At the cell-to-cell level, bacteria and fungi interact at many different levels of intimacy that can be considered from two perspectives: in terms of physical associations and in terms of molecular communication. More often than not, multiple mechanism of interaction can be employed by one microorganism. Cumulative cell-to-cell interactions among organisms belonging to very different taxa levels and origins, such as animals, protists, fungi, bacteria, archaea, and viruses, determine the overall microbial community activity in a given habitat. These interactions have an effect not just on their surrounding environment (i.e., microenvironment), but also influence large-scale fluxes and, thus, impact global ecosystem processes.
- soil microbiology
- microenvironment
- heterogeneity
- methods
- fungi
- bacteria
1. Fungi, Bacteria and Their Microenvironment
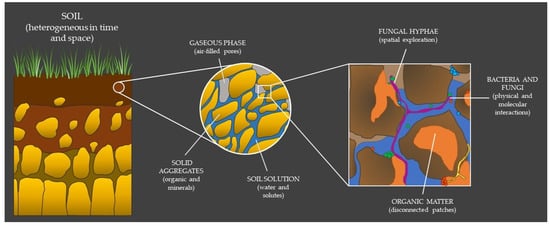
2. The Hidden Properties of Soil
3. Bacterial–Fungal Interactions: A Harsh Existence
References
- Young, I.M. Interactions and self-organization in the soil-microbe complex. Science 2004, 304, 1634–1637.
- Dexter, A. Advances in characterization of soil structure. Soil Tillage Res. 1988, 11, 199–238.
- Hillel, D. Introduction to Environmental Soil Physics; Elsevier: Amsterdam, The Netherlands, 2003.
- Young, I.; Crawford, J.; Rappoldt, C. New methods and models for characterising structural heterogeneity of soil. Soil Tillage Res. 2001, 61, 33–45.
- Baveye, P.C.; Otten, W.; Kravchenko, A.; Balseiro-Romero, M.; Beckers, É.; Chalhoub, M.; Darnault, C.; Eickhorst, T.; Garnier, P.; Hapca, S.; et al. Emergent properties of microbial activity in heterogeneous soil microenvironments: Different research approaches are slowly converging, yet major challenges remain. Front. Microbiol. 2018, 9, 1929.
- Tisdall, J.M.; Oades, J.M. Organic matter and water-stable aggregates in soils. Eur. J. Soil Sci. 1982, 33, 141–163.
- Horn, R.; Smucker, A. Structure formation and its consequences for gas and water transport in unsaturated arable and forest soils. Soil Tillage Res. 2005, 82, 5–14.
- Mueller, C.W.; Koelbl, A.; Hoeschen, C.; Hillion, F.; Heister, K.; Herrmann, A.M.; Kögel-Knabner, I. Submicron scale imaging of soil organic matter dynamics using NanoSIMS—From single particles to intact aggregates. Org. Geochem. 2012, 42, 1476–1488.
- Rawlins, B.G.; Wragg, J.; Reinhard, C.; Atwood, R.C.; Houston, A.; Lark, R.M.; Rudolph, S. Three-dimensional soil organic matter distribution, accessibility and microbial respiration in macroaggregates using osmium staining and synchrotron X-ray computed tomography. SOIL 2016, 2, 659–671.
- Jasinska, E.; Wetzel, H.; Baumgartl, T.; Horn, R. Heterogeneity of physico-chemical properties in structured soils and its consequences. Pedosphere 2006, 16, 284–296.
- Schurgers, G.; Dörsch, P.; Bakken, L.; Leffelaar, P.; Haugen, L. Modelling soil anaerobiosis from water retention characteristics and soil respiration. Soil Biol. Biochem. 2006, 38, 2637–2644.
- Ranjard, L.; Richaume, A. Quantitative and qualitative microscale distribution of bacteria in soil. Res. Microbiol. 2001, 152, 707–716.
- Otten, W.; Gilligan, C.; Watts, C.; Dexter, A.; Hall, D. Continuity of air-filled pores and invasion thresholds for a soil-borne fungal plant pathogen, Rhizoctonia solani. Soil Biol. Biochem. 1999, 31, 1803–1810.
- Otten, W.; Gilligan, C.A. Soil structure and soil-borne diseases: Using epidemiological concepts to scale from fungal spread to plant epidemics. Eur. J. Soil Sci. 2006, 57, 26–37.
- Kaisermann, A.; Maron, P.; Beaumelle, L.; Lata, J. Fungal communities are more sensitive indicators to non-extreme soil moisture variations than bacterial communities. Appl. Soil Ecol. 2015, 86, 158–164.
- Baveye, P.C.; Ebaveye, J.; Egowdy, J. Soil “ecosystem” services and natural capital: Critical appraisal of research on uncertain ground. Front. Environ. Sci. 2016, 4, 1–49.
- Demanèche, S.; Jocteur-Monrozier, L.; Quiquampoix, H.; Simonet, P. Evaluation of biological and physical protection against nuclease degradation of clay-bound plasmid DNA. Appl. Environ. Microbiol. 2001, 67, 293–299.
- Pietramellara, G.; Ascher, J.; Borgogni, F.; Ceccherini, M.T.; Guerri, G.; Nannipieri, P. Extracellular DNA in soil and sediment: Fate and ecological relevance. Biol. Fertil. Soils 2009, 45, 219–235.
- Romanowski, G.; Lorenz, M.G.; Sayler, G.; Wackernagel, W. Persistence of free plasmid DNA in soil monitored by various methods, including a transformation assay. Appl. Environ. Microbiol. 1992, 58, 3012–3019.
- Stotzky, G. Influence of soil mineral colloids on metabolic processes, growth, adhesion, and ecology of microbes and viruses. Geochem. Soil Radionucl. 2015, 17, 305–428.
- Nielsen, K.M.; Calamai, L.; Pietramellara, G. Stabilization of extracellular DNA and proteins by transient binding to various soil components. Soil Biol. 2006, 8, 141–157.
- Konhauser, K. Introduction to Geomicrobiology, 1st ed.; John Wiley & Sons: Hoboken, NJ, USA; Blackwell Publishing: Hoboken, NJ, USA, 2007.
- Rappoldt, C.; Crawford, J. The distribution of anoxic volume in a fractal model of soil. Geoderma 1999, 88, 329–347.
- Lombard, N.; Prestat, E.; van Elsas, J.D.; Simonet, P. Soil-specific limitations for access and analysis of soil microbial communities by metagenomics. FEMS Microbiol. Ecol. 2011, 78, 31–49.
- Nunan, N.; Schmidt, H.; Raynaud, X. The ecology of heterogeneity: Soil bacterial communities and C dynamics. Philos. Trans. R. Soc. B Biol. Sci. 2020, 375, 20190249.
- Strong, D.T.; Wever, H.D.E.; Merckx, R.; Recous, S. Spatial location of carbon decomposition in the soil pore system. Eur. J. Soil Sci. 2004, 55, 739–750.
- Salomé, C.; Nunan, N.; Pouteau, V.; Lerch, T.Z.; Chenu, C. Carbon dynamics in topsoil and in subsoil may be controlled by different regulatory mechanisms. Glob. Change Biol. 2010, 16, 416–426.
- Ruamps, L.S.; Nunan, N.; Pouteau, V.; Leloup, J.; Raynaud, X.; Roy, V.; Chenu, C. Regulation of soil organic C mineralisation at the pore scale. FEMS Microbiol. Ecol. 2013, 86, 26–35.
- Ruamps, L.S.; Nunan, N.; Chenu, C. Microbial biogeography at the soil pore scale. Soil Biol. Biochem. 2011, 43, 280–286.
- Frey-Klett, P.; Burlinson, P.; Deveau, A.; Barret, M.; Tarkka, M.; Sarniguet, A. Bacterial–fungal interactions: Hyphens between agricultural, clinical, environmental, and food microbiologists. Microbiol. Mol. Biol. Rev. 2011, 75, 583–609.
- Jansson, J.K.; Hofmockel, K.S. The soil microbiome—From metagenomics to metaphenomics. Curr. Opin. Microbiol. 2018, 43, 162–168.
- Bonfante, P.; Anca, I.-A. Plants, mycorrhizal fungi, and bacteria: A network of interactions. Annu. Rev. Microbiol. 2009, 63, 363–383.
- Deveau, A.; Bonito, G.; Uehling, J.; Paoletti, M.; Becker, M.; Bindschedler, S.; Hacquard, S.; Hervé, V.; Labbé, J.; Lastovetsky, O.A.; et al. Bacterial–fungal interactions: Ecology, mechanisms and challenges. FEMS Microbiol. Rev. 2018, 42, 335–352.
- Clarholm, M. Protozoan grazing of bacteria in soil—Impact and importance. Microb. Ecol. 1981, 7, 343–350.
- Kuikman, P.; van Elsas, J.; Jansen, A.; Burgers, S.; van Veen, J. Population dynamics and activity of bacteria and protozoa in relation to their spatial distribution in soil. Soil Biol. Biochem. 1990, 22, 1063–1073.
- Rutherford, P.M.; Juma, N.G. Influence of texture on habitable pore space and bacterial-protozoan populations in soil. Biol. Fertil. Soils 1992, 12, 221–227.
- Wright, D.A.; Killham, K.; Glover, L.A.; Prosser, J.I. Role of pore size location in determining bacterial activity during predation by protozoa in soil. Appl. Environ. Microbiol. 1995, 61, 3537–3543.
- Wright, D.; Killham, K.; Glover, L.; Prosser, J. The effect of location in soil on protozoal grazing of a genetically modified bacterial inoculum. Geoderma 1993, 56, 633–640.
- Männik, J.; Driessen, R.; Galajda, P.; Keymer, J.E.; Dekker, C. Bacterial growth and motility in sub-micron constrictions. Proc. Natl. Acad. Sci. USA 2009, 106, 14861–14866.
- Kravchenko, A.N.; Negassa, W.C.; Guber, A.K.; Hildebrandt, B.; Marsh, T.; Rivers, M.L. Intra-aggregate pore structure influences phylogenetic composition of bacterial community in macroaggregates. Soil Sci. Soc. Am. J. 2014, 78, 1924–1939.
- Dechesne, A.; Wang, G.; Gulez, G.; Or, D.; Smets, B.F. Hydration-controlled bacterial motility and dispersal on surfaces. Proc. Natl. Acad. Sci. USA 2010, 107, 14369–14372.
- Vos, M.; Wolf, A.B.; Jennings, S.J.; Kowalchuk, G.A. Micro-scale determinants of bacterial diversity in soil. FEMS Microbiol. Rev. 2013, 37, 936–954.
- Raynaud, X.; Nunan, N. Spatial ecology of bacteria at the microscale in soil. PLoS ONE 2014, 9, e87217.
- Pennell, K.D. Specific surface area. In Reference Module in Earth Systems and Environmental Sciences; Elias, S.A., Ed.; Elsevier: Amsterdam, The Netherlands, 2016; pp. 1–8.
- Boswell, G.P.; Jacobs, H.; Ritz, K.; Gadd, G.M.; Davidson, F. The development of fungal networks in complex environments. Bull. Math. Biol. 2007, 69, 605–634.
- Fricker, M.D.; Heaton, L.L.M.; Jones, N.S.; Boddy, L. The mycelium as a network. Fungal Kingd. 2017, 5, 335–367.
- Kohlmeier, S.; Smits, T.H.; Ford, R.M.; Keel, C.; Harms, H.; Wick, L.Y. Taking the fungal highway: Mobilization of pollutant-degrading bacteria by fungi. Environ. Sci. Technol. 2005, 39, 4640–4646.
- Warmink, J.A.; Nazir, R.; van Elsas, J.D. Universal and species-specific bacterial ‘fungiphiles’ in the mycospheres of different basidiomycetous fungi. Environ. Microbiol. 2009, 11, 300–312.
- Simon, A.; Hervé, V.; Al-Dourobi, A.; Verrecchia, E.; Junier, P. An in situ inventory of fungi and their associated migrating bacteria in forest soils using fungal highway columns. FEMS Microbiol. Ecol. 2016, 93, fiw217.
- Worrich, A.; Stryhanyuk, H.; Musat, N.; König, S.; Banitz, T.; Centler, F.; Frank, K.; Thullner, M.; Harms, H.; Richnow, H.-H.; et al. Mycelium-mediated transfer of water and nutrients stimulates bacterial activity in dry and oligotrophic environments. Nat. Commun. 2017, 8, 15472.
- Guhr, A.; Borken, W.; Spohn, M.; Matzner, E. Redistribution of soil water by a saprotrophic fungus enhances carbon mineralization. Proc. Natl. Acad. Sci. USA 2015, 112, 14647–14651.
- Pion, M.; Spangenberg, J.E.; Simon, A.; Bindschedler, S.; Flury, C.; Chatelain, A.; Bshary, R.; Job, D.; Junier, P. Bacterial farming by the fungus Morchella crassipes. Proc. R. Soc. B Boil. Sci. 2013, 280, 20132242.
- Haq, I.U.; Zhang, M.; Yang, P.; van Elsas, J.D. The interactions of bacteria with fungi in soil. Adv. Appl. Microbiol. 2014, 89, 185–215.
- Berthold, T.; Centler, F.; Hübschmann, T.; Remer, R.; Thullner, M.; Harms, H.; Wick, L.Y. Mycelia as a focal point for horizontal gene transfer among soil bacteria. Sci. Rep. 2016, 6, 36390.
- Nazir, R.; Mazurier, S.; Yang, P.; Lemanceau, P.; van Elsas, J.D. The ecological role of type three secretion systems in the interaction of bacteria with fungi in soil and related habitats is diverse and context-dependent. Front. Microbiol. 2017, 8, 38.
- Otto, S.; Bruni, E.P.; Harms, H.; Wick, L.Y. Catch me if you can: Dispersal and foraging of Bdellovibrio bacteriovorus 109J along mycelia. ISME J. 2016, 11, 386–393.
- Esser, D.S.; Leveau, J.H.; Meyer, K.M.; Wiegand, K. Spatial scales of interactions among bacteria and between bacteria and the leaf surface. FEMS Microbiol. Ecol. 2015, 91, 1–13.
- Gantner, S.; Schmid, M.; Dürr, C.; Schuhegger, R.; Steidle, A.; Hutzler, P.; Langebartels, C.; Eberl, L.; Hartmann, A.; Dazzo, F.B. In situ quantitation of the spatial scale of calling distances and population density-independent N-acylhomoserine lactone-mediated communication by rhizobacteria colonized on plant roots. FEMS Microbiol. Ecol. 2006, 56, 188–194.
- Cai, P.; Sun, X.; Wu, Y.; Gao, C.; Mortimer, M.; Holden, P.A.; Redmile-Gordon, M.; Huang, Q. Soil biofilms: Microbial interactions, challenges, and advanced techniques for ex-situ characterization. Soil Ecol. Lett. 2019, 1, 85–93.
- Schmidt, R.; Cordovez, V.; de Boer, W.; Raaijmakers, J.; Garbeva, P. Volatile affairs in microbial interactions. ISME J. 2015, 9, 2329–2335.
- Effmert, U.; Kalderás, J.; Warnke, R.; Piechulla, B. Volatile mediated interactions between bacteria and fungi in the soil. J. Chem. Ecol. 2012, 38, 665–703.
- Nazir, R.; Warmink, J.A.; Boersma, H.; van Elsas, J.D. Mechanisms that promote bacterial fitness in fungal-affected soil microhabitats. FEMS Microbiol. Ecol. 2009, 71, 169–185.
- Bignell, E. The molecular basis of pH sensing, signaling, and homeostasis in fungi. Adv. Appl. Microbiol. 2012, 79, 1–18.
- Braunsdorf, C.; Mailänder-Sánchez, D.; Schaller, M. Fungal sensing of host environment. Cell. Microbiol. 2016, 18, 1188–1200.
- Lurthy, T.; Cantat, C.; Jeudy, C.; Declerck, P.; Gallardo, K.; Barraud, C.; Leroy, F.; Ourry, A.; Lemanceau, P.; Salon, C.; et al. Impact of bacterial siderophores on iron status and ionome in pea. Front. Plant Sci. 2020, 11, 1–12.
- Haas, H. Fungal siderophore metabolism with a focus on Aspergillus fumigatus. Nat. Prod. Rep. 2014, 31, 1266–1276.
