MiRNAs are ~22-nucleotide long noncoding sequences of RNA that are located across the genome, within an intron or untranslated region (UTR) of a coding gene. Pri-miRNAs are transcribed from their genes in longer primary transcripts which are processed by two RNase III proteins—Drosha and Dicer—to form a functional miRISC complex that binds to the 3′ UTR of target mRNAs and induces their degradation and translational repression . miRNAs were found to be highly stable in blood and other body fluids, where they circulate in a cell-free form, bound to other proteins, lipids, or lipoprotein or encapsulated in exosomes. The development of specific high-throughput detection methods allowing miRNA detection in extracellular fluids, besides the fact that profiles of miRNAs were shown to be either downregulated or overexpressed across several cancer types compared to normal counterparts, has paved the way for serum miRNAs to be developed as biomarkers for early detection and monitoring of tumor evolution. However, significant challenges remain, such as the low concentration of miRNAs released in the blood, especially in early-stage disease, and the difficult identification of biomarker release sites.
- miRNAs
- biomarkers
- prostate cancer
- ultrasounds
1. Ultrasound Treatment Increases the Release of Known Biomarkers in PCa Cell Lines
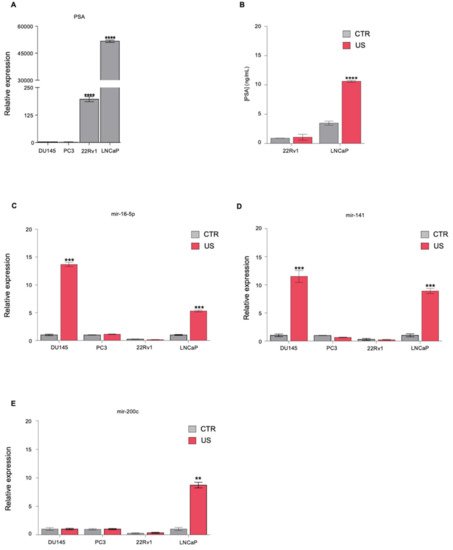
2. Identification of Novel miRNAs in the Supernatant of PCa Cells following US Treatment
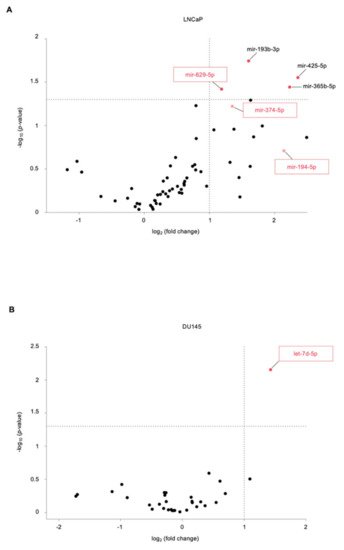
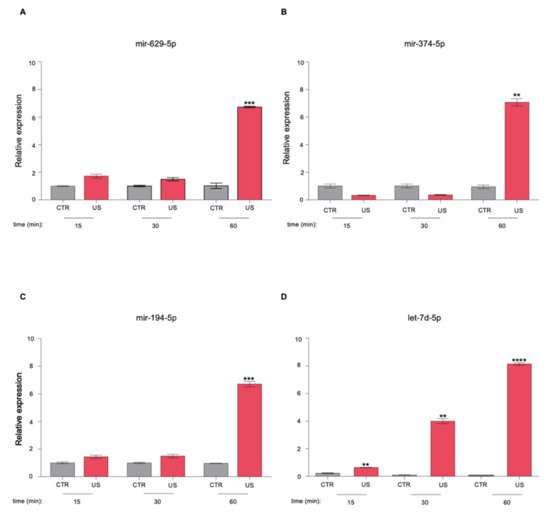
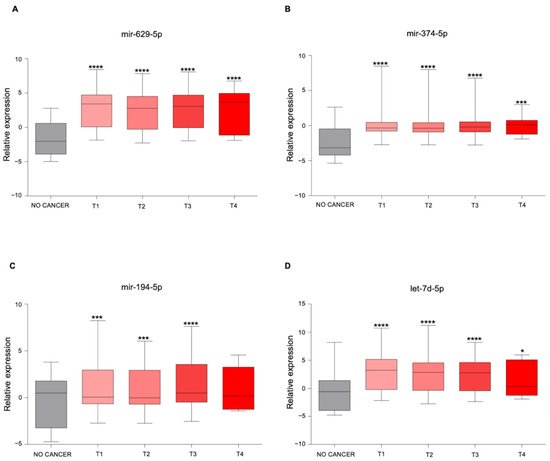
3. The Newly Identified miRNAs Are Upregulated in the Serum from PCa Patients
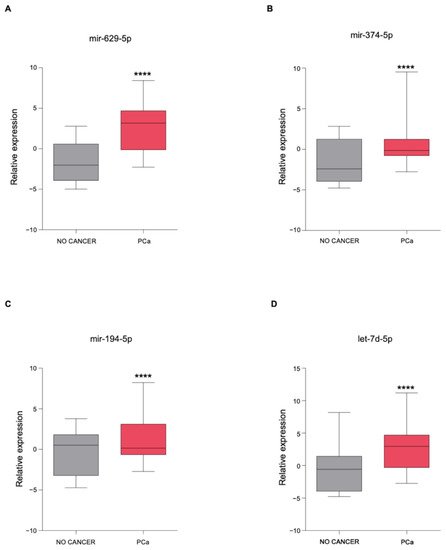
4. In Silico miRNA: Gene Interaction Analysis
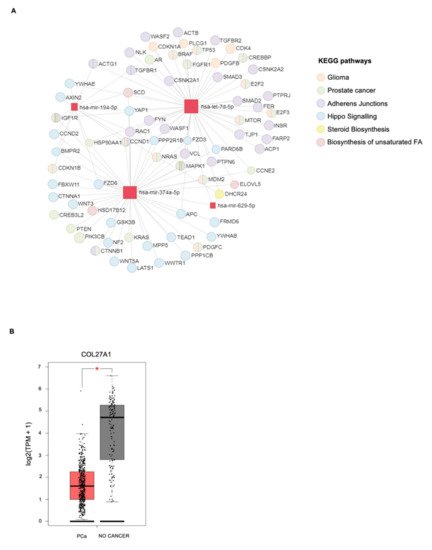
| miRNAs | KEGG Pathways | p-Value | Target Genes |
|---|---|---|---|
| miR-629-5p | Prolactin signaling pathway | 4.27 × 10−3 | PRLR AKT3 |
| miR-194-5p | Valine, leucine, and isoleucine degradation |
6.99 × 10−4 | BCKDHA ACADSB BCAT1 |
| let-7d-5p | ECM–receptor interaction | 1.07 × 10−2 | COL27A1 COL3A1 COL1A1 COL1A2 |
