Fish gut represents a peculiar ecological niche where bacteria can transit and reside to play vital roles by producing bio-compounds with nutritional, immunomodulatory and other functions. This complex microbial ecosystem reflects several factors (environment, feeding regimen, fish species etc.). The objective of the present study was the identification of intestinal microbial strains able to produce molecules called biosurfactants (BSs) which were tested for surface and antibacterial activity in order to select a group of probiotic bacteria for aquaculture use. This works indicated that fish gut is a source of bioactive compounds which deserves to be explored for applicative purposes.
- grey mullets
- natural antibiotics
- gut microbiota
- probiotics
1. Enumeration of Bacteria and Colony Isolation
2. Screening of Bacteria for BS Production
| Strain | Fish Species |
Bacterial Affiliation |
GeneBank Accession Number |
Drop Collapse | E-24 (%) |
Surface TensionmN·m | −1 | BS Type |
|---|---|---|---|---|---|---|---|---|
| 1 | CR | Pseudomonas aeruginosa | MW369461 | +++ | 70.5 ± 9.1 | 36.5 ± 0.1 | Rhamnolipid | |
| 3 | CR | Pseudomonas aeruginosa | OK342256 | +++ | 68.0 ± 12.7 | 37.1 ± 0.1 | Rhamnolipid | |
| 5 | CR | Pseudomonas aeruginosa | OK342257 | +++ | 77.0 ± 0.0 | 36.9 ± 0.4 | Rhamnolipid | |
| 6 | CR | Pseudomonas aeruginosa | MW369462 | +++ | 56.4 ± 0.0 | 37.1 ± 0.1 | Less polar compound | |
| 8 | CR | Pseudomonas aeruginosa | OK342258 | ++ | 0.0 ± 0.0 | 37.1 ± 0.1 | nd | |
| 9 | CR | Pseudomonas aeruginosa | OK342259 | +++ | 57.7 ± 1.8 | 37.2 ± 0.3 | nd | |
| 10 | CR | Pseudomonas alcaligenes | MW369463 | + | 50.0 ± 1.8 | 37.2 ± 0.3 | nd | |
| 11 | CR | Aeromonas caviae | MW369464 | - | 0.0 ± 0.0 | 43.0 ± 0.1 | nd | |
| 12 | CR | - | - | +++ | 15.4 ± 21.8 | 36.9 ± 0.1 | nd | |
| 13 | CR | Pseudomonas aeruginosa | MW369465 | +++ | 51.3 ± 3.6 | 36.9 ± 0.1 | Rhamnolipid | |
| 15 | CR | Pseudomonas aeruginosa | MW369466 | +++ | 59.0 ± 3.6 | 36.6 ± 0.6 | Rhamnolipid | |
| 16 | CR | - | - | weak | 0.0 ± 0.0 | 35.35 ± 0.6 | nd | |
| 17 | CR | Pseudomonas mendocina | MW369467 | - | 20.5 ± 0 | nd | nd | |
| 18 | CR | Pseudomonas putida | OK342260 | weak | 33.3 ± 3.6 | 36.1 ± 0.1 | Less polar compound | |
| 19 | MC | Pseudomonas | sp. | OK342261 | + | 28.2 ± 3.6 | 35.2 ± 0.0 | Less polar compound |
| 20 | MC | Pseudomonas alcaliphila | MW369468 | weak | 0.0 ± 0.0 | 35.0 ± 0.4 | nd | |
| 21 | MC | - | - | weak | 25.6 ± 14.5 | nd | nd | |
| 22 | ||||||||
| sp. | ||||||||
| OK342267 | ||||||||
| + | ||||||||
| 0.0 ± 0.0 | ||||||||
| 39.9 ± 0.1 | ||||||||
| nd | ||||||||
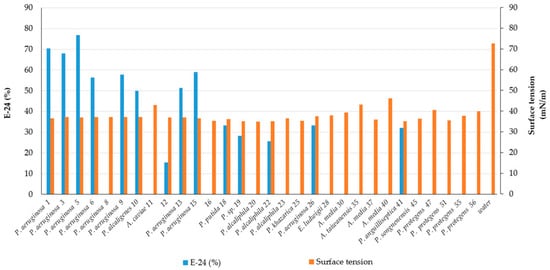
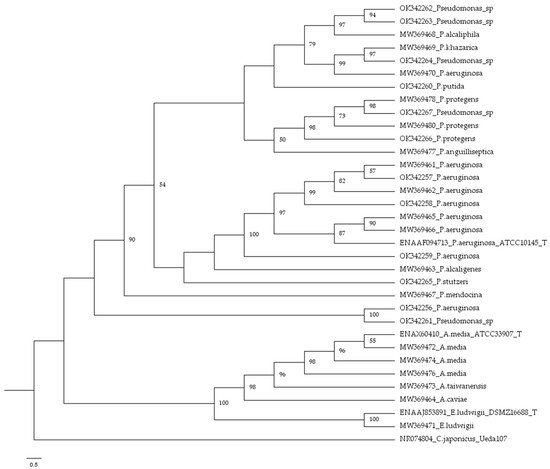
3. Bacterial Identification

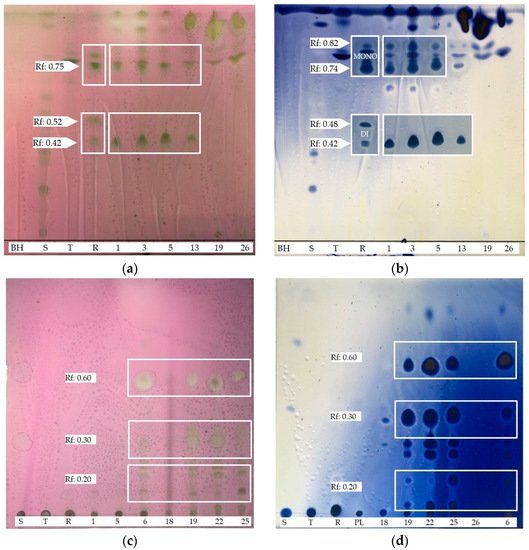
4. BSs yield and Thin Layer Chromatography (TLC)
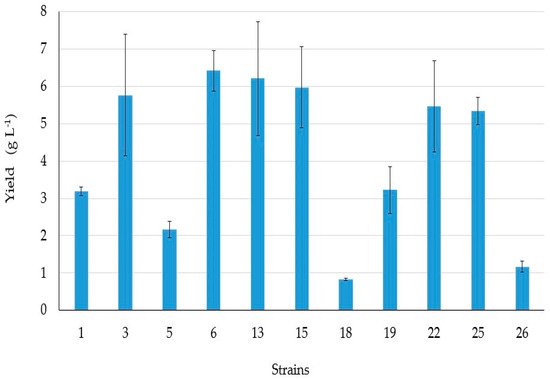

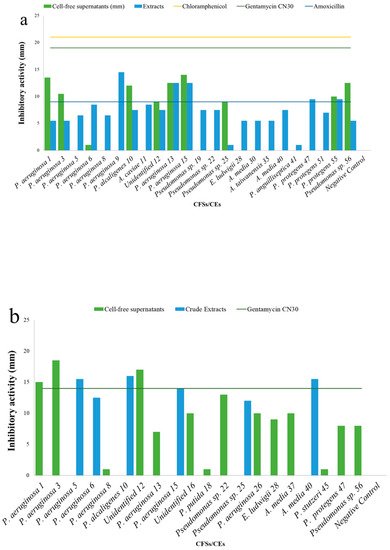
5. Antibacterial Activities
| Cell-Free Supernatants (CFSs) and Crude Extracts (CEs) (mm) | |||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Test | S. aureus | H1610 | P. mirabilis | H1643 | K. pneumoniae | H1637 | A. hydrophila | H1563 | |||||||||||
| CFSs | CEs | CFSs | CEs | CFSs | CEs | CFSs | CEs | ||||||||||||
| Pseudomonas aeruginosa | 1 | 13.5 ± 0.7 | 5.5 ± 0.7 | 15 ± 0.0 | - | - | 17.5 ± 0.7 | - | - | ||||||||||
| Pseudomonas aeruginosa | 3 | 10.5 ± 2.1 | 5.5 ± 0.7 | 18.5 ± 0.7 | - | - | 8.0 ± 0.0 | - | - | ||||||||||
| Pseudomonas aeruginosa | 5 | - | 5.5 ± 0.7 | - | 15.5 ± 0.7 | - | 6.0 ± 0.0 | - | - | ||||||||||
| Pseudomonas aeruginosa | 6 | + | 8.5 ± 0.7 | - | 12.5 ± 0.7 | - | 11.5 ± 0.7 | - | - | ||||||||||
| Pseudomonas aeruginosa | 8 | - | 6.5 ± 0.7 | + | - | - | 7.0 ± 0.0 | - | 6.5 ± 0.7 | ||||||||||
| Pseudomonas aeruginosa | 9 | - | 14.5 ± 0.7 | - | - | - | 12.0 ± 0.0 | - | - | ||||||||||
| Pseudomonas alcaligenes | 10 | 12 ± 0.0 | 7.5 ± 0.7 | - | 16.0 ± 0.0 | - | + | - | - | ||||||||||
| Aeromonas caviae | 11 | - | 8.5 ± 0.7 | - | - | - | + | - | - | ||||||||||
| Unidentified 12 | 9 ± 1.4 | 7.5 ± 0.7 | 17.0 ± 1.4 | - | - | 7.5 ± 0.7 | - | - | |||||||||||
| Pseudomonas aeruginosa | 13 | 12.5 ± 0.7 | 12.5 ± 0.7 | 7.0 ± 0.0 | - | - | 12.5 ± 0.7 | - | 7.0 ± 0.0 | ||||||||||
| Pseudomonas aeruginosa | 15 | 14 ± 0.0 | 12.5 ± 0.7 | - | 14.0 ± 0.0 | - | 12.5 ± 0.7 | - | - | ||||||||||
| Unidentified 16 | - | - | 10.0 ± 0.0 | - | - | - | - | - | |||||||||||
| Pseudomonas putida | 18 | - | - | + | - | - | - | - | - | ||||||||||
| Pseudomonas | sp. 19 | - | 7.5 ± 0.7 | - | - | - | 8.0 ± 0.0 | - | + | ||||||||||
| Pseudomonas alcaliphila | 20 | - | - | - | - | - | - | - | - | ||||||||||
| Pseudomonas | sp. 22 | - | 7.5 ± 0.7 | 13.0 ± 0.0 | - | - | |||||||||||||
| MC | Pseudomonas | sp. | OK342262 | + | 25.6 ± 0.0 | 35.1 ± 0.2 | Less polar compound | ||||||||||||
| 5.5 ± 0.7 | - | 23 | MC | Pseudomonas | sp. | OK342263 | weak | 0.0 ± 0.0 | 36.5 ± 0.1 | nd | |||||||||
| 24 | MC | Pseudomonas khazarica | MW369469 | weak | 0.0 ± 0.0 | nd | nd | ||||||||||||
| 25 | MC | Pseudomonas | sp. | OK342264 | + | 0.0 ± 0.0 | 35.3 ± 0.1 | Less polar compounds | |||||||||||
| - | |||||||||||||||||||
| Pseudomonas | sp. 23 | - | + | + | - | - | - | - | - | ||||||||||
| Pseudomonas | sp. 25 | 9 ± 0.0 | + | - | 12.0 ± 0.0 | - | 6.5 ± 0.7 | - | - | ||||||||||
| Pseudomonas aeruginosa | 26 | - | - | 10 ± 0.0 | - | - | - | 26 | MC | Pseudomonas aeruginosa | MW369470 | ++ | 33.3 ± 0.0 | 37.6 ± 0.3 | Less polar compound | ||||
| - | - | ||||||||||||||||||
| Enterococcus ludwigii | 28 | - | 5.5 ± 0.7 | 9.0 ± 0.0 | - | + | + | - | - | ||||||||||
| Aeromonas media | 30 | - | 5.5 ± 0.7 | - | - | - | 6.0 ± 0.0 | - | - | - | 6.0 ± 0.0 | 28 | CS | Enterobacter ludwigii | MW369471 | + | 0.0 ± 0.0 | 37.9 ± 0.1 | nd |
| - | - | - | |||||||||||||||||
| Pseudomonas protegens | 37 | - | - | 10.0 ± 0.0 | - | - | - | - | - | 30 | CS | Aeromonas media | MW369472 | weak | 0.0 ± 0.0 | 39.4 ± 0.9 | nd | ||
| - | |||||||||||||||||||
| Aeromonas taiwanensis | 35 | - | |||||||||||||||||
| Aeromonas media | 40 | - | 7.5 ± 0.7 | - | 15.5 ± 0.7 | - | + | - | + | 35 | CS | Aeromonas taiwanensis | MW369473 | weak | 0.0 ± 0.0 | 43.2 ± 0.1 | nd | ||
| Pseudomonas anguilliseptica | 41 | - | + | - | - | - | - | - | 8.0 ± 0.0 | 37 | |||||||||
| Pseudomonas stutzeri | CL | 45 | Aeromonas media | -MW369474 | - | 0.0 ± 0.0 | 35.9 ± 0.1 | nd | |||||||||||
| 40 | CL | Aeromonas media | MW369476 | - | 0.0 ± 0.0 | 46.1 ± 0.3 | nd | ||||||||||||
| 41 | CL | Pseudomonas anguilliseptica | MW369477 | + | 32.1 ± 5.4 | 35.2 ± 0.6 | nd | ||||||||||||
| 5.5 ± 0.7 | - | + | - | - | - | - | - | ||||||||||||
| Pseudomonas protegens | 47 | - | 9.5 ± 0.7 | 8.0 ± 0.0 | - | - | - | - | 6.5 ± 0.7 | ||||||||||
| Pseudomonas protegens | 51 | - | 7.0 ± 0.7 | - | - | - | 7.5 ± 0.7 | - | - | 45 | CL | Pseudomonas stutzeri | OK342265 | + | 0.0 ± 0.0 | 36.3 ± 0.1 | nd | ||
| Pseudomonas protegens | 55 | 10 ± 0.0 | 9.5 ± 0.7 | - | - | - | - | - | - | 47 | CL | Pseudomonas protegens | MW369478 | weak | 0.0 ± 0.0 | 40.5 ± 0.4 | nd | ||
| Pseudomonas | sp. 56 | 13.5 ± 0.7 | 5.5 ± 0.7 | 8.0 ± 0.0 | - | - | + | - | - | 51 | CL | Pseudomonas protegens | OK342266 | + | 0.0 ± 0.0 | 35.5 ± 0.1 | nd | ||
| Negative control | 0.0 ± 0.0 | 0.0 ± 0.0 | 0.0 ± 0.0 | 0.0 ± 0.0 | 55 | CL | Pseudomonas protegens | MW369480 | - | 0.0 ± 0.0 | 37.7 ± 0.1 | nd | |||||||
| Chloramphenicol | 21 ± 0.0 | - | + | 30.0 ± 0.0 | 56 | CL | |||||||||||||
| Gentamycin CN30 | - | Pseudomonas | 14 | 8.0 ± 0.0 | 18.0 ± 0.0 | ||||||||||||||
| Amoxycillin | - | - | - | - | |||||||||||||||
 [1]
[1]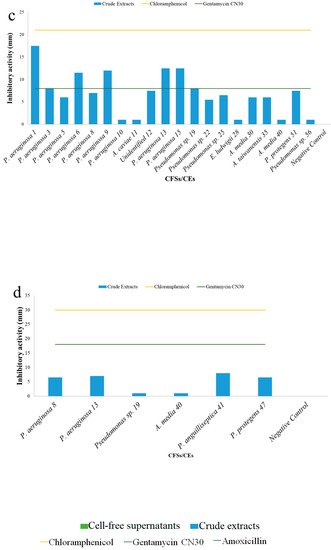
6. Conclusions
The present research let to select bacterial strains from the gut of grey mullets with interesting biotechnologically traits and has confirmed that intestinal microbiota is a promising source of new and biologically active pharmaceutical agents to control fish health and to preserve the environment. Additionally, the study of BS-producing bacteria associated with fish intestine is of relevance for our understanding of their ecological role in the symbiotic and antagonist interaction with the host and between themselves and for understanding whether the production of bioactive compounds might represent a biological strategy for protecting fish against gut and liver inflammations, an immune response and for survival with respect to the surrounding environmentThe present entry let to select bacterial strains from the gut of grey mullets with interesting biotechnologically traits and has confirmed that intestinal microbiota is a promising source of new and biologically active pharmaceutical agents to control fish health and to preserve the environment. Additionally, the study of BS-producing bacteria associated with fish intestine is of relevance for our understanding of their ecological role in the symbiotic and antagonist interaction with the host and between themselves and for understanding whether the production of bioactive compounds might represent a biological strategy for protecting fish against gut and liver inflammations, an immune response and for survival with respect to the surrounding environment
References
- Bodour, A.A.; Drees, K.P.; Maier R.M.; Distribution of biosurfactant-producing bacteria in undisturbed contaminated arid southwestern soils. . Appl. Environ. Microbiol. 2003, 69, 3280-3287, 10.1128/AEM.69.6.3280-3287.2003.
