One of the major health concerns for human society is skin cancer. When the pigments producing skin color turn carcinogenic, this disease gets contracted. A skin cancer diagnosis is a challenging process for dermatologists as many skin cancer pigments may appear similar in appearance. Hence, early detection of lesions (which form the base of skin cancer) is definitely critical and useful to completely cure the patients suffering from skin cancer. Significant progress has been made in developing automated tools for the diagnosis of skin cancer to assist dermatologists. The worldwide acceptance of artificial intelligence-supported tools has permitted usage of the enormous collection of images of lesions and benevolent sores approved by histopathology.
- image classification
- skin lesion
- CNN
- transfer learning
- artificial intelligence
1. Introduction
Literature Background
-
To classify the images from HAM10000 dataset into seven different types of skin cancer.
-
To use transfer learning nets for feature selection and classification so as to identify all types of lesions found in skin cancer.
-
To properly balance the dataset using replication on only training data and perform a detailed analysis using different transfer learning models.
2. Methods
2.1. Dataset Description for Skin Lesion
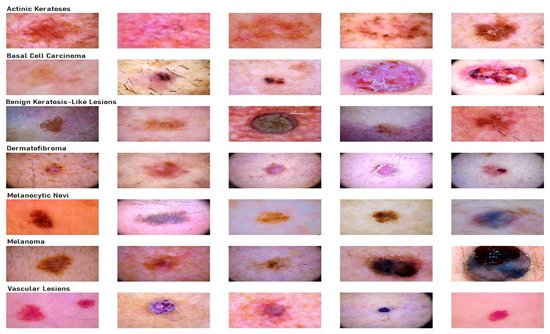
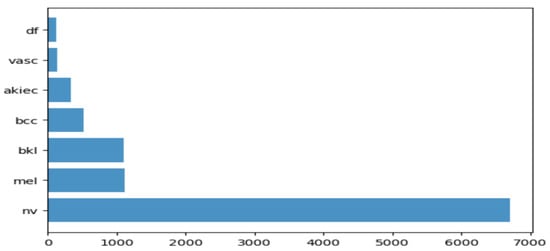
2.2. Transfer Learning Nets
2.2.1. VGG19
2.2.2. InceptionV3
2.2.3. InceptionResnetv2
2.2.4. ResNet50
2.2.5. Xception
2.2.6. MobileNet
2.3. Proposed Methodology
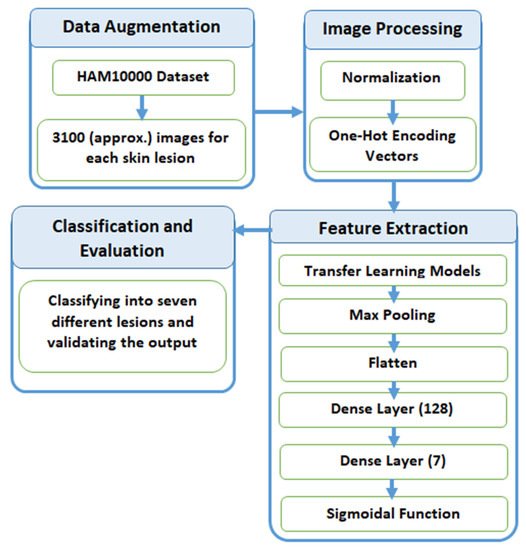
2.3.1. Data Augmentation
| Disease | Frequency before Augmentation | Multiply Factor (k) | Frequency after Augmentation |
|---|---|---|---|
| Melanocytic Nevi | 3179 | 1 | 3179 |
| Benign Keratosis | 317 | 10 | 3170 |
| Melanoma | 165 | 19 | 3135 |
| Basal Cell Carcinoma | 126 | 25 | 3150 |
| Actinic Keratosis | 109 | 29 | 3161 |
| Vascular Skin Lesions | 46 | 69 | 3174 |
| Dermatofibroma | 28 | 110 | 3080 |
2.3.2. Preprocessing
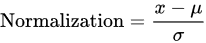
2.3.3. Feature Extraction
2.3.4. Classification and Evaluation
References
- Kricker, A.; Armstrong, B.K.; English, D.R. Sun exposure and non-melanocytic skin cancer. Cancer Causes Control 1994, 5, 367–392.
- Armstrong, B.K.; Kricker, A. The epidemiology of UV induced skin cancer. J. Photochem. Photobiol. B 2001, 63, 8–18.
- American Cancer Society. Cancer Facts and Figures. 2020. Available online: https://www.cancer.org/content/dam/cancer-org/research/cancer-facts-and-statistics/annual-cancer-facts-and-figures/2020/cancer-facts-and-figures-2020.pdf (accessed on 5 July 2021).
- Silberstein, L.; Anastasi, J. Hematology: Basic Principles and Practice; Elsevier: Amsterdam, The Netherlands, 2017; p. 2408.
- Kadampur, M.A.; Al Riyaee, S. Skin cancer detection: Applying a deep learning based model driven architecture in the cloud for classifying dermal cell images. Inform. Med. Unlocked 2020, 18, 100282.
- Haenssle, H.A.; Fink, C.; Schneiderbauer, R.; Toberer, F.; Buhl, T.; Blum, A.; Kalloo, A.; Hassen, A.B.H.; Thomas, L.; Enk, A.; et al. Man against machine: Diagnostic performance of a deep learning convolutional neural network for dermoscopic melanoma recognition in comparison to 58 dermatologists. Ann. Oncol. 2018, 29, 1836–1842.
- Bhattacharyya, S.; Snasel, V.; Hassanien, A.E.; Saha, S.; Tripathy, B.K. Deep Learning: Research and Applications; De Gruyter: Berlin, Germany; Boston, MA, USA, 2020.
- Dhanamjayulu, C.; Nizhal, N.U.; Maddikunta, P.K.R.; Gadekallu, T.R.; Iwendi, C.; Wei, C.; Xin, Q. Identification of malnutrition and prediction of BMI from facial images using real-time image processing and machine learning. IET Image Process. 2021.
- Adate, A.; Arya, D.; Shaha, A.; Tripathy, B.K. Deep Learning: Research and Applications. In 4 Impact of Deep Neural Learning on Artificial Intelligence Research; Siddhartha, B., Vaclav, S., Aboul Ella, H., Satadal, S., Tripathy, B.K., Eds.; De Gruyter: Berlin, Germany, 2020; pp. 69–84.
- Adate, A.; Tripathy, B.K. Deep Learning Techniques for Image Processing Machine Learning for Big Data Analysis; Bhattacharyya, S., Bhaumik, H., Mukherjee, A., De, S., Eds.; De Gruyter: Berlin, Germany, 2018; pp. 69–90.
- Ayoub, A.; Mahboob, K.; Javed, A.R.; Rizwan, M.; Gadekallu, T.R.; Alkahtani, M.; Abidi, M.H. Classification and Categorization of COVID-19 Outbreak in Pakistan. Comput. Mater. Contin. 2021, 69, 1253–1269.
- Abidi, M.H.; Alkhalefah, H.; Mohammed, M.K.; Umer, U.; Qudeiri, J.E.A. Optimal Scheduling of Flexible Manufacturing System Using Improved Lion-Based Hybrid Machine Learning Approach. IEEE Access 2020, 8, 96088–96114.
- Abidi, M.H.; Umer, U.; Mohammed, M.K.; Aboudaif, M.K.; Alkhalefah, H. Automated Maintenance Data Classification Using Recurrent Neural Network: Enhancement by Spotted Hyena-Based Whale Optimization. Mathematics 2020, 8, 2008.
- Ch, R.; Gadekallu, T.R.; Abidi, M.H.; Al-Ahmari, A. Computational System to Classify Cyber Crime Offenses using Machine Learning. Sustainability 2020, 12, 4087.
- Abidi, M.H.; Alkhalefah, H.; Umer, U. Fuzzy harmony search based optimal control strategy for wireless cyber physical system with industry 4.0. J. Intell. Manuf. 2021.
- Kumar, P.A.; Shankar, G.S.; Maddikunta, P.K.R.; Gadekallu, T.R.; Al-Ahmari, A.; Abidi, M.H. Location Based Business Recommendation Using Spatial Demand. Sustainability 2020, 12, 4124.
- Marks, R. The epidemiology of non-melanoma skin cancer: Who, why and what can we do about it. J. Dermatol. 1995, 22, 853–857.
- Farooq, A.; Jia, X.; Hu, J.; Zhou, J. Transferable Convolutional Neural Network for Weed Mapping With Multisensor Imagery. IEEE Trans. Geosci. Remote Sens. 2021, 1–16.
- Farooq, A.; Jia, X.; Hu, J.; Zhou, J. Knowledge Transfer via Convolution Neural Networks for Multi-Resolution Lawn Weed Classification. In Proceedings of the 2019 10th Workshop on Hyperspectral Imaging and Signal Processing: Evolution in Remote Sensing (WHISPERS), Amsterdam, The Netherlands, 24–26 September 2019; pp. 1–5.
- Bose, A.; Tripathy, B.K. Deep Learning for Audio Signal Classification. In Deep Learning: Research and Applications; Bhattacharyya, S., Snasel, V., Ella Hassanien, A., Saha, S., Tripathy, B.K., Eds.; De Gruyter: Berlin, Germany, 2020; pp. 105–136.
- Ghayvat, H.; Pandya, S.N.; Bhattacharya, P.; Zuhair, M.; Rashid, M.; Hakak, S.; Dev, K. CP-BDHCA: Blockchain-based Confidentiality-Privacy preserving Big Data scheme for healthcare clouds and applications. IEEE J. Biomed. Health Inform. 2021, 1.
- Shah, A.; Ahirrao, S.; Pandya, S.; Kotecha, K.; Rathod, S. Smart Cardiac Framework for an Early Detection of Cardiac Arrest Condition and Risk. Front. Public Health 2021, 9, 762303.
- Ghayvat, H.; Awais, M.; Gope, P.; Pandya, S.; Majumdar, S. ReCognizing SUspect and PredictiNg ThE SpRead of Contagion Based on Mobile Phone LoCation DaTa (COUNTERACT): A system of identifying COVID-19 infectious and hazardous sites, detecting disease outbreaks based on the internet of things, edge computing, and artificial intelligence. Sustain. Cities Soc. 2021, 69, 102798.
- Kaul, D.; Raju, H.; Tripathy, B.K. Deep Learning in Healthcare. In Deep Learning in Data Analytics: Recent Techniques, Practices and Applications; Acharjya, D.P., Mitra, A., Zaman, N., Eds.; Springer International Publishing: Cham, Switzerland, 2022; pp. 97–115.
- Nugroho, A.A.; Slamet, I.; Sugiyanto. Skins cancer identification system of HAMl0000 skin cancer dataset using convolutional neural network. AIP Conf. Proc. 2019, 2202, 020039.
- Voulodimos, A.; Doulamis, N.; Doulamis, A.; Protopapadakis, E. Deep Learning for Computer Vision: A Brief Review. Comput. Intell. Neurosci. 2018, 2018, 7068349.
- Guo, Y.; Liu, Y.; Oerlemans, A.; Lao, S.; Wu, S.; Lew, M.S. Deep learning for visual understanding: A review. Neurocomputing 2016, 187, 27–48.
- Garcia-Garcia, A.; Orts-Escolano, S.; Oprea, S.; Villena-Martinez, V.; Garcia-Rodriguez, J. A Review on Deep Learning Techniques Applied to Semantic Segmentation. arXiv 2017, arXiv:1704.06857. Available online: https://ui.adsabs.harvard.edu/abs/2017arXiv170406857G (accessed on 1 April 2017).
- Maheshwari, K.; Shaha, A.; Arya, D.; Rajasekaran, R.; Tripathy, B.K. Deep Learning: Research and Applications. In Convolutional Neural Networks: A Bottom-Up Approach; Siddhartha, B., Vaclav, S., Aboul Ella, H., Satadal, S., Tripathy, B.K., Eds.; De Gruyter: Berlin, Germany, 2020; pp. 21–50.
- Mateen, M.; Wen, J.; Nasrullah; Song, S.; Huang, Z. Fundus Image Classification Using VGG-19 Architecture with PCA and SVD. Symmetry 2019, 11, 1.
- Canziani, A.; Paszke, A.; Culurciello, E. An Analysis of Deep Neural Network Models for Practical Applications. arXiv 2016, arXiv:1605.07678.
- Codella, N.C.F.; Gutman, D.; Celebi, M.E.; Helba, B.; Marchetti, M.A.; Dusza, S.W.; Kalloo, A.; Liopyris, K.; Mishra, N.; Kittler, H.; et al. Skin lesion analysis toward melanoma detection: A challenge at the 2017 International symposium on biomedical imaging (ISBI), hosted by the international skin imaging collaboration (ISIC). In Proceedings of the 2018 IEEE 15th International Symposium on Biomedical Imaging (ISBI 2018), Washington, DC, USA, 4–7 April 2018; pp. 168–172.
- Li, Y.; Shen, L. Skin Lesion Analysis towards Melanoma Detection Using Deep Learning Network. Sensors 2018, 18, 556.
- Han, S.S.; Kim, M.S.; Lim, W.; Park, G.H.; Park, I.; Chang, S.E. Classification of the Clinical Images for Benign and Malignant Cutaneous Tumors Using a Deep Learning Algorithm. J. Investig. Dermatol. 2018, 138, 1529–1538.
- Chaturvedi, S.S.; Gupta, K.; Prasad, P.S. Skin Lesion Analyser: An Efficient Seven-Way Multi-class Skin Cancer Classification Using MobileNet. Adv. Mach. Learn. Technol. Appl. 2020, 1141, 165–176.
- Milton, M.A.A. Automated Skin Lesion Classification Using Ensemble of Deep Neural Networks in ISIC 2018: Skin Lesion Analysis Towards Melanoma Detection Challenge. arXiv 2019, arXiv:1901.10802.
- Esteva, A.; Kuprel, B.; Novoa, R.A.; Ko, J.; Swetter, S.M.; Blau, H.M.; Thrun, S. Dermatologist-level classification of skin cancer with deep neural networks. Nature 2017, 542, 115–118.
- Goyal, M.; Oakley, A.; Bansal, P.; Dancey, D.; Yap, M.H. Skin Lesion Segmentation in Dermoscopic Images With Ensemble Deep Learning Methods. IEEE Access 2020, 8, 4171–4181.
- Tschandl, P.; Rosendahl, C.; Kittler, H. The HAM10000 dataset, a large collection of multi-source dermatoscopic images of common pigmented skin lesions. Sci. Data 2018, 5, 180161.
- Adate, A.; Tripathy, B.K. A Survey on Deep Learning Methodologies of Recent Applications. In Deep Learning in Data Analytics: Recent Techniques, Practices and Applications; Acharjya, D.P., Mitra, A., Zaman, N., Eds.; Springer International Publishing: Cham, Switzerland, 2022; pp. 145–170.
- Simonyan, K.; Zisserman, A. Very Deep Convolutional Networks for Large-Scale Image Recognition. arXiv 2014, arXiv:1409.1556.
- Szegedy, C.; Vanhoucke, V.; Ioffe, S.; Shlens, J.; Wojna, Z. Rethinking the Inception Architecture for Computer Vision. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, 27–30 June 2016; pp. 2818–2826.
- Szegedy, C.; Wei, L.; Yangqing, J.; Sermanet, P.; Reed, S.; Anguelov, D.; Erhan, D.; Vanhoucke, V.; Rabinovich, A. Going deeper with convolutions. In Proceedings of the 2015 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Boston, MA, USA, 7–12 June 2015; pp. 1–9.
- Szegedy, C.; Ioffe, S.; Vanhoucke, V.; Alemi, A.A. Inception-v4, inception-ResNet and the impact of residual connections on learning. In Proceedings of the Thirty-First AAAI Conference on Artificial Intelligence, San Francisco, CA, USA, 4–9 February 2017; pp. 4278–4284.
- Debgupta, R.; Chaudhuri, B.B.; Tripathy, B.K. A Wide ResNet-Based Approach for Age and Gender Estimation in Face Images. In Proceedings of the International Conference on Innovative Computing and Communications, Singapore, 24 October 2020; pp. 517–530.
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. In Proceedings of the 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778.
- Chollet, F. Xception: Deep Learning with Depthwise Separable Convolutions. In Proceedings of the 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 21–26 July 2017; pp. 1800–1807.
- Howard, A.G.; Zhu, M.; Chen, B.; Kalenichenko, D.; Wang, W.; Weyand, T.; Andreetto, M.; Adam, H. Mobilenets: Efficient convolutional neural networks for mobile vision applications. arXiv 2017, arXiv:1704.04861.
