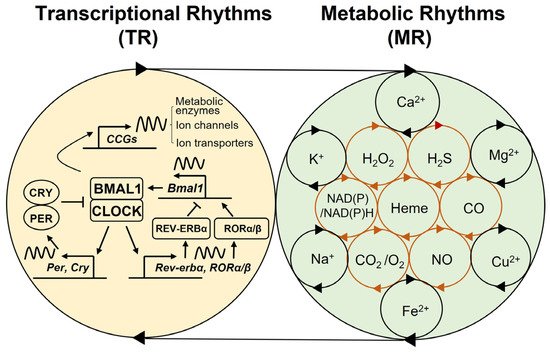
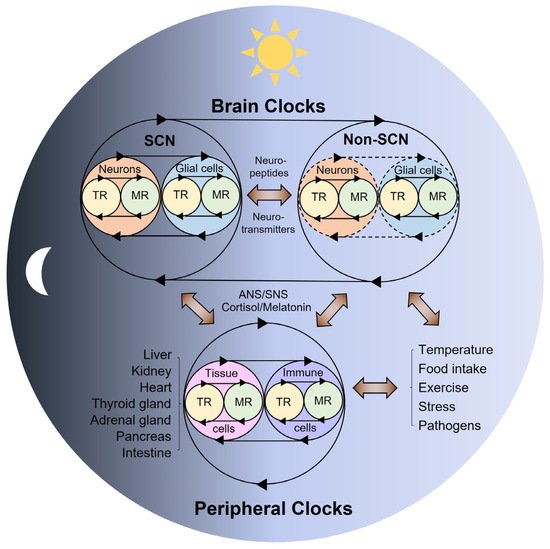
2. Multi-Modal Mechanisms of Circadian Physiology
3. Conclusions
References
- Dunlap, J.C. Molecular bases for circadian clocks. Cell 1999, 96, 271–290.
- Bass, J.; Lazar, M.A. Circadian time signatures of fitness and disease. Science 2016, 354, 994–999.
- Chaix, A.; Zarrinpar, A.; Panda, S. The circadian coordination of cell biology. J. Cell Biol. 2016, 215, 15–25.
- Kalmbach, D.A.; Schneider, L.D.; Cheung, J.; Bertrand, S.J.; Kariharan, T.; Pack, A.I.; Gehrman, P.R. Genetic Basis of Chronotype in Humans: Insights From Three Landmark GWAS. Sleep 2017, 40, zsw048.
- Refinetti, R.; Wassmer, T.; Basu, P.; Cherukalady, R.; Pandey, V.K.; Singaravel, M.; Giannetto, C.; Piccione, G. Variability of behavioral chronotypes of 16 mammalian species under controlled conditions. Physiol. Behav. 2016, 161, 53–59.
- Bloch, G.; Barnes, B.M.; Gerkema, M.P.; Helm, B. Animal activity around the clock with no overt circadian rhythms: Patterns, mechanisms and adaptive value. Proc. Biol. Sci. 2013, 280, 20130019.
- Dominoni, D.M.; Borniger, J.C.; Nelson, R.J. Light at night, clocks and health: From humans to wild organisms. Biol. Lett. 2016, 12, 20160015.
- Cederroth, C.R.; Albrecht, U.; Bass, J.; Brown, S.A.; Dyhrfjeld-Johnsen, J.; Gachon, F.; Green, C.B.; Hastings, M.H.; Helfrich-Förster, C.; Hogenesch, J.B.; et al. Medicine in the Fourth Dimension. Cell Metab. 2019, 30, 238–250.
- Patke, A.; Young, M.W.; Axelrod, S. Molecular mechanisms and physiological importance of circadian rhythms. Nat. Rev. Mol. Cell. Biol. 2020, 21, 67–84.
- Lee, Y.; Field, J.M.; Sehgal, A. Circadian Rhythms, Disease and Chronotherapy. J. Biol. Rhythms 2021, 7487304211044301.
- Logan, R.W.; McClung, C.A. Rhythms of life: Circadian disruption and brain disorders across the lifespan. Nat. Rev. Neurosci. 2019, 20, 49–65.
- Scheiermann, C.; Gibbs, J.; Ince, L.; Loudon, A. Clocking in to immunity. Nat. Rev. Immunol. 2018, 18, 423–437.
- Segers, A.; Depoortere, I. Circadian clocks in the digestive system. Nat. Rev. Gastroenterol. Hepatol. 2021, 18, 239–251.
- Masri, S.; Sassone-Corsi, P. The emerging link between cancer, metabolism, and circadian rhythms. Nat. Med. 2018, 24, 1795–1803.
- Aiello, I.; Fedele, M.L.M.; Roman, F.; Marpegan, L.; Caldart, C.; Chiesa, J.J.; Golombek, D.A.; Finkielstein, C.V.; Paladino, N. Circadian disruption promotes tumor-immune microenvironment remodeling favoring tumor cell proliferation. Sci. Adv. 2020, 6, eaaz4530.
- Hadadi, E.; Taylor, W.; Li, X.M.; Aslan, Y.; Villote, M.; Riviere, J.; Duvallet, G.; Auriau, C.; Dulong, S.; Raymond-Letron, I.; et al. Chronic circadian disruption modulates breast cancer stemness and immune microenvironment to drive metastasis in mice. Nat. Commun. 2020, 11, 3193.
- Mattis, J.; Sehgal, A. Circadian Rhythms, Sleep, and Disorders of Aging. Trends Endocrinol. Metab. 2016, 27, 192–203.
- Shafi, A.A.; McNair, C.M.; McCann, J.J.; Alshalalfa, M.; Shostak, A.; Severson, T.M.; Zhu, Y.; Bergman, A.; Gordon, N.; Mandigo, A.C.; et al. The circadian cryptochrome, CRY1, is a pro-tumorigenic factor that rhythmically modulates DNA repair. Nat. Commun. 2021, 12, 401.
- Lee, Y.; Lahens, N.F.; Zhang, S.; Bedont, J.; Field, J.M.; Sehgal, A. G1/S cell cycle regulators mediate effects of circadian dysregulation on tumor growth and provide targets for timed anticancer treatment. PLoS Biol. 2019, 17, e3000228.
- Lee, Y.; Fong, S.Y.; Shon, J.; Zhang, S.L.; Brooks, R.; Lahens, N.F.; Chen, D.; Dang, C.V.; Field, J.M.; Sehgal, A. Time-of-day specificity of anticancer drugs may be mediated by circadian regulation of the cell cycle. Sci. Adv. 2021, 7, eabd2645.
- Diallo, A.B.; Coiffard, B.; Leone, M.; Mezouar, S.; Mege, J.L. For Whom the Clock Ticks: Clinical Chronobiology for Infectious Diseases. Front. Immunol. 2020, 11, 1457.
- Gabriel, B.M.; Zierath, J.R. Circadian rhythms and exercise—Re-setting the clock in metabolic disease. Nat. Rev. Endocrinol. 2019, 15, 197–206.
- Lewis, P.; Oster, H.; Korf, H.W.; Foster, R.G.; Erren, T.C. Food as a circadian time cue—Evidence from human studies. Nat. Rev. Endocrinol. 2020, 16, 213–223.
- Johnson, C.H.; Zhao, C.; Xu, Y.; Mori, T. Timing the day: What makes bacterial clocks tick? Nat. Rev. Microbiol. 2017, 15, 232–242.
- Takahashi, J.S. Transcriptional architecture of the mammalian circadian clock. Nat. Rev. Genet. 2017, 18, 164–179.
- Koronowski, K.B.; Sassone-Corsi, P. Communicating clocks shape circadian homeostasis. Science 2021, 371, eabd0951.
- Anafi, R.C.; Lee, Y.; Sato, T.K.; Venkataraman, A.; Ramanathan, C.; Kavakli, I.H.; Hughes, M.E.; Baggs, J.E.; Growe, J.; Liu, A.C.; et al. Machine learning helps identify CHRONO as a circadian clock component. PLoS Biol. 2014, 12, e1001840.
- Lee, Y.; Lee, J.; Kwon, I.; Nakajima, Y.; Ohmiya, Y.; Son, G.H.; Lee, K.H.; Kim, K. Coactivation of the CLOCK-BMAL1 complex by CBP mediates resetting of the circadian clock. J. Cell Sci. 2010, 123, 3547–3557.
- Lee, J.; Lee, Y.; Lee, M.J.; Park, E.; Kang, S.H.; Chung, C.H.; Lee, K.H.; Kim, K. Dual modification of BMAL1 by SUMO2/3 and ubiquitin promotes circadian activation of the CLOCK/BMAL1 complex. Mol. Cell. Biol. 2008, 28, 6056–6065.
- Lee, Y.; Shen, Y.; Francey, L.J.; Ramanathan, C.; Sehgal, A.; Liu, A.C.; Hogenesch, J.B. The NRON complex controls circadian clock function through regulated PER and CRY nuclear translocation. Sci. Rep. 2019, 9, 11883.
- Lee, Y.; Jang, A.R.; Francey, L.J.; Sehgal, A.; Hogenesch, J.B. KPNB1 mediates PER/CRY nuclear translocation and circadian clock function. eLife 2015, 4, e08647.
- Lee, Y.; Chun, S.K.; Kim, K. Sumoylation controls CLOCK-BMAL1-mediated clock resetting via CBP recruitment in nuclear transcriptional foci. Biochim. Biophys. Acta 2015, 1853, 2697–2708.
- Korge, S.; Maier, B.; Bruning, F.; Ehrhardt, L.; Korte, T.; Mann, M.; Herrmann, A.; Robles, M.S.; Kramer, A. The non-classical nuclear import carrier Transportin 1 modulates circadian rhythms through its effect on PER1 nuclear localization. PLoS Genet. 2018, 14, e1007189.
- Zhang, R.; Lahens, N.F.; Ballance, H.I.; Hughes, M.E.; Hogenesch, J.B. A circadian gene expression atlas in mammals: Implications for biology and medicine. Proc. Natl. Acad. Sci. USA 2014, 111, 16219–16224.
- Mure, L.S.; Le, H.D.; Benegiamo, G.; Chang, M.W.; Rios, L.; Jillani, N.; Ngotho, M.; Kariuki, T.; Dkhissi-Benyahya, O.; Cooper, H.M.; et al. Diurnal transcriptome atlas of a primate across major neural and peripheral tissues. Science 2018, 359, eaao0318.
- Ruben, M.D.; Wu, G.; Smith, D.F.; Schmidt, R.E.; Francey, L.J.; Lee, Y.Y.; Anafi, R.C.; Hogenesch, J.B. A database of tissue-specific rhythmically expressed human genes has potential applications in circadian medicine. Sci. Transl. Med. 2018, 10, eaat8806.
- Thaiss, C.A.; Zeevi, D.; Levy, M.; Zilberman-Schapira, G.; Suez, J.; Tengeler, A.C.; Abramson, L.; Katz, M.N.; Korem, T.; Zmora, N.; et al. Transkingdom control of microbiota diurnal oscillations promotes metabolic homeostasis. Cell 2014, 159, 514–529.
- Dyar, K.A.; Lutter, D.; Artati, A.; Ceglia, N.J.; Liu, Y.; Armenta, D.; Jastroch, M.; Schneider, S.; de Mateo, S.; Cervantes, M.; et al. Atlas of Circadian Metabolism Reveals System-wide Coordination and Communication between Clocks. Cell 2018, 174, 1571–1585.e11.
- Solanas, G.; Peixoto, F.O.; Perdiguero, E.; Jardi, M.; Ruiz-Bonilla, V.; Datta, D.; Symeonidi, A.; Castellanos, A.; Welz, P.S.; Caballero, J.M.; et al. Aged Stem Cells Reprogram Their Daily Rhythmic Functions to Adapt to Stress. Cell 2017, 170, 678–692.e20.
- Wang, J.; Mauvoisin, D.; Martin, E.; Atger, F.; Galindo, A.N.; Dayon, L.; Sizzano, F.; Palini, A.; Kussmann, M.; Waridel, P.; et al. Nuclear Proteomics Uncovers Diurnal Regulatory Landscapes in Mouse Liver. Cell Metab. 2017, 25, 102–117.
- Wang, Y.; Song, L.; Liu, M.; Ge, R.; Zhou, Q.; Liu, W.; Li, R.; Qie, J.; Zhen, B.; Wang, Y.; et al. A proteomics landscape of circadian clock in mouse liver. Nat. Commun. 2018, 9, 1553.
- Robles, M.S.; Humphrey, S.J.; Mann, M. Phosphorylation Is a Central Mechanism for Circadian Control of Metabolism and Physiology. Cell Metab. 2017, 25, 118–127.
- Chiang, C.K.; Xu, B.; Mehta, N.; Mayne, J.; Sun, W.Y.; Cheng, K.; Ning, Z.; Dong, J.; Zou, H.; Cheng, H.M.; et al. Phosphoproteome Profiling Reveals Circadian Clock Regulation of Posttranslational Modifications in the Murine Hippocampus. Front. Neurol. 2017, 8, 110.
- Noya, S.B.; Colameo, D.; Bruning, F.; Spinnler, A.; Mircsof, D.; Opitz, L.; Mann, M.; Tyagarajan, S.K.; Robles, M.S.; Brown, S.A. The forebrain synaptic transcriptome is organized by clocks but its proteome is driven by sleep. Science 2019, 366, eaav2642.
- Malik, D.M.; Paschos, G.K.; Sehgal, A.; Weljie, A.M. Circadian and Sleep Metabolomics Across Species. J. Mol. Biol. 2020, 432, 3578–3610.
- Hofman, M.A.; Fliers, E.; Goudsmit, E.; Swaab, D.F. Morphometric analysis of the suprachiasmatic and paraventricular nuclei in the human brain: Sex differences and age-dependent changes. J. Anat. 1988, 160, 127–143.
- Hofman, M.A.; Swaab, D.F. A brain for all seasons: Cellular and molecular mechanisms of photoperiodic plasticity. Prog. Brain Res. 2002, 138, 255–280.
- Hofman, M.A.; Swaab, D.F. Living by the clock: The circadian pacemaker in older people. Ageing Res. Rev. 2006, 5, 33–51.
- Harmar, A.J.; Marston, H.M.; Shen, S.; Spratt, C.; West, K.M.; Sheward, W.J.; Morrison, C.F.; Dorin, J.R.; Piggins, H.D.; Reubi, J.C.; et al. The VPAC(2) receptor is essential for circadian function in the mouse suprachiasmatic nuclei. Cell 2002, 109, 497–508.
- Brown, T.M.; Colwell, C.S.; Waschek, J.A.; Piggins, H.D. Disrupted neuronal activity rhythms in the suprachiasmatic nuclei of vasoactive intestinal polypeptide-deficient mice. J. Neurophysiol. 2007, 97, 2553–2558.
- Yamaguchi, Y.; Suzuki, T.; Mizoro, Y.; Kori, H.; Okada, K.; Chen, Y.; Fustin, J.M.; Yamazaki, F.; Mizuguchi, N.; Zhang, J.; et al. Mice genetically deficient in vasopressin V1a and V1b receptors are resistant to jet lag. Science 2013, 342, 85–90.
- Mieda, M.; Ono, D.; Hasegawa, E.; Okamoto, H.; Honma, K.; Honma, S.; Sakurai, T. Cellular clocks in AVP neurons of the SCN are critical for interneuronal coupling regulating circadian behavior rhythm. Neuron 2015, 85, 1103–1116.
- Patton, A.P.; Edwards, M.D.; Smyllie, N.J.; Hamnett, R.; Chesham, J.E.; Brancaccio, M.; Maywood, E.S.; Hastings, M.H. The VIP-VPAC2 neuropeptidergic axis is a cellular pacemaking hub of the suprachiasmatic nucleus circadian circuit. Nat. Commun. 2020, 11, 3394.
- Todd, W.D.; Venner, A.; Anaclet, C.; Broadhurst, R.Y.; De Luca, R.; Bandaru, S.S.; Issokson, L.; Hablitz, L.M.; Cravetchi, O.; Arrigoni, E.; et al. Suprachiasmatic VIP neurons are required for normal circadian rhythmicity and comprised of molecularly distinct subpopulations. Nat. Commun. 2020, 11, 4410.
- Wen, S.; Ma, D.; Zhao, M.; Xie, L.; Wu, Q.; Gou, L.; Zhu, C.; Fan, Y.; Wang, H.; Yan, J. Spatiotemporal single-cell analysis of gene expression in the mouse suprachiasmatic nucleus. Nat. Neurosci. 2020, 23, 456–467.
- Park, J.; Zhu, H.; O’Sullivan, S.; Ogunnaike, B.A.; Weaver, D.R.; Schwaber, J.S.; Vadigepalli, R. Single-Cell Transcriptional Analysis Reveals Novel Neuronal Phenotypes and Interaction Networks Involved in the Central Circadian Clock. Front. Neurosci. 2016, 10, 481.
- Morris, E.L.; Patton, A.P.; Chesham, J.E.; Crisp, A.; Adamson, A.; Hastings, M.H. Single-cell transcriptomics of suprachiasmatic nuclei reveal a Prokineticin-driven circadian network. EMBO J. 2021, 40, e108614.
- Brancaccio, M.; Patton, A.P.; Chesham, J.E.; Maywood, E.S.; Hastings, M.H. Astrocytes Control Circadian Timekeeping in the Suprachiasmatic Nucleus via Glutamatergic Signaling. Neuron 2017, 93, 1420–1435.e5.
- Tso, C.F.; Simon, T.; Greenlaw, A.C.; Puri, T.; Mieda, M.; Herzog, E.D. Astrocytes Regulate Daily Rhythms in the Suprachiasmatic Nucleus and Behavior. Curr. Biol. 2017, 27, 1055–1061.
- Brancaccio, M.; Edwards, M.D.; Patton, A.P.; Smyllie, N.J.; Chesham, J.E.; Maywood, E.S.; Hastings, M.H. Cell-autonomous clock of astrocytes drives circadian behavior in mammals. Science 2019, 363, 187–192.
- Buijs, R.M.; Guzman Ruiz, M.A.; Mendez Hernandez, R.; Rodriguez Cortes, B. The suprachiasmatic nucleus; a responsive clock regulating homeostasis by daily changing the setpoints of physiological parameters. Auton. Neurosci. 2019, 218, 43–50.
- Gizowski, C.; Zaelzer, C.; Bourque, C.W. Clock-driven vasopressin neurotransmission mediates anticipatory thirst prior to sleep. Nature 2016, 537, 685–688.
- Paul, S.; Hanna, L.; Harding, C.; Hayter, E.A.; Walmsley, L.; Bechtold, D.A.; Brown, T.M. Output from VIP cells of the mammalian central clock regulates daily physiological rhythms. Nat. Commun. 2020, 11, 1453.
- Collins, B.; Pierre-Ferrer, S.; Muheim, C.; Lukacsovich, D.; Cai, Y.; Spinnler, A.; Herrera, C.G.; Wen, S.; Winterer, J.; Belle, M.D.C.; et al. Circadian VIPergic Neurons of the Suprachiasmatic Nuclei Sculpt the Sleep-Wake Cycle. Neuron 2020, 108, 486–499.e5.
- Ishida, A.; Mutoh, T.; Ueyama, T.; Bando, H.; Masubuchi, S.; Nakahara, D.; Tsujimoto, G.; Okamura, H. Light activates the adrenal gland: Timing of gene expression and glucocorticoid release. Cell Metab. 2005, 2, 297–307.
- Buijs, R.M.; Kalsbeek, A. Hypothalamic integration of central and peripheral clocks. Nat. Rev. Neurosci. 2001, 2, 521–526.
- Russell, G.; Lightman, S. The human stress response. Nat. Rev. Endocrinol. 2019, 15, 525–534.
- Buijs, F.N.; León-Mercado, L.; Guzmán-Ruiz, M.; Guerrero-Vargas, N.N.; Romo-Nava, F.; Buijs, R.M. The Circadian System: A Regulatory Feedback Network of Periphery and Brain. Physiology 2016, 31, 170–181.
- Firsov, D.; Bonny, O. Circadian rhythms and the kidney. Nat. Rev. Nephrol. 2018, 14, 626–635.
- Ikegami, K.; Refetoff, S.; Van Cauter, E.; Yoshimura, T. Interconnection between circadian clocks and thyroid function. Nat. Rev. Endocrinol. 2019, 15, 590–600.
- Stenvers, D.J.; Scheer, F.; Schrauwen, P.; la Fleur, S.E.; Kalsbeek, A. Circadian clocks and insulin resistance. Nat. Rev. Endocrinol. 2019, 15, 75–89.
- Challet, E. The circadian regulation of food intake. Nat. Rev. Endocrinol. 2019, 15, 393–405.
- Tahara, Y.; Shibata, S. Circadian rhythms of liver physiology and disease: Experimental and clinical evidence. Nat. Rev. Gastroenterol. Hepatol. 2016, 13, 217–226.
- Hasegawa, S.; Fukushima, H.; Hosoda, H.; Serita, T.; Ishikawa, R.; Rokukawa, T.; Kawahara-Miki, R.; Zhang, Y.; Ohta, M.; Okada, S.; et al. Hippocampal clock regulates memory retrieval via Dopamine and PKA-induced GluA1 phosphorylation. Nat. Commun. 2019, 10, 5766.
- Myung, J.; Schmal, C.; Hong, S.; Tsukizawa, Y.; Rose, P.; Zhang, Y.; Holtzman, M.J.; De Schutter, E.; Herzel, H.; Bordyugov, G.; et al. The choroid plexus is an important circadian clock component. Nat. Commun. 2018, 9, 1062.
- Son, G.H.; Chung, S.; Choe, H.K.; Kim, H.D.; Baik, S.M.; Lee, H.; Lee, H.W.; Choi, S.; Sun, W.; Kim, H.; et al. Adrenal peripheral clock controls the autonomous circadian rhythm of glucocorticoid by causing rhythmic steroid production. Proc. Natl. Acad. Sci. USA 2008, 105, 20970–20975.
- Myung, J.; Wu, M.Y.; Lee, C.Y.; Rahim, A.R.; Truong, V.H.; Wu, D.; Piggins, H.D.; Wu, M.S. The Kidney Clock Contributes to Timekeeping by the Master Circadian Clock. Int. J. Mol. Sci. 2019, 20, 2765.
- Sinturel, F.; Gos, P.; Petrenko, V.; Hagedorn, C.; Kreppel, F.; Storch, K.F.; Knutti, D.; Liani, A.; Weitz, C.; Emmenegger, Y.; et al. Circadian hepatocyte clocks keep synchrony in the absence of a master pacemaker in the suprachiasmatic nucleus or other extrahepatic clocks. Genes Dev. 2021, 35, 329–334.
- Bano-Otalora, B.; Piggins, H.D. Contributions of the lateral habenula to circadian timekeeping. Pharmacol. Biochem. Behav. 2017, 162, 46–54.
- Van Drunen, R.; Eckel-Mahan, K. Circadian Rhythms of the Hypothalamus: From Function to Physiology. Clocks Sleep 2021, 3, 189–226.
- Antle, M.C.; Mistlberger, R.E. Circadian clock resetting by sleep deprivation without exercise in the Syrian hamster. J. Neurosci. 2000, 20, 9326–9332.
- Mendoza, J. Circadian clocks: Setting time by food. J. Neuroendocrinol. 2007, 19, 127–137.
- Marchant, E.G.; Mistlberger, R.E. Morphine phase-shifts circadian rhythms in mice: Role of behavioural activation. Neuroreport 1995, 7, 209–212.
- Guerrero-Vargas, N.N.; Salgado-Delgado, R.; Basualdo, M.e.C.; García, J.; Guzmán-Ruiz, M.; Carrero, J.C.; Escobar, C.; Buijs, R.M. Reciprocal interaction between the suprachiasmatic nucleus and the immune system tunes down the inflammatory response to lipopolysaccharide. J. Neuroimmunol. 2014, 273, 22–30.
- Saderi, N.; Salgado-Delgado, R.; Avendaño-Pradel, R.; Basualdo, M.e.C.; Ferri, G.L.; Chávez-Macías, L.; Roblera, J.E.; Escobar, C.; Buijs, R.M. NPY and VGF immunoreactivity increased in the arcuate nucleus, but decreased in the nucleus of the Tractus Solitarius, of type-II diabetic patients. PLoS ONE 2012, 7, e40070.
- Buijs, F.N.; Cazarez, F.; Basualdo, M.C.; Scheer, F.A.; Perusquía, M.; Centurion, D.; Buijs, R.M. The suprachiasmatic nucleus is part of a neural feedback circuit adapting blood pressure response. Neuroscience 2014, 266, 197–207.
- Shioiri, T.; Takahashi, K.; Yamada, N.; Takahashi, S. Motor activity correlates negatively with free-running period, while positively with serotonin contents in SCN in free-running rats. Physiol. Behav. 1991, 49, 779–786.
- Yi, C.X.; van der Vliet, J.; Dai, J.; Yin, G.; Ru, L.; Buijs, R.M. Ventromedial arcuate nucleus communicates peripheral metabolic information to the suprachiasmatic nucleus. Endocrinology 2006, 147, 283–294.
- Acosta-Galvan, G.; Yi, C.X.; van der Vliet, J.; Jhamandas, J.H.; Panula, P.; Angeles-Castellanos, M.; Del Carmen Basualdo, M.; Escobar, C.; Buijs, R.M. Interaction between hypothalamic dorsomedial nucleus and the suprachiasmatic nucleus determines intensity of food anticipatory behavior. Proc. Natl. Acad. Sci. USA 2011, 108, 5813–5818.
- Thaiss, C.A.; Levy, M.; Korem, T.; Dohnalova, L.; Shapiro, H.; Jaitin, D.A.; David, E.; Winter, D.R.; Gury-BenAri, M.; Tatirovsky, E.; et al. Microbiota Diurnal Rhythmicity Programs Host Transcriptome Oscillations. Cell 2016, 167, 1495–1510.e12.
- Buhr, E.D.; Yoo, S.H.; Takahashi, J.S. Temperature as a universal resetting cue for mammalian circadian oscillators. Science 2010, 330, 379–385.
- Reinke, H.; Asher, G. Crosstalk between metabolism and circadian clocks. Nat. Rev. Mol. Cell Biol. 2019, 20, 227–241.
- Parmalee, N.L.; Aschner, M. Metals and Circadian Rhythms. Adv. Neurotoxicol. 2017, 1, 119–130.
- Rey, G.; Valekunja, U.K.; Feeney, K.A.; Wulund, L.; Milev, N.B.; Stangherlin, A.; Ansel-Bollepalli, L.; Velagapudi, V.; O’Neill, J.S.; Reddy, A.B. The Pentose Phosphate Pathway Regulates the Circadian Clock. Cell Metab. 2016, 24, 462–473.
- Ch, R.; Rey, G.; Ray, S.; Jha, P.K.; Driscoll, P.C.; Dos Santos, M.S.; Malik, D.M.; Lach, R.; Weljie, A.M.; MacRae, J.I.; et al. Rhythmic glucose metabolism regulates the redox circadian clockwork in human red blood cells. Nat. Commun. 2021, 12, 377.
