Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is a comparison between Version 1 by Nan Chen and Version 2 by Beatrix Zheng.
The German cockroach, Blattella germanica, and the American cockroach, Periplaneta americana are the most common and synanthropic household pests of interest to public health. While they have increasingly served as model systems in hemimetabolous insects for studying many biological issues, there is still a lack of stable reference gene evaluation for reliable quantitative real-time PCR (qPCR) outputs and functional genomics.
- cockroaches
- reference genes
- qPCR normalization
- gene expression
- functional genomics
1. Introduction
The Blattaria cockroaches have evolved as an ancient and highly successful form of insect life. Some species (less than 1%) in this group serve as public health pests, of which the German cockroach, Blattella germanica, and the American cockroach, Periplaneta americana, are the most common and troublesome household pests worldwide [1]. These two species are strictly synanthropic and usually infest human-built structures, including homes, apartments, restaurants, hospitals, and other places where food is available. They harbor and mechanically transmit various pathogens and trigger asthma and allergic diseases [2][3]. During the last decades, an increasing number of studies on B. germanica and P. americana have shown them to be valuable organisms for exploring a variety of biological issues. In particular, they serve as model systems for studies of developmental biology and endocrinology in hemimetabolous insects [4][5][6][7][8][9][10] and nutrition and reproduction physiology [11][12][13][14]. As excellent chemical communicators, they have long served as important models for studying chemical ecology, especially in the aspects of sex and aggregation pheromones [15][16][17][18][19][20]. These omnivorous cockroach species are also wildly used for examining host-gut microbiota interactions with regard to their development and behavior [21][22][23][24].
Quantitative real-time polymerase chain reaction (qPCR) is a powerful molecular tool that allows the detection and measurement of messenger RNA (mRNA) at the transcriptional level. Being a faster and more sensitive method over the traditional northern blotting and semi-quantitative PCR, qPCR has developed as the most widely used approach for gene expression profiling and validation of transcriptome data [25][26][27][28]. The accuracy and reliability of qPCR outputs strongly depend on many biological and technical factors, such as sample quality, RNA integrity, cDNA synthesis efficiency, and laboratory procedures involved. Therefore, normalization of the data with appropriate reference genes, also known as housekeeping genes, is needed for minimizing variability [29]. Ideal reference genes are assumed to have constant and stable expression across biotic and abiotic factors. However, it is hard and almost impossible to use universal reference genes under all conditions (e.g., developmental stages, tissues, and experimental treatments). Evaluation and identification of appropriate reference genes prior to qPCR analyses is hence crucial for normalization. Importantly, this is also an indispensable step of the MIQE guideline that currently serves as the golden criteria of qPCR experiments [29].
With the advent of next-generation sequencing technology, many research fields in entomology have been profoundly developed into the Genomic Era. In 2018, the genomes of both B. germanica and P. americana were published [9][30]. Depending on the genome availability, functional genomic studies could provide an in-depth understanding of cockroaches and novel insights into old issues at the molecular level and on a genome-wide scale. Assessment of gene function by silencing gene expression (e.g., highly efficient RNAi in cockroaches) and accurate measurement of gene expression are needed for successful functional genomics. Hitherto, only actin has been used historically and extensively as a reference gene for qPCR normalization in these two species [6][7][8][9][10][11][12][13][14][28][31][32][33]. However, its stability under specific experimental conditions was not empirically validated, yet there is no stable reference gene quantification system for B. germanica and P. americana.
2. Current Insights
Many qPCR studies have reached a consensus that it is unrealistic to find a ‘universal’ reference gene showing constant expression across all species and experimental conditions [25]. Identification of appropriate reference genes under different conditions (spatiotemporal and experimental treatments) is therefore mandatory for reliable qPCR analysis in a given species [29]. Hitherto, a stable reference gene system has been established in a variety of insect orders but not the Blattaria cockroaches. In the present study, we evaluated the stability of several reference genes in B. germanica and P. americana, which are important model systems in hemimetabolous insects. We sampled several tissues and developmental stages that have their own advantages against others on studying biological issues of interest. For example, the antennae should be preferred for exploring chemosensory mechanisms, as with legs for limb regeneration, and the last nymphal instar for metamorphosis. We found that the obtained Ct values from either different tissues, stages, or experimental treatments showed a much higher variation in P. americana than those in B. germanica (Figure 1, Figure 2 and Figure 3). A possible explanation is that the mixed specimens at MN (N9–11) and LN (N12–14) might introduce higher sample variations since P. americana harbors a much longer molting cycle at each instar. Previous studies have demonstrated significant impacts of tissue types and developmental stages on reference gene expression, in some cases, even greater than the experimental treatments [34][35][36]. Based on the comprehensive RefFinder ranking that integrates the results of four individual algorithms, RPL32 showed the lowest variation in most tissues or at most developmental stages in B. germanica. By contrast, actin was most frequently preferred in P. americana tissue types, whereas EF1α performed well at most stages (Figure 2 and Figure 3). It is of note that actin was most stably expressed across various developmental stages in the P. americana gut but was the least stable gene in the ovary (Figure 2M,N). Nevertheless, these varied data highlight the importance of screening condition-specific reference genes in a given species.
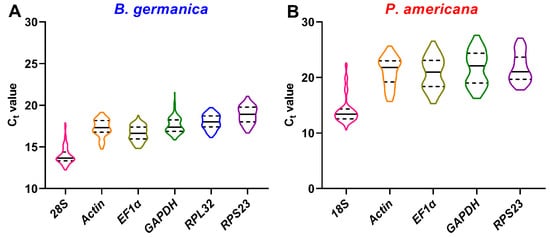
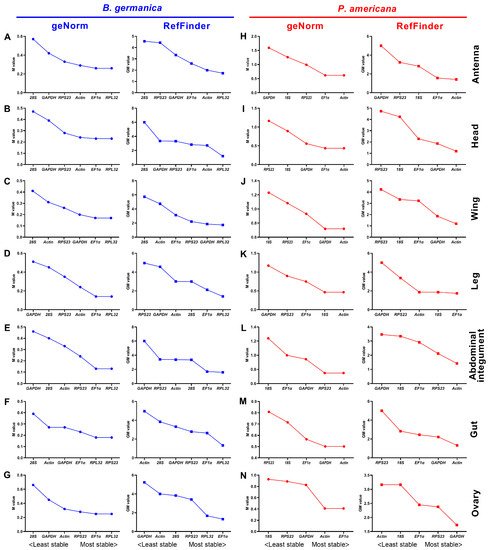

Figure 1. Ct value distribution of candidate reference genes in all examined samples from B. germanica (A) or P. americana (B). The median and quartiles are indicated by dashed and solid lines.

Figure 2. Expression stability rankings of candidate reference genes across different developmental stages in specific tissue types. The M and GM values were calculated by the geNorm and RefFinder algorithms, respectively, for candidate reference genes across different stages in the tissue of antenna, head, wing, abdominal integument, gut, and ovary in either B. germanica (A–G) or P. americana (H–N).
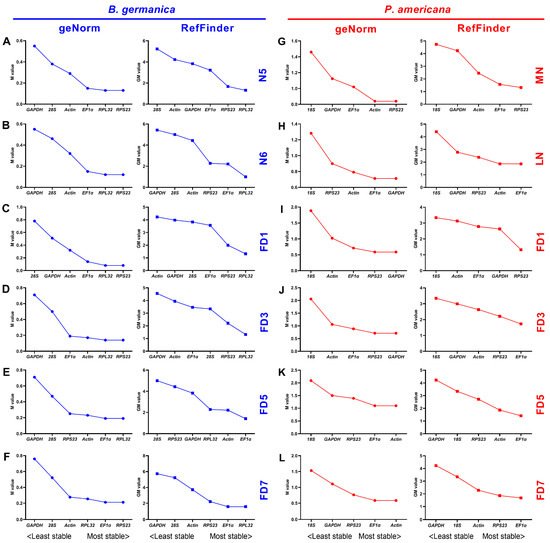

Figure 3. Expression stability rankings of candidate reference genes across various tissues at given developmental stages. The M and GM values were calculated by the geNorm and RefFinder algorithms, respectively, for candidate reference genes across various tissue types at the developmental stages of nymphs N5 (A) and N6 (B), and adults FD1 (C), FD3 (D), FD5 (E), and FD7 (F) in B. germanica. In P. americana, gene expression stability was calculated at the developmental stages of nymphs MN (G) and LN (H), and also adults FD1 (I), FD3 (J), FD5 (K), and FD7 (L).
While the actin gene has been extensively and empirically used in both B. germanica and P. americana, regardless of tissue types, developmental stages, and experimental treatments, its stability under specific conditions has never been evaluated. Our data showed that actin was rarely selected as the most stable reference gene by all four algorithms (only preferred by NormFinder under JH treatment) (Figure S4A) in B. germanica. However, it showed the highest stability in several tissue types, but not at various examined stages in P. americana (Figure 2 and Figure S2). Therefore, we conclude that actin was not appropriate for gene expression analyses, at least in B. germanica, for all the examined tissue types and developmental stages, nor under hormone treatment and RNAi conditions.
Overall, the present study has identified appropriate reference genes across different tissue types, developmental stages, and experimental treatments, including hormone application and dsRNA injection in B. germanica, and inhibitor and antibiotic feeding in P. americana. In B. germanica, we recommend RPL32 as an appropriate internal control for most spatiotemporal conditions, and the combination of RPL32 and EF1α might be ideal for all the tested tissue types and developmental stages. In addition, GAPDH and EF1α were recommended for the quantification of gene expression under JH treatment and RNAi conditions, respectively. In P. americana, actin and EF1α were appropriate in most tissue types and developmental stages, respectively, while no efficient reference gene combination was sufficient for spatiotemporal normalization. RPS23 and 18S was the best choice under inhibitor and antibiotic treatment, respectively. Clearly, more investigations are needed for qPCR analysis under other experimental conditions not tested at this time. This study is the first step toward facilitating functional genomics and an in-depth understanding of cockroaches from aspects of interest at the molecular level.
References
- Bell, W.J.; Roth, L.M.; Nalepa, C.A. Cockroaches: Ecology, Behavior, and Natural History; The Johns Hopkins University Press: Baltimore, MD, USA, 2007; pp. 1–6.
- Gore, J.C.; Schal, C. Cockroach Allergen Biology and Mitigation in the Indoor Environment. Annu. Rev. Entomol. 2007, 52, 439–463.
- Ramirez, P.J. The Cockroach as a Vector of Pathogenic Agents. Bol. Ofcina. Sanit. Panam. 1989, 107, 41–53.
- Treiblmayr, K.; Pascual, N.; Piulachs, M.-D.; Keller, T.; Belles, X. Juvenile Hormone Titer versus Juvenile Hormone Synthesis in Female Nymphs and Adults of the German Cockroach, Blattella germanica. J. Insect Sci. 2006, 6, 43.
- Mané-Padrós, D.; Cruz, J.; Vilaplana, L.; Pascual, N.; Belles, X.; Martín, D. The Nuclear Hormone Receptor BgE75 Links Molting and Developmental Progression in the Direct-Developing Insect Blattella germanica. Dev. Biol. 2008, 315, 147–160.
- Gomez-Orte, E.; Belles, X. MicroRNA-Dependent Metamorphosis in Hemimetabolan Insects. Proc. Natl. Acad. Sci. USA 2009, 106, 21678–21682.
- Lozano, J.; Belles, X. Conserved Repressive Function of Krüppel Homolog 1 on Insect Metamorphosis in Hemimetabolous and Holometabolous Species. Sci. Rep. 2011, 1, 163.
- Belles, X.; Santos, C. The MEKRE93 (Methoprene Tolerant-Krüppel Homolog 1-E93) Pathway in the Regulation of Insect Metamorphosis, and the Homology of the Pupal Stage. Insect Biochem. Mol. Biol. 2014, 52, 60–68.
- Li, S.; Zhu, S.; Jia, Q.; Yuan, D.; Ren, C.; Li, K.; Liu, S.; Cui, Y.; Zhao, H.; Cao, Y.; et al. The Genomic and Functional Landscapes of Developmental Plasticity in the American Cockroach. Nat. Commun. 2018, 9, 1008.
- Wexler, J.; Delaney, E.K.; Belles, X.; Shcal, C.; Wada-Katsumata, A.; Amicucci, K.; Kopp, A. Hemimetabolous Insects Elucidate the Origin of Sexual Development via Alternative Splicing. eLife 2019, 8, e47490.
- Maestro, J.; Cobo, J.; Belles, X. Target of Rapamycin (TOR) Mediates the Transduction of Nutritional Signals into Juvenile Hormone Production. J. Biol. Chem. 2009, 284, 5506–5513.
- Süren-Castillo, S.; Abrisqueta, M.; Maestro, J. FoxO Inhibits Juvenile Hormone Biosynthesis and Vitellogenin Production in the German Cockroach. Insect Biochem. Mol. Biol. 2012, 42, 491–498.
- Abrisqueta, M.; Süren-Castillo, S.; Maestro, J. Insulin Receptor-Mediated Nutritional Signalling Regulates Juvenile Hormone Biosynthesis and Vitellogenin Production in the German Cockroach. Insect Biochem. Mol. Biol. 2014, 49, 14–23.
- Zhu, S.; Liu, F.; Zeng, H.; Li, N.; Ren, C.; Su, Y.; Zhou, S.; Wang, G.; Palli, S.R.; Wang, J.; et al. Insulin/IGF Signaling and TORC1 Promote Vitellogenesis via Inducing Juvenile Hormone Biosynthesis in the American Cockroach. Development 2020, 147, dev188805.
- Okada, K.; Mori, M.; Shimazaki, K.; Chuman, T. Behavioral Responses of Male Periplaneta americana L. to Female Sex Pheromone Components, Periplanone-A and Periplanone-B. J. Chem. Ecol. 1990, 16, 2605–2614.
- Schal, C.; Burns, E.; Gadot, M.; Chase, J.; Blomquist, G.J. Biochemistry and Regulation of Pheromone Production in Blattella germanica (L.) (Dictyoptera, Blattellidae). Insect Biochem. 1991, 21, 73–79.
- Chase, J.; Touhara, K.; Prestwich, G.D.; Schal, C.; Blomquist, G.J. Biosynthesis and Endocrine Control of the Production of the German Cockroach Sex Pheromone 3,11-Dimethylnonacosan-2-One. Proc. Natl. Acad. Sci. USA 1992, 89, 6050–6054.
- Nojima, S.; Schal, C.; Webster, F.X.; Santangelo, R.G.; Roelofs, W.L. Identification of the Sex Pheromone of the German Cockroach, Blattella germanica. Science 2005, 307, 1104–1106.
- Eliyahu, D.; Nojima, S.; Mori, K.; Schal, C. New Contact Sex Pheromone Components of the German Cockroach, Blattella Germanica, Predicted from the Proposed Biosynthetic Pathway. J. Chem. Ecol. 2008, 34, 229–237.
- Wada-Katsumata, A.; Schal, C. Antennal Grooming Facilitates Courtship Performance in a Group-Living Insect, the German Cockroach Blattella germanica. Sci. Rep. 2019, 9, 2942.
- Tinker, K.A.; Ottesen, E.A. The Core Gut Microbiome of the American Cockroach, Periplaneta americana, is Stable and Resilient to Dietary Shifts. Appl. Environ. Microbiol. 2016, 82, 6603–6610.
- Jahnes, B.C.; Herrmann, M.; Sabree, Z.L. Conspecific Coprophagy Stimulates Normal Development in a Germ-Free Model Invertebrate. PeerJ 2019, 7, e6914.
- Guzman, J.; Vilcinskas, A. Bacteria Associated with Cockroaches: Health Risk or Biotechnological Opportunity? Appl. Microbiol. Biotechnol. 2020, 104, 10369–10387.
- Vera-Ponce de Leon, A.; Jahnes, B.C.; Otero-Bravo, A.; Sabree, Z.L. Microbiota Perturbation or Elimination Can Inhibit Normal Development and Elicit a Starvation-like Response in an Omnivorous Model Invertebrate. mSystems 2021, 6, e00802–e00821.
- Bustin, S.A. Quantification of mRNA Using Real-Time Reverse Transcription PCR (RT-PCR): Trends and Problems. J. Mol. Endocrinol. 2002, 29, 23–39.
- Zhou, X.; Qian, K.; Tong, Y.; Zhu, J.; Qiu, X.; Zeng, X. De Novo Transcriptome of the Hemimetabolous German Cockroach (Blattella germanica). PLoS ONE 2014, 9, e106932.
- Niu, D.-J.; Liu, Y.; Dong, X.-T.; Dong, S.-L. Transcriptome Based Identification and Tissue Expression Profiles of Chemosensory Genes in Blattella germanica (Blattaria: Blattidae). Comp. Biochem. Physiol. D 2016, 18, 30–43.
- Chen, N.; Pei, X.; Li, S.; Fan, Y.-L.; Liu, T.-X. Involvement of integument-rich CYP4G19 in hydrocarbon biosynthesis and cuticular penetration resistance in Blattella germanica (L.). Pest. Manag. Sci. 2019, 76, 215–226.
- Bustin, S.A.; Benes, V.; Garson, J.A.; Hellemans, J.; Huggett, J.; Kubista, M.; Mueller, R.; Nolan, T.; Pfaffl, M.W.; Shipley, G.L.; et al. The MIQE Guidelines: Minimum Information for Publication of Quantitative Real-Time PCR Experiments. Clin. Chem. 2009, 55, 611–622.
- Harrison, M.C.; Jongepier, E.; Robertson, H.M.; Arning, M.; Bitard-Feildel, T.; Chao, H.; Childers, C.P.; Dinh, H.; Doddapaneni, H.; Dugan, S.; et al. Hemimetabolous genomes reveal molecular basis of termite eusociality. Nat. Ecol. Evol. 2018, 2, 557–566.
- Clynen, E.; Ciudad, L.; Bellés, X.; Piulachs, M.-D. Conservation of fruitless’ role as master regulator of male courtship behaviour from cockroaches to flies. Dev. Genes Evol. 2011, 221, 43–48.
- Ylla, G.; Piulachs, M.-D.; Belles, X. Comparative Transcriptomics in Two Extreme Neopterans Reveal General Trends in the Evolution of Modern Insects. Iscience 2018, 4, 164–179.
- Pei, X.-J.; Fan, Y.-L.; Bai, Y.; Bai, T.-T.; Schal, C.; Zhang, Z.-F.; Chen, N.; Li, S.; Liu, T.-X. Modulation of Fatty Acid Elongation in Cockroaches Sustains Sexually Dimorphic Hydrocarbons and Female Attractiveness. PLoS Biol. 2021, 19, e3001330.
- Jacobsen, A.V.; Yemaneab, B.T.; Jass, J.; Scherbak, N. Reference Gene Selection for qPCR Is Dependent on Cell Type Rather Than Treatment in Colonic and Vaginal Human Epithelial Cell Lines. PLoS ONE 2014, 9, e115592.
- Rocha-Martins, M.; Njaine, B.; Silveira, M.S. Avoiding Pitfalls of Internal Controls: Validation of Reference Genes for Analysis by qRT-PCR and Western Blot throughout Rat Retinal Development. PLoS ONE 2012, 7, e43028.
- Uddin, M.J.; Cinar, M.U.; Tesfaye, D.; Looft, C.; Tholen, E.; Schellander, K. Age-Related Changes in Relative Expression Stability of Commonly Used Housekeeping Genes in Selected porciNE Tissues. BMC Res. Notes 2011, 4, 441.
More
