Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is a comparison between Version 2 by Vivi Li and Version 1 by Gasim Abd-Elfarag.
Nodding syndrome (NS) is a debilitating yet often neglected neurological disease affecting thousands of children in several sub-Saharan African countries.
- nodding syndrome
- disease
- sub-Saharan Africa
1. Introduction
Nodding syndrome (NS) is a devastating but often neglected neurological condition that affects thousands of individuals in remote and resource-poor regions in several countries throughout sub-Saharan Africa, with major public health, psychosocial, and economic consequences [1,2,3,4][1][2][3][4].
2. Epidemiology
Twenty-two studies in our dataset reported on the epidemiology of NS. The first reports of NS cases came from southern Tanzania between the 1930s and the 1960s [10][5]. These early reports were followed decades later by reports from Liberia [11][6], southern Sudan (now officially known as the Republic of South Sudan) [3[3][7],12], western Uganda [13][8], and northern Uganda [14][9] (Figure 31). Recently, NS was reported in regions within the Democratic Republic of Congo (DRC) [15][10], Cameroon, and the Central African Republic [16,17][11][12] (Figure 31).
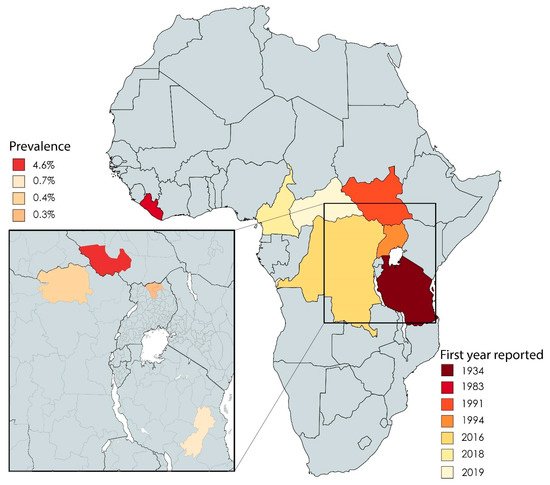

Figure 31. African countries in which cases of NS have been reported, including the first year in which they were reported. The inset shows the reported prevalence of NS in the indicated countries.
On a regional level, the overall prevalence of NS was reported to be 4.6%, 0.7%, 0.4%, and 0.3% for South Sudan (Western Equatoria) [3[3][13],18], northern Uganda (Kitgum, Pader, and Lamwu districts) [19][14], the DRC (town of Aketi) [20][15], and Tanzania (Ulang district and Morogoro region) [21][16], respectively. In Cameroon (Bilomo, Kelleng, Ngongol, Nyamongo, and Bayomen), the reported prevalence of NS among a subgroup of patients with epilepsy was 21.8% [16][11]. Finally, in the Central African Republic, a total of five cases were identified among 6175 individuals [17][12].
In villages, NS has been reported to cluster around rapidly flowing rivers infested with blackflies (Simulium spp.) [12,15,16,18,20,22,23,24,25][7][10][11][13][15][17][18][19][20] and in families, with some families having two or more affected members [3,10,19,26,27,28,29][3][5][14][21][22][23][24]. In addition, NS has also been associated with poverty, food shortage, and a history of displacement [12,18,23,25][7][13][18][20].
3. Aetiology
The aetiology of NS was investigated and reported by 18 studies in our dataset, which were further subdivided into the following seven categories (with some studies reporting more than one category): infections (n = 11), nutritional deficiencies (n = 4), toxins (n = 5), and autoimmune (n = 4), hormonal (n = 3), metabolic (n = 2), and genetic factors (n = 1). These categories are discussed in detail below.
3.1. Infection
With respect to parasitic infection, most studies focused on Onchocerca volvulus (OV) infection (Table 31) and studied the association between NS and OV infection using skin snip microscopy, serology (Ov16 IgG detection using ELISA and OvFAR/MSA detection using the luciferase immunoprecipitation system), and PCR analysis. We found that four case–control studies using skin snip microscopy measured a significantly higher prevalence of OV infection in NS cases (range: 71.1–96.7%) compared to controls (range: 43.7–53.9%) [12,18,30,31][7][13][25][26]. In addition, one case study using skin snip microscopy found a higher density of OV microfilaria in NS cases compared to patients with other forms of epilepsy [32][27]. In contrast, case studies using PCR on cerebrospinal fluid (CSF) samples did not detect either genomic material of OV in 139 cases [12,29,33,34][7][24][28][29] or species of Wolbachia (an endosymbiotic bacteria that occurs together with OV) in CSF samples taken from 10 NS cases [33][28]. In addition, Mansonella perstans infection, which was detected using microscopy on blood samples, was significantly associated with NS in one case–control study, with an odds ratio (OR) of 3.2 (p = 0.005) [12][7] (Table 31). Finally, we found no association between NS and the presence of any other parasitic infections, including Loa loa, Wuchereria bancrofti, Trypanosoma gambiense (a protozoan that causes human African trypanosomiasis, or sleeping sickness), and Taenia solium (an intestinal tapeworm that causes cysticercosis) [12,31][7][26].
Table 31. Case–control studies reporting associations between pathogens and nodding syndrome.
| Pathogen | Location | Test | Cases | Controls | Odds Ratio (95% CI) | p-Value | Reference | |||
|---|---|---|---|---|---|---|---|---|---|---|
53. Case–control studies reporting associations between toxins and nodding syndrome.
| Toxins | Location | Test | Cases | Controls | Odds Ratio (95% CI) | p-Value | Reference | |||
|---|---|---|---|---|---|---|---|---|---|---|
| Country | Area (Year) | N | % | N | % | |||||
| Country | Area (Year) | N | % | N | % | |||||
| Onchocerca volvulus | South Sudan | Lui (2001) | SSM | 39 | 89.7 | 31 | 48.3 | 9.2 (2.7–3.26) | 0.00003 | [12,18][7 |
| Mouldy maize | Uganda | Kitgum (2014) | DtH | ] | [ | 50 | 13 | ] | ||
| 50 | Amadi (2001) | SSM | 30 | 96.7 | 34 | 50 | 29 (3.5–237.7) | - | [18][13] | |
| Lui (2002) | SSM | 13 | 92.3 | 16 | 43.7 | 15.4 (1.6–148.8) | 0.008 | [12,18][7][13] | ||
| Maridi and Witto (2011) | SSM | 38 | 76.3 | 38 | 47.4 | 3.2 (1.2–8.7) | 0.02 | [30][25] | ||
| Uganda | Kitgum (2009) | SSM | 45 | 71.1 | 39 | 53.9 | 1.11 (0.37–3.27) | - | [31][26] | |
| 4.33 (1.4–18.9) | 0.009 | [ | 35 | ] | [ | 31 | ] | |||
| Maize | Uganda | Kitgum (2014) | Ov16 IgG | 39 | 66.7 | 44 | 31.8 | 3.14 (1.08–9.13) | - | |
| OvFAR/MSA | 39 | 94.9 | 41 | 48.8 | 14.4 (2.65–78.3) | |||||
| Kitgum and Pader (2016/17) | Ov16 IgG | 154 | 93.5 | 153 | 54.9 | 8.79 (4.15–18.65) | 0.001 | [37][30] | ||
| Mansonella perstans | South Sudan | Lui (2001) | BM | 39 | 41 | 31 | 9.6 | 3.2 | 0.005 | [12][7] |
| Amadi (2001) | BM | 30 | 66.6 | 34 | 50 | |||||
| Loa loa | South Sudan | Lui and Amadi (2001) | BM | 69 | 0 | 65 | 0 | - | - | [12][7] |
| Wuchereria bancrofti | South Sudan | Lui (2001) | BM | 39 | 0 | 31 | 9 | - | 0.47 | [12][7] |
| Amadi (2001) | 30 | 0 | 34 | 7.6 | - | |||||
| Trypanosoma brucei | South Sudan | Lui (2021) | CATT | 39 | 12.8 | 31 | 9.6 | 0.84 | 0.94 | [12][7] |
| Amadi (2001) | 30 | 0 | 34 | 5.8 | ||||||
| Uganda | Kitgum (2009) | CATT | 36 | 0 | 40 | 0 | - | - | [31][26] | |
| Taenia soleum | Uganda | Kitgum (2009) | Antibody | 36 | 0 | 40 | 0 | - | - | [31][26] |
| Measles virus | Uganda | Kitgum (2009) | Past history | 23.5 | 6.1 | 3.3 (0.8–13.6) | [31][26] | |||
| PCR | 16 | 0 | 0 | - | - | - | ||||
| South Sudan | Lui and Amadi (2002) | Past history | 13 | 15.38 | 19 | 58 | 0.13 | 0.025 | [12][7] | |
| Uganda | Kitgum (2014) | Past history | 50 | 100 | 50 | - | 6 (1.02–113) | 0.047 | [35][31] | |
| Hepatitis E virus | Kitgum (2009) | IgM | 38 | 31.6 | 31 | 16.1 | 1.45 (0.37–5.58) | - | [31][26] | |
| IgG | 38 | 26.3 | 30 | 33.3 | 0.81 (0.24–2.75) | - | ||||
SSM: skin snip microscopy; BM: blood microscopy; CATT: card agglutination test.
With respect to viruses, three case–control studies investigated measles infection using a previous history of measles infection as reported by the parents or legal guardians, with conflicting results (Table 31). The first study (conducted in South Sudan in 2002) reported an inverse association between prior measles infection and NS (OR: 0.13; p = 0.025) [12][7]. The second study (conducted in Uganda in 2009) found no significant association (OR: 3.3; 95% CI: 0.8–13.6) [31][26], and the third study (also conducted in Uganda, but in 2014) found a significant positive association between measles infection and NS (OR: 6; p = 0.047) [35][31]. In addition, no association was identified between hepatitis E infection and NS [31][26]. Finally, a recent case–control study found no association between NS and known viruses including Anneloviridae, Hepadnaviridae (hepatitis B), Flaviviridae, Herpesviridae, Polyomaviridae (human polyomavirus), Papillomaviridae, and Virgaviridae [36][32].
Our search revealed no studies regarding bacterial infections. Finally, there was no association between prion disease and NS (based on a reported history of eating monkey meat) [31][26].
3.2. Nutritional Deficiency
A case–control study in Uganda found that compared to controls, NS cases had lower plasma levels of vitamin B6 (OR: 7.2; p = 0.001) and higher plasma 3-hydroxykynurenine levels (OR: 4.50; p = 0.013) [38][33] (Table 42). In contrast, however, three other studies conducted in Uganda and Tanzania found no such associations [31,39,40][26][34][35].
Table 42. Case–control studies that studied nutritional deficiencies in nodding syndrome.
| Micronutrient | Location | Cases | Controls | Odds Ratio (95% CI) | p-Value | Reference | |||
|---|---|---|---|---|---|---|---|---|---|
| Country | Area (Year) | N | % | N | % | ||||
| Vitamin B6 (P5P) | Uganda | Gulu & Amuru district (2013) | 66 | 73 | 7.22 (2.24–23.26) | 0.001 | [38][33] | ||
| Uganda | Kitgum (2009) | 49 | 73 | 42 | 64 | 1.22 (0.41–3.59) | |||
. In the majority of cases, head nodding is reported to be triggered by the sight of food or by cold weather [4,14,16,18,29,30,46,57][4][9][11][13][24][25][41][52].
In over 80% of cases, NS progresses to include other seizure types, including generalised tonic–clonic seizures, partial complex seizures, and atypical absence seizures [4,12,14,16,18,21,26,29,53,56][4][7][9][11][13][16][21][24][48][51]. In addition, approximately 30% of patients develop cognitive impairment, which can present as a slow reaction time, depression, and/or dropping out of school [4,12,13,14,16,18,24,26,30,56,57,58][4][7][8][9][11][13][19][21][25][51][52][53]. Impaired physical development is also reported, including stunted growth with delayed bone age (41% of cases), wasting (73% of cases), and delayed puberty [4,13,16,24,46,56,58,59][4][8][11][19][41][51][53][54].
Further complications reported to be associated with NS include psychiatric symptoms such as mood changes, aggressiveness, sleep disturbances, and wandering in 36%, 27%, 23%, and 9% of cases, respectively, as well as catatonia symptoms (e.g., staring, mutism, stupor, and grimacing) in some cases [4,58,60][4][53][55]. In contrast, neither focal neurological abnormalities nor cranial nerve palsies have been reported in NS [4,13,14,30][4][8][9][25].
Some patients were described to progress to even more severe forms of NS, although the percentage of patients who do so is currently unknown. Children with a severe form of NS are often severely mentally retarded with impaired speech or a complete loss of speech, the inability to stand, urinary incontinence, a tendency to wander, and Parkinson-like features such as drooling, facial masking, facial tics when speaking, and slow speech patterns [4,10][4][5]. These patients may die due to uncontrolled generalised seizures or other events such as falling into the fire while cooking or drowning [4,10,12,13,14,18,29,56,58][4][5][7][8][9][13][24][51][53]. To date, no reports of children making a full recovery from NS have been published.
6. Diagnosis and Management
Currently, no laboratory test is available to confirm the clinical diagnosis of NS. However, in 2012 a consensus case definition was established during the International Scientific Meeting on Nodding Syndrome held in Kampala, Uganda [9][56], which was modified in 2013 during the single-stage cluster survey conducted by the CDC and the Ugandan Ministry of Health to determine the prevalence of NS in Uganda (Table 64) [19][14]. This definition is now used widely by clinicians and scientists. Prior to 2012, however, characteristic repetitive head-nodding seizures were used to diagnose NS.
Table 64. The 2013 modified consensus case definition for NS.
| Suspected Case: | Reported Head Nodding in a Previously Healthy Person. Head Nodding Is Defined as Repetitive, Involuntary Drops of the Head towards the Chest on Two or More Occasions | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Probable case | Suspected case of head nodding, with one major criterion plus at least one minor criterion Major criteria:
|
||||||||||||
| Confirmed case | Probable case, with documented head-nodding episodes based on:
| - | [ | 31 | ] | [ | DtH | 26] | |||||
| 50 | 50 | 4 (1.0–26.5) | 0.05 | [ | 35 | ] | [ | 31] | Uganda | - | 3 | ||
| Emergency/relief food supplies | Uganda | 100 | 5 | Kitgum (2014) | DtH | 47100 | - | - | [ | 39][ | 5034] | ||
| 4 (1.3–17.6) | 0.016 | [ | 35 | ] | [ | 31 | ] | Vitamin A | Uganda | Kitgum (2009) | 25 | ||
| Gulu and Amuru (2016) | DtH | 40 | 67 | 40 | 12 | 33 | 2.15 (0.41–11.12) | 18- | 27 | 4.05 (1.23–13.28) | 0.021 | [38][31][26] | |
| [ | 33 | ] | Vitamin B12 | Uganda | Kitgum (2009) | 25 | 8 | [ | 31][26] | ||||
| Red/brown sorghum | South Sudan | Mundri (2002) | 12 | Folate | Uganda | Kitgum (2009) | 11 | 9 | 9 | 0 | - | - | [31][26] |
| Zinc | Uganda | Kitgum (2009) | 17 | 47 | 12 | 67 | 0.72 (0.13–3.94) | - | [31][26] | ||||
| Selenium | Uganda | Kitgum (2009) | 17 | 100 | 12 | 100 | - | - | [31][26] |
P5P: pyridoxal-5-phosphate; PL: plasma level. Note: all nutrients were measured as plasma levels.
No significant association was found between NS and other micronutrient deficiencies, including vitamin A, vitamin B12, folate, zinc, and selenium [31][26].
3.3. Toxins
With respect to food-related toxins, a history of consuming maize (OR: 4.0; p = 0.05), mouldy maize (OR: 4.3; p = 0.009) [35][31], relief foods (OR: 4.0; p = 0.02) [35[31][33],38], and either red or brown sorghum (OR: 6.22; p = 0.05) [12][7] was significantly associated with NS (Table 53). One study also investigated the possible underlying aetiology associated with consuming mouldy foods by examining the presence of various mycotoxins (aflatoxin, ochratoxin, and ribotoxin deoxynivalenol) present in contaminated foods (maize, sorghum, millet, and groundnuts), but found no association [41][36].
Table
| DtH | ||||||||||
| 13 | ||||||||||
| 54 | 8 | 1.46 (0.09–22.82) | - | 19 | 16 | 6.22 (1.2–32.3) | 0.049 | [ | 12 | ][7] |
| Uganda | Kitgum (2009) | DtH | - | 98 | - | 100 | 1.3 (0.0–125.9) | - | [31][26] | |
| Spoiled relief food | Uganda | Kitgum (2009) | DtH | - | 43 | - | 47 | 0.3 (0.1–1.3) | - | [31][26] |
| Seeds meant for planting | Uganda | Kitgum (2009) | DtH | - | 61 | - | 65 | 0.6 (0.1–2.3) | - | [31][26] |
| South Sudan | Mundri (2002) | DtH | - | - | - | - | 5 (0.82–30.4) | 0.11 | [12][7] | |
| River fish | Uganda | Kitgum (2009) | DtH | - | 96 | - | 100 | 0.3 (0.0–11.6) | - | [31][26] |
| Insects | Uganda | Kitgum (2009) | DtH | - | 41 | - | 33 | 0.8 (0.2–2.9) | - | [31][26] |
| Rodent brain | Uganda | Kitgum (2009) | DtH | - | 55 | - | 51 | 1.8 (0.3–12.3) | - | [31][26] |
| Baboon brain | South Sudan | Mundri (2002) | DtH | - | - | - | - | 3 (0.63–14.2) | 0.25 | [12][7] |
| Baboon meat | South Sudan | Mundri (2002) | DtH | - | - | - | - | 4.5 (0.97–20.8) | 0.07 | [12][7] |
| Crushed roots as traditional medicines | Uganda | Kitgum (2009) | DrH | - | 39 | - | 16 | 5.4 (1.3–22.1) | - | [31][26] |
| Uganda | Kitgum (2014) | DrH | 50 | - | 50 | - | 1.29 (0.47–3.6) | 0.617 | [35][31] | |
| Crushed leaves | Uganda | Kitgum (2009) | DrH | - | 8 | - | 2 | 3.4 (0.2–45.8) | - | [31][26] |
| Crushed flowers | Uganda | Kitgum (2009) | DrH | - | 0 | - | 2 | 0.9 (0.1–5.6) | - | [31][26] |
| Inhaled medicines | Uganda | Kitgum (2009) | DrH | - | 2 | - | 0 | 0.2 (0.0–1.5) | - | [31][26] |
| Exposure to chemicals from munitions | Uganda | Kitgum (2009) | EH | - | 70 | - | 51 | 13.9 (1.4–135) | - | [31][26] |
DtH: dietary history; DrH: drug history; EH: exposure history.
Another case–control study failed to confirm the association between red/brown sorghum and NS [31][26]. Interestingly, one study reported an association—albeit not significant—between NS and a history of eating baboon meat [18][13], whereas other studies found no association between NS and the consumption of agricultural seeds, cassava, river fish, insects, rodent meat, or bush meat (including brains) [12,31][7][26]. Finally, one case–control study detected the mycotoxins α-zearalenone, aflatoxin M1, and T-2 toxin in the urine of both cases and controls, but found no significant difference between cases and controls [42][37].
Aside from food-related toxins, one study in Uganda found a positive association between NS and the use of crushed plant roots as traditional medicines (OR: 5.4; 95% CI: 1.3–22.1) [31][26] (Table 53). However, another study conducted in the same area found no such association (OR: 1.29; 95% CI: 0.47–3.6) [35][31]. Exposure to toxins from war munitions was associated with NS in Uganda (OR: 13.9; 95% CI: 1.4–135) [31][26], but not in South Sudan [12,30][7][25]. Finally, no association was found between NS and exposure to copper or mercury and the source of water for domestic use (river, borehole, spring, shallow well, or pipe) [31][26].
3.4. Autoimmunity
One case–control study used a protein array to screen for a large number of autoantibodies, revealing that autoantibodies against the protein leiomodin-1 had the strongest association with NS; specifically, leiomodin-1 autoantibodies were found in 53% of NS cases compared to 31% of controls (OR: 2.7; 95% CI: 1.1–6.5) [43][38]. However, a recent study could not confirm the association between autoantibodies against leiomodin-1 and NS [44][39].
One study found no autoantibodies against N-methyl-D-aspartic acid (NMDA) receptors or the voltage-gated potassium channel (VGKC) complex in the serum of any NS cases tested [40][35].
Finally, another case–control study found that the mean plasma levels of macrophage migration inhibitory factor (MIF) were significantly elevated (47.3 ± 25 ng/mL vs. 17.8 ± 6 ng/mL) in NS cases compared to the healthy controls, which was hypothesized to play a role in the development and disease progression as a result of autoimmunity and neuroinflammation [45][40]. However, the frequency of MIF −173 C genotypes (CC/CG) was significantly lower in NS cases compared to the healthy controls (OR 0.33; 95% CI 0.14–0.8) [45][40].
3.5. Hormonal, Metabolic, and Genetic Factors
With respect to hormonal imbalances, a case series of Ugandan adolescents reported low levels of the peptide hormone somatomedin C (also known as insulin-like growth factor-1) in two out of eight NS cases. This study also reported low levels of luteinising hormone, follicle-stimulating hormone, and the sex hormones testosterone and oestrogen in seven of eight cases [46][41]. In contrast, all eight cases had normal levels of other hormones, including thyroid hormone, parathyroid hormone, growth hormone, adrenocorticotropic hormone, adrenal corticosteroids, and mineralocorticoids [46,47][41][42]. Finally, a case–control study found no association between serum serotonin level and NS [48][43].
Altered metabolism was identified as a possible cause of NS by a case series involving 48 Ugandan patients with low mean levels of biotinidase and acetyl carnitine, but normal urate levels [49][44]. In addition, another case series involving 10 Ugandans reported high anion gap metabolic acidosis [47][42].
With respect to genetic aetiological factors, one case–control study in South Sudan (with 48 cases and 51 controls) found that the presence of specific amino acids at specific positions in the HLA-B (Ala11, Ala24, Asn63, and Phe67), DRB1 (Ala73 and Thr77), and DQB1 (Pro56, Glu66, and Val67) proteins were significantly associated with increased susceptibility to NS, while other specific amino acids in the HLA-B (Ser11 and Glu63), HLA-C (Trp156 and Glu163), DRB1 (Lys71, Gly73, Arg74, and Asn77), and DQB1 (Leu56, Asp66, Ile67, Glu70, and Asp71) proteins were associated with a decreased risk of NS [50][45]. A separate, relatively small case–control study (with three cases and five controls) of Ugandan children reported normal mitochondrial DNA and negative oligonucleotide microassay results for consanguinity, deletion, and duplication [39][34].
4. Pathophysiology
One autopsy study of five deceased NS cases in Uganda suggested that NS may represent a new form of tauopathy based on the presence of tau-immunoreactive neuronal neurofibrillary tangles, pre-tangles, neuropil threads, and dot-like tau in the cerebral cortex, brain stem, and basal ganglia of all five cases [51][46]. However, another autopsy study of another five Ugandan cases could not confirm these findings; instead, the authors proposed a neuroinflammatory pathophysiology, as they found no signs of generalised tauopathy, but rather found cerebellar atrophy, a loss of cerebellar Purkinje cells, cortical gliosis, and features indicating previous ventriculitis and/or meningitis [52][47]. This conclusion based on neuroinflammation was consistent with the findings of six separate magnetic resonance imaging (MRI) studies involving 66 cases, which found gliosis (mainly in the occipital and parieto-occipital areas) and cerebellar atrophy, with no focal changes in the cerebral cortex or hippocampus [4,14,29,34,39,53][4][9][24][29][34][48]. This neuroinflammation did not appear to be the result of a chronic infection in the central nervous system (CNS), as six studies (with a total of 148 cases) reported normal cell counts, protein levels, and glucose concentration in the CSF [4,14,29,31,33,39][4][9][24][26][28][34]. Interestingly, one of these six studies reported the presence of oligoclonal bands in the CSF—which indicates the presence of immunoglobulins that are commonly found in a wide range of neuroinflammatory diseases—in one of the three cases in their study [39][34].
No cytokine-specific profile was identified in any of the NS cases; however, one case–control study reported higher levels of the cytokine-like polypeptide C5a in the CSF of NS cases compared to controls, suggesting complement activation [54][49]. The same study also found lower plasma levels of CCL2, CCL5, CXCL13, IL-10, APRIL (also known as TNFSF13), and MMP-9 (matrix metallopeptidase 9) in NS cases compared to controls [54][49]. Another study reported decreased levels of IL-13 in NS cases, but could not detect IL-5, IL-6, or TNF-α [55][50].
Seven studies (with a total of 111 cases) reported findings based on electroencephalogram (EEG) recordings and found evidence of atonic seizures (i.e., presence of electro-decrement and paraspinal electromyographic dropout) in patients who experienced nodding episodes during the EEG recordings [14,53][9][48]. In addition, EEG recordings also showed evidence of generalised and focal seizures [4,14,18,26,29,39,53][4][9][13][21][24][34][48].
With respect to nonspecific laboratory findings, several case series reported low haemoglobin levels [4,10,47][4][5][42], elevated eosinophil counts [4[4][24],29], and increased erythrocyte sedimentation rates [4] in NS cases. In contrast, liver function, kidney function, white blood cell counts, and platelet counts were all normal [4,29,47][4][24][42].
5. Clinical Features
Prior to the onset of typical head-nodding seizures, 20% of NS cases seem to develop prodromal signs, including an expressionless stare, excessive sleepiness, dizziness, and loss of attention [4,30,56][4][25][51]. The characteristic involuntary repetitive head nodding—which may be present in all NS cases—likely results from a periodic loss of neck muscle tone [13,14,18,30][8][9][13][25]. The onset of these symptoms has been described in both children and young adults ranging from 2 to 22 years of age, with a slightly higher prevalence among males (55%) [4,10,13,14,16,19,25,26,31][4][5][8][9][11][14][20][21][26]
| Video/EEG/EMG documenting head nodding as atonic seizures |
Seven studies in our dataset reported on the management of NS. Five of these studies found that the use of anticonvulsants—including phenobarbitone, carbamazepine, sodium valproate, and phenytoin (either individually or in combination)—reduced seizure frequency in 70% of cases, with complete seizure control achieved in 25% of cases [13,29,53,59,61][8][24][48][54][57]. In addition, anticonvulsants were reported to increase basic self-care, behaviour, and school attendance in 80%, 59%, and 40% of cases, respectively, thus reflecting an increase in independence [61][57]. In one study, the combined use of anticonvulsants and multivitamins was reported to reduce wasting and stunting [59][54]; however, whether these interventions affect disease progression in NS remains unknown. Finally, community-directed treatment with ivermectin combined with larviciding of rivers was reported to coincide with a reduction in the incidence of NS in northern Uganda [62][58].
7. Community Perceptions and Psychosocial and Economic Impact
Twelve qualitative studies (all of which were performed in Uganda) investigated community perceptions and the psychosocial and economic burden of NS.
7.1. Perceptions and Beliefs
Within communities, NS was perceived to be associated with a wide variety of factors, including living in a camp for internally displaced people [63,64,65][59][60][61]; consumption of expired and/or contaminated relief foods [64,65,66][60][61][62]; exposure to chemicals from war munitions [63,64,65,67,68][59][60][61][63][64]; evil spirits, being cursed, or punishment from the gods as a result of bad deeds committed during a time of war [58,63,64,65,66,67,69][53][59][60][61][62][63][65]; open-gold mining in Karamoja region (northeastern Uganda) polluting the rivers with heavy metals such as mercury [65][61]; and blackflies breeding in the rivers in northern Uganda [63,67][59][63].
7.2. Transmission, Presentation, and Treatment
In some studies, communities reported the belief that NS can be transmitted from person to person through saliva and the air [58,63,64,66,68][53][59][60][62][64]; therefore, patients with NS are often isolated from their family members and peers (e.g., eating and sleeping separately) in NS-affected communities. Some communities reportedly perceive NS as a distinct disease entity from epilepsy, given that NS is generally more severe and can include severe mental and/or physical disabilities [66,68][62][64]. Furthermore, local communities in northern Uganda reported believing that NS cannot be cured [58,63,69][53][59][65] and that anticonvulsive medications—particularly sodium valproate—are associated with promiscuous behaviour, resulting in low treatment compliance [70][66].
7.3. Health-Seeking Behaviour of Caretakers
Caretakers generally seek healthcare from government health facilities; however, a shortage of healthcare workers, long waiting hours, frequent lack of medicines, and transport-related issues (e.g., a lack of money for transportation and/or difficulties associated with transporting mentally and physically disabled patients with NS) often drives caretakers to seek care from traditional healers and witch doctors [70,71][66][67].
7.4. Psychosocial and Economic Burden
The mental and physical impairments caused by NS often lead to personal and family shame [70[66][68],72], discrimination, social isolation [65[61][62][63][68],66,67,72], and economic constraints due to reduced livelihood activities [66,67,68,72][62][63][64][68]. In addition, caretakers experience poor sleep and are often depressed, which can lead to domestic violence, substance abuse, and suicidal and homicidal thoughts, potentially ending in family separation [66,67,68,72][62][63][64][68]. Some caretakers report feeling that their children with NS are useless [65,69][61][65]. In addition, children often drop out of school due to fears related to the person-to-person transmission of NS [66,67][62][63]. Finally, some individuals with NS report being sexually abused, exploited, or forced to engage in child labour [70][66].
References
- Dowell, S.F.; Sejvar, J.J.; Riek, L.; Vandemaele, K.A.; Lamunu, M.; Kuesel, A.C.; Schmutzhard, E.; Matuja, W.; Bunga, S.; Foltz, J.; et al. Nodding syndrome. Emerg. Infect. Dis. 2013, 19, 1374–1384.
- Lacey, M. Nodding disease: Mystery of southern Sudan. Lancet Neurol. 2003, 2, 714.
- Nodding Syndrome. Available online: http://www.who.int/onchocerciasis/symptoms/nodding_syndrome/en/ (accessed on 27 September 2019).
- Idro, R.; Opoka, R.O.; Aanyu, H.T.; Kakooza-Mwesige, A.; Piloya-Were, T.; Namusoke, H.; Musoke, S.B.; Nalugya, J.; Bangirana, P.; Mwaka, A.D.; et al. Nodding syndrome in Ugandan children—Clinical features, brain imaging and complications: A case series. BMJ Open 2013, 3, 3.
- Spencer, P.S.; Palmer, V.S.; Jilek-Aall, L. Nodding syndrome: Origins and natural history of a longstanding epileptic disorder in sub-Saharan Africa. Afr. Health Sci. 2013, 13, 176–182.
- Waals, F.W.V.D.; Goudsmit, J.; Gajdusek, D.C. Characteristics of Highly Prevalent Seizure Disorders in the Gbawein and Wroughbarh Clan Region of Grand Bassa County, Liberia. Neuroepidemiology 1983, 1983, 35–44.
- Spencer, P.S.; Vandemaele, K.; Richer, M.; Palmer, V.S.; Chungong, S.; Anker, M.; Ayana, Y.; Opoka, M.L.; Klaucke, B.N.; Quarello, A.; et al. Nodding syndrome in Mundri county, South Sudan: Environmental, nutritional and infectious factors. Afr. Health Sci. 2013, 13, 183–204.
- Kaiser, C.; Rubaale, T.; Tukesiga, E.; Kipp, W.; Asaba, G. Nodding syndrome, western Uganda, 1994. Am. J. Trop. Med. Hyg. 2015, 93, 198–202.
- Sejvar, J.J.; Kakooza, A.M.; Foltz, J.L.; Makumbi, I.; Atai-Omoruto, A.D.; Malimbo, M.; Ndyomugyenyi, R.; Alexander, L.N.; Abang, B.; Downing, R.G.; et al. Clinical, neurological, and electrophysiological features of nodding syndrome in Kitgum, Uganda: An observational case series. Lancet Neurol. 2013, 12, 166–174.
- Lenaerts, E.; Mandro, M.; Mukendi, D.; Suykerbuyk, P.; Dolo, H.; Wonya’Rossi, D.; Ngave, F.; Ensoy-Musoro, C.; Laudisoit, A.; Hotterbeekx, A.; et al. High prevalence of epilepsy in onchocerciasis endemic health areas in Democratic Republic of the Congo. Infect. Dis. Poverty 2018, 7, 68.
- Siewe, J.F.N.; Ngarka, L.; Tatah, G.; Mengnjo, M.K.; Nfor, L.N.; Chokote, E.S.; Boullé, C.; Nkouonlack, C.; Dema, F.; Nkoro, G.A.; et al. Clinical presentations of onchocerciasis-associated epilepsy (OAE) in Cameroon. Epilepsy Behav. 2019, 90, 70–78.
- Metanmo, S.; Boumediene, F.; Preux, P.M.; Colebunders, R.; Siewe Fodjo, J.N.; de Smet, E.; Yangatimbi, E.; Winkler, A.S.; Mbelesso, P.; Ajzenberg, D. First description of Nodding Syndrome in the Central African Republic. PLoS Negl. Trop. Dis. 2021, 15, e0009430.
- Tumwine, J.K.; Vandemaele, K.; Chungong, S.; Richer, M.; Anker, M.; Ayana, Y.; Opoka, M.L.; Klaucke, D.N.; Quarello, A.; Spencer, P.S. Clinical and epidemiologic characteristics of nodding syndrome in Mundri County, southern Sudan. Afr. Health Sci. 2012, 12, 242–248.
- Iyengar, P.J.; Wamala, J.; Ratto, J.; Blanton, C.; Malimbo, M.; Lukwago, L.; Becknell, S.; Downing, R.; Bunga, S.; Sejvar, J.; et al. Prevalence of nodding syndrome—Uganda, 2012–2013. CDC MMWR—Morb. Mortal. Wkly. Rep. 2014, 63, 603–606.
- Mukendi, D.; Tepage, F.; Akonda, I.; Siewe, J.N.F.; Rotsaert, A.; Ndibmun, C.N.; Laudisoit, A.; Couvreur, S.; Kabutako, B.; Menon, S.; et al. High prevalence of epilepsy in an onchocerciasis endemic health zone in the Democratic Republic of the Congo, despite 14 years of community-directed treatment with ivermectin: A mixed-method assessment. Int. J. Infect. Dis. 2019, 79, 187–194.
- Mmbando, B.P.; Suykerbuyk, P.; Mnacho, M.; Kakorozya, A.; Matuja, W.; Hendy, A.; Greter, H.; Makunde, W.H.; Colebunders, R. High prevalence of epilepsy in two rural onchocerciasis endemic villages in the Mahenge area, Tanzania, after 20 years of community directed treatment with ivermectin. Infect. Dis. Poverty 2018, 7, 64.
- Colebunders, R.; Carter, J.Y.; Olore, P.C.; Puok, K.; Bhattacharyya, S.; Menon, S.; Abd-Elfarag, G.; Ojok, M.; Ensoy-Musoro, C.; Lako, R.; et al. High prevalence of onchocerciasis-associated epilepsy in villages in Maridi County, Republic of South Sudan: A community-based survey. Seizure 2018, 63, 93–101.
- Kitara, D.L.; Oh, J.; Mwaka, A.D. Nodding Syndrome in Uganda—A Disease Cluster: An Epidemiological Dilemma. Pac. J. Med Sci. 2013, 11, 21–33.
- Kaiser, C.; Asaba, G.; Rubaale, T.; Tukesiga, E.; Kipp, W. Onchocerciasis-Associated Epilepsy with Head Nodding Seizures-Nodding Syndrome: A Case Series of 15 Patients from Western Uganda, 1994. Am. J. Trop. Med. Hyg. 2018, 99, 1211–1218.
- Lakwo, T.L.; Raimon, S.; Tionga, M.; Siewe Fodjo, J.N.; Alinda, P.; Sebit, W.J.; Carter, J.Y.; Colebunders, R. The Role of the Maridi Dam in Causing an Onchocerciasis-Associated Epilepsy Epidemic in Maridi, South Sudan: An Epidemiological, Sociological, and Entomological Study. Pathogens 2020, 9, 315.
- De Polo, G.; Romaniello, R.; Otim, A.; Benjamin, K.; Bonanni, P.; Borgatti, R. Neurophysiological and clinical findings on Nodding Syndrome in 21 South Sudanese children and a review of the literature. Seizure 2015, 31, 64–71.
- Colebunders, R.; Hendy, A.; Mokili, J.L.; Wamala, J.F.; Kaducu, J.; Kur, L.; Tepage, F.; Mandro, M.; Mucinya, G.; Mambandu, G.; et al. Nodding syndrome and epilepsy in onchocerciasis endemic regions: Comparing preliminary observations from South Sudan and the Democratic Republic of the Congo with data from Uganda. BMC Res. Notes 2016, 9, 182.
- Idro, R. Nodding syndrome in Uganda: Prevalence, clinical characteristics and management. Trop. Med. Int. Health 2013, 18, 29.
- Winkler, A.S.; Friedrich, K.; Konig, R.; Meindl, M.; Helbok, R.; Unterberger, I.; Gotwald, T.; Dharsee, J.; Velicheti, S.; Kidunda, A.; et al. The head nodding syndrome--clinical classification and possible causes. Epilepsia 2008, 49, 2008–2015.
- CDC. Nodding syndrome—South Sudan, 2011. MMWR Morb. Mortal. Wkly. Rep. 2012, 61, 52–54.
- Foltz, J.L.; Makumbi, I.; Sejvar, J.J.; Malimbo, M.; Ndyomugyenyi, R.; Atai-Omoruto, A.D.; Alexander, L.N.; Abang, B.; Melstrom, P.; Kakooza, A.M.; et al. An Epidemiologic Investigation of Potential Risk Factors for Nodding Syndrome in Kitgum District, Uganda. PLoS ONE 2013, 8, e66419.
- Abd-Elfarag, G.; Carter, J.Y.; Raimon, S.; Sebit, W.; Suliman, A.; Fodjo, J.N.S.; Olore, P.C.; Biel, K.P.; Ojok, M.; Logora, M.Y.; et al. Persons with onchocerciasis-associated epilepsy and nodding seizures have a more severe form of epilepsy with more cognitive impairment and higher levels of Onchocerca volvulus infection. Epileptic Disord. 2020, 22, 301–308.
- Hotterbeekx, A.; Raimon, S.; Abd-Elfarag, G.; Carter, J.Y.; Sebit, W.; Suliman, A.; Siewe Fodjo, J.N.; De Witte, P.; Logora, M.Y.; Colebunders, R.; et al. Onchocerca volvulus is not detected in the cerebrospinal fluid of persons with onchocerciasis-associated epilepsy. Int. J. Infect. Dis. 2020, 91, 119–123.
- Winkler, A.S.; Friedrich, K.; Velicheti, S.; Dharsee, J.; Konig, R.; Nassri, A.; Meindl, M.; Kidunda, A.; Muller, T.H.; Jilek-Aall, L.; et al. MRI findings in people with epilepsy and nodding syndrome in an area endemic for onchocerciasis: An observational study. Afr. Health Sci. 2013, 13, 529–540.
- Gumisiriza, N.; Kugler, M.; Brusselaers, N.; Mubiru, F.; Anguzu, R.; Ningwa, A.; Ogwang, R.; Akun, P.; Mwaka, A.D.; Abbo, C.; et al. Risk Factors for Nodding Syndrome and Other Forms of Epilepsy in Northern Uganda: A Case-Control Study. Pathogens 2021, 10, 1451.
- Spencer, P.S.; Mazumder, R.; Palmer, V.S.; Lasarev, M.R.; Stadnik, R.C.; King, P.; Kabahenda, M.; Kitara, D.L.; Stadler, D.; McArdle, B.; et al. Environmental, dietary and case-control study of Nodding Syndrome in Uganda: A post-measles brain disorder triggered by malnutrition? J. Neurol. Sci. 2016, 369, 191–203.
- Roach, M.; Cantu, A.; Vieri, M.K.; Cotten, M.; Kellam, P.; Phan, M.; Hoek, L.V.; Mandro, M.; Tepage, F.; Mambandu, G.; et al. No Evidence Known Viruses Play a Role in the Pathogenesis of Onchocerciasis-Associated Epilepsy. An Explorative Metagenomic Case-Control Study. Pathogens 2021, 10, 787.
- Obol, J.H.; Arony, D.A.; Wanyama, R.; Luryama Moi, K.; Bodo, B.; Odong, P.O.; Odida, M. Reduced plasma concentrations of vitamin B6 and increased plasma concentrations of the neurotoxin 3-hydroxykynurenine are associated with nodding syndrome: A case control study in Gulu and Amuru Districts, Northern Uganda. Trans. R. Soc. Trop. Med. Hyg. 2019, 113, S102–S103.
- Soldatos, A.; Nutman, T.; Groden, C.; Wahl, C.; Inati, S.; Buckler, G.; O’Connell, E.; Opar, B.; Aryek-Kwe, J.; Downing, R.; et al. Evaluation and immunomodulatory treatment at the NIH of children with nodding syndrome from northern Uganda. Neurology 2015, 84, S37.005.
- Dietmann, A.; Wallner, B.; Konig, R.; Friedrich, K.; Pfausler, B.; Deisenhammer, F.; Griesmacher, A.; Seger, C.; Matuja, W.; JilekAall, L.; et al. Nodding syndrome in Tanzania may not be associated with circulating anti-NMDA-and anti-VGKC receptor antibodies or decreased pyridoxal phosphate serum levels-a pilot study. Afr. Health Sci. 2014, 14, 434–438.
- Echodu, R.; Edema, H.; Malinga, G.M.; Hendy, A.; Colebunders, R.; Moriku Kaducu, J.; Ovuga, E.; Haesaert, G. Is nodding syndrome in northern Uganda linked to consumption of mycotoxin contaminated food grains? BMC Res. Notes 2018, 11, 678.
- Duringer, J.; Mazumder, R.; Palmer, V.; Craig, M.; Spencer, P. Case-Control Study of Nodding Syndrome in Acholiland: Urinary Multi-Mycotoxin Screening. Toxins 2021, 13, 313.
- Johnson, T.P.; Tyagi, R.; Lee, P.R.; Lee, M.H.; Johnson, K.R.; Kowalak, J.; Elkahloun, A.; Medynets, M.; Hategan, A.; Kubofcik, J.; et al. Nodding syndrome may be an autoimmune reaction to the parasitic worm Onchocerca volvulus. Sci. Transl. Med. 2017, 9, 15.
- Hotterbeekx, A.; Vieri, M.K.; Ramberger, M.; Jozefzoon-Aghai, A.; Mandro, M.; Tepage, F.; Dusabimana, A.; Kumar-Singh, S.; Titulaer, M.J.; Colebunders, R. No Evidence for the Involvement of Leiomodin-1 Antibodies in the Pathogenesis of Onchocerciasis-Associated Epilepsy. Pathogens 2021, 10, 845.
- Benedek, G.; Abed El Latif, M.; Miller, K.; Rivkin, M.; Ahmed Ramadhan Lasu, A.; Riek, L.P.; Lako, R.; Edvardson, S.; Arbel-Alon, S.; Galun, E.; et al. Macrophage migration inhibitory factor in Nodding syndrome. PLoS Negl. Trop. Dis. 2021, 15, e0009821.
- Piloya-Were, T.; Odongkara-Mpora, B.; Namusoke, H.; Idro, R. Physical growth, puberty and hormones in adolescents with Nodding Syndrome; a pilot study. BMC Res. Notes 2014, 7, 858.
- Kitara, D.L.; Mwaka, A.D.; Kigonya, E. High Anion Gap Metabolic Acidosis among Children with Nodding Syndrome (NS) in Northern Uganda: Case Series. Br. J. Med. Med. Res. 2014, 4, 1301–1314.
- Vieri, M.K.; Hotterbeekx, A.; Mandro, M.; Siewe Fodjo, J.N.; Dusabimana, A.; Nyisi, F.; Mukendi, D.; Gwatsvaira, J.; Kumar-Singh, S.; Colebunders, R. Serotonin Levels in the Serum of Persons with Onchocerciasis-Associated Epilepsy: A Case-Control Study. Pathogens 2021, 10, 720.
- Denis, A.A.; Galloway, P.; Collines, A.; Frederick, M.E.; David, D.P.K.L. Metabolic analysis of nodding syndrome in Uganda: A pilot study is a biotinidase and acetyl carnitine deficiency; a metabolic disorder. An observational study design. World J. Pharm. Med. Res. 2018, 4, 160–174.
- Benedek, G.; Abed El Latif, M.; Miller, K.; Rivkin, M.; Ramadhan Lasu, A.A.; Riek, L.P.; Lako, R.; Edvardson, S.; Alon, S.A.; Galun, E.; et al. Protection or susceptibility to devastating childhood epilepsy: Nodding Syndrome associates with immunogenetic fingerprints in the HLA binding groove. PLoS Negl. Trop. Dis. 2020, 14, e0008436.
- Pollanen, M.S.; Onzivua, S.; Robertson, J.; McKeever, P.M.; Olawa, F.; Kitara, D.L.; Fong, A. Nodding syndrome in Uganda is a tauopathy. Acta Neuropathol. 2018, 136, 691–697.
- Hotterbeekx, A.; Lammens, M.; Idro, R.; Akun, P.R.; Lukande, R.; Akena, G.; Nath, A.; Taylor, J.; Olwa, F.; Kumar-Singh, S.; et al. Neuroinflammation and Not Tauopathy Is a Predominant Pathological Signature of Nodding Syndrome. J. Neuropathol. Exp. Neurol. 2019, 78, 1049–1058.
- Winkler, A.S.; Wallner, B.; Friedrich, K.; Pfausler, B.; Unterberger, I.; Matuja, W.; Jilek-Aall, L.; Schmutzhard, E. A longitudinal study on nodding syndrome--a new African epilepsy disorder. Epilepsia 2014, 55, 86–93.
- Ogwang, R.; Muhanguzi, D.; Mwikali, K.; Anguzu, R.; Kubofcik, J.; Nutman, T.B.; Taylor, M.; Newton, C.R.; Vincent, A.; Conroy, A.L.; et al. Systemic and cerebrospinal fluid immune and complement activation in Ugandan children and adolescents with long-standing nodding syndrome: A case-control study. Epilepsia Open 2021, 6, 297–309.
- Vieri, M.K.; Hotterbeekx, A.; Raimon, S.; Abd-Elfarag, G.; Mukendi, D.; Carter, J.Y.; Kumar-Singh, S.; Colebunders, R. Cytokines and Onchocerciasis-Associated Epilepsy, a Pilot Study and Review of the Literature. Pathogens 2021, 10, 310.
- Idro, R.; Ogwang, R.; Kayongo, E.; Gumisiriza, N.; Lanyero, A.; Kakooza-Mwesige, A.; Opar, B. The natural history of nodding syndrome. Epileptic Disord. 2018, 20, 508–516.
- Colebunders, R.; Abd-Elfarag, G.; Carter, J.Y.; Olore, P.C.; Puok, K.; Menon, S.; Fodjo Siewe, J.N.; Bhattacharyya, S.; Ojok, M.; Lako, R.; et al. Clinical characteristics of onchocerciasis-associated epilepsy in villages in Maridi County, Republic of South Sudan. Seizure 2018, 62, 108–115.
- Musisi, S.; Akena, D.; Nakimuli-Mpungu, E.; Abbo, C.; Okello, J. Neuropsychiatric perspectives on nodding syndrome in northern Uganda: A case series study and a review of the literature. Afr. Health Sci. 2013, 13, 205–218.
- Gazda, S.; Kitara, D.L. Treatment and rehabilitation outcomes of children affected with nodding syndrome in Northern Uganda: A descriptive case series. Pan. Afr. Med. J. 2018, 29, 228.
- Kakooza-Mwesige, A.; Dhossche, D.M.; Idro, R.; Akena, D.; Nalugya, J.; Opar, B.T. Catatonia in Ugandan children with nodding syndrome and effects of treatment with lorazepam: A pilot study. BMC Res. Notes 2015, 8, 825.
- WHO. International Scientific Meeting on Nodding Syndrome Kampala, Uganda; WHO: Geneva, Switzerland, 2012; pp. 1–42.
- Idro, R.; Namusoke, H.; Abbo, C.; Mutamba, B.B.; Kakooza-Mwesige, A.; Opoka, R.O.; Musubire, A.K.; Mwaka, A.D.; Opar, B.T. Patients with nodding syndrome in Uganda improve with symptomatic treatment: A cross-sectional study. BMJ Open 2014, 4, e006476.
- Gumisiriza, N.; Mubiru, F.; Siewe Fodjo, J.N.; Mbonye Kayitale, M.; Hotterbeekx, A.; Idro, R.; Makumbi, I.; Lakwo, T.; Opar, B.; Kaducu, J.; et al. Prevalence and incidence of nodding syndrome and other forms of epilepsy in onchocerciasis-endemic areas in northern Uganda after the implementation of onchocerciasis control measures. Infect. Dis. Poverty 2020, 9, 12.
- Van Bemmel, K.; Derluyn, I.; Stroeken, K. Nodding syndrome or disease? On the conceptualization of an illness-in-the-making. Ethn. Health 2014, 19, 100–118.
- Mitchell, K.B.; Kornfeld, J.; Adiama, J.; Mugenyi, A.; Schmutzhard, E.; Ovuga, E.; Kamstra, J.; Winkler, A.S. Nodding syndrome in northern Uganda: Overview and community perspectives. Epilepsy Behav. 2013, 26, 22–24.
- Kitara, D.L.; Amone, C. Perception of the population in Northern Uganda to nodding syndrome. J. Med. Med. Sci. 2012, 3, 464–470.
- Buchmann, K. ‘These nodding people’: Experiences of having a child with nodding syndrome in postconflict Northern Uganda. Epilepsy Behav. 2015, 42, 71–77.
- Buchmann, K. ‘You sit in fear’: Understanding perceptions of nodding syndrome in post-conflict northern Uganda. Glob. Health Action 2014, 7, 25069.
- Irani, J.; Rujumba, J.; Mwaka, A.D.; Arach, J.; Lanyuru, D.; Idro, R.; Gerrets, R.; Grietens, K.P.; O’Neill, S. “Those who died are the ones that are cured”. Walking the political tightrope of Nodding Syndrome in northern Uganda: Emerging challenges for research and policy. PLoS Negl. Trop. Dis. 2019, 13, e0007344.
- Mutamba, B.; Abbo, C.; Muron, J.; Idro, R.; Mwaka, A.D. Stereotypes on Nodding syndrome: Responses of health workers in the affected region of northern Uganda. Afr. Health Sci. 2013, 13, 986–991.
- Abbo, C.; Mwaka, A.D.; Opar, B.T.; Idro, R. Qualitative evaluation of the outcomes of care and treatment for children and adolescents with nodding syndrome and other epilepsies in Uganda. Infect. Dis. Poverty 2019, 8, 30.
- Mwaka, A.D.; Okello, E.S.; Abbo, C.; Odwong, F.O.; Olango, W.; Etolu, J.W.; Oriyabuzu, R.; Lagoro, D.K.; Mutamba, B.B.; Idro, R.; et al. Is the glass half full or half empty? A qualitative exploration on treatment practices and perceived barriers to biomedical care for patients with nodding syndrome in post-conflict northern Uganda. BMC Res. Notes 2015, 8, 386.
- Nakigudde, J.; Mutamba, B.B.; Bazeyo, W.; Musisi, S.; James, O. An exploration of caregiver burden for children with nodding syndrome (lucluc) in Northern Uganda. BMC Psychiatry 2016, 16, 255.
More
