Therapeutic bacteriophages, commonly called as phages, are a promising potential alternative to antibiotics in the management of bacterial infections of a wide range of organisms including cultured fish. Their natural immunogenicity often induces the modulation of a variated collection of immune responses within several types of immunocytes while promoting specific mechanisms of bacterial clearance.
- aquaculture
- bacteriophages
- disease management
- fish
- immunology
- lytic enzymes
- pathogens
1. Phage Biology and Spatial Distribution
2. Phage’s Life Cycle
The phages like any other viruses depend on the metabolism of their bacterial host for reproduction. During the reproductive process, most phage types completely consume the resources of their host and kill them when releasing their progeny [6]. Initially, phages must infect their host bacteria through the binding of specific receptors that selectively sense specific components of the target bacterial cell wall such as the lipopolysaccharide in Gram-negative, or peptidoglycan in Gram-positive, capsular polysaccharides, and superficial appendages such as pili and flagella [7][8][9]. Following the classical viral reproductive strategies, once the phage inserts their nucleic acid into the bacterium’s cytoplasm, the host cellular machinery is highjacked to induce extensive replication through the lytic cycle (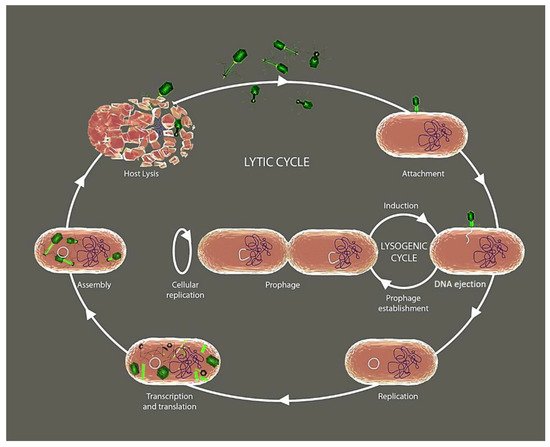
3. Phage Lytic Enzymes and Depolymerases
Lysins derived from phages degrade bacterial peptidoglycans and are classified into five groups, depending on the bonds these enzymatic proteins cleave in the bacterial peptidoglycan [11][16]. Although their function is exclusively to degrade the cell wall of bacteria, the lytic enzymes of phages present a tremendous structural diversity and a significant number of different mechanisms of action [12][13][14][15][17,18,19,20].
In general, lysins are more likely to lyse Gram-positive bacteria because their cell wall peptidoglycan is directly exposed on the cell surface unlike Gram-negative bacteria. However, the study of phages or their lysins has been limited to a few fish pathogens such as Streptococcus agalactiae, Lactococcus garvieae, Renibacterium salmoninarum, Streptococcus iniae, and S. dysgalactiae, which are highly associated with disease outbreaks in fish farms.
4. Interactions between Phage and the Fish Immune System
4.1. Phage-Mediated Activation of Inflammation
Bacteriophage treatment was associated with opposite shifts in the inflammatory response in several test models, both in vivo and in vitro [16][17][18][19]. However, the results seem to depend not only on the cellular or animal model used but also on the type of phage applied and the panel of cytokines analyzed. Phage therapy in humans can also modify the levels of some cytokines produced by blood cells in treated patients [20]. In fish, some researchers have analyzed the cytokines’ response to the presence of bacteriophages alone or the coinfection of phages with their target bacteria. For example, phage therapy reduced the expression of the proinflammatory cytokines tnfa and il1b in the inflammatory response generated by Pseudomonas aeruginosa infection in zebrafish embryos [21][22]. Besides, using the adult zebrafish (Danio rerio) and the E. tarda model of infection, other authors also showed that although a phage treatment induced the expression of cytokine genes at specific time points, a robust proinflammatory response was undetected in the host [23]. Furthermore, a recent study has shown that a phage lysate of A. hydrophila induced a more robust immune response in Cyprinus carpio when compared to a formalin killed vaccine [24]. As a proof-of-concept, a novel commercial preparation containing three bacterial phages (BAFADOR4.1. Phage-Mediated Activation of Inflammation
4.2. Phage-Specific Adaptive Responses
Due to the protein structure of the phage envelope, these proteins are the target of the adaptive immune system, which response with the production of neutralizing antibodies against them. Early studies with mice and even amphibians showed that phage exposure of the animals induced primary and secondary antibody responses [26][27][28]. It is expected that some phage epitopes stimulate an antibody response in experimental models. However, antibody production depends on the route of phage administration, the application schedule and dose, and individual features of a phage. Consequently, the results of studies where an antibody response to phages has been verified are very heterogeneous. Phagocytosis by immune patrolling cells seems to be a significant process of bacteriophage neutralization within animal bodies [29]. Moreover, although blood in humans and animals, including fish, is deemed sterile, genomic analysis has shown a rich phage community, which inevitably comes into continuous contact with immune cells in this rich fluid [30]. Despite these mechanisms of phagocytosis, antigen presentation, and antibody production by the immune cells against phages, the number of antibodies produced does not affect phage therapy outcomes. On the other hand, due to the numerous and constant presence of large numbers of phages in our microbiota, it is not surprising that a low but stable background of antibodies against them is produced. Therefore, in some human or animal tests, high antibody levels have not been found against the phages used. Phage-derived RNA and ssDNA could directly contribute to B cell activation and the synthesis of anti-bacteriophage antibodies [31][32]. Despite the production of antibodies by animals against phage core or tail proteins, the induction of antibodies seems irrelevant for treating infections because the antibacterial effects of phages are faster than antibody formation in acute infections [33]. Conversely, the production of antibodies against phages could interfere with the outcome of the infection in chronic infections [34]. However, no robust studies have demonstrated an antibody-mediated immune response after inoculation or experimental infection with phages in fish.4.2. Phage-Specific Adaptive Responses
5. Potential of Phage Therapy in Aquaculture Settings
During the fish and shellfish production cycle, these animals are already in daily contact with billions of bacteriophages, which assures us that they are safe. However, in their use against bacterial infections where massive phage production is required, we must consider several factors. As phage treatments constantly require isolating the bacterium causing the disease, once a helpful phage is characterized against this bacterial strain, a stable batch of technically challenging preparations must be produced for field use. Consequently, one of the most critical challenge for microbiologists working directly or indirectly with aquaculture is the standardization of stocks used to treat infections or combat biofilms in aquaculture facilities. These stocks require strict quality control for purity, viability, and stability, implying that the correct conservation of the stocks is necessary for preparations containing single or mixed phages (phage cocktail). Titer, dosage, and quality of phage preparations are crucial parameters in standardizing experiments in the laboratory and experimental infections in field trials. Since we know that while some phages can grow exponentially inside a bacterial population from a low initial concentration, other phages need to maintain a relationship between the number of bacteria and the number of phage particles to achieve an adequate performance. Therefore, we must empirically verify this critical parameter. Very recently, a phage cocktail containing seven bacteriophages (three against A. hydrophila and four against P. fluorescens) has been tested in the European eel (Anguilla anguilla) and rainbow trout (Oncorhynchus mykiss), reducing the mortality of fish challenged with strains of these two species of bacteria [25][35]. Cocktails have also been used successfully in laboratory tests or small field trials in food protection or veterinary and human medicine [36][37][38][39]. In these and other studies, many phages (cocktail) are used to carry out the experiments, but in most cases, only the phage that has presented better results in vitro is subsequently characterized [40][41][42][43]. Second, it would be desirable to know phage genetics with sufficient precision. After all, we must consider that when we intend to use bacteriophages in aquaculture, they may contain genes for resistance to antibiotics or bacterial virulence genes that can produce noticeable side effects because they replicate exponentially in contact with their target bacteria. We must also remember that many antibiotic residues end up in continental or oceanic waters due to anthropogenic activities. Therefore, we must be aware that even phages isolated from aquatic environments can carry antibiotic resistance genes or virulence factors [44][45]. At present, although each time their number increases, not all phages used in in vitro or in vivo assays against fish or shellfish bacterial pathogens have been entirely genetically analyzed or characterized (| Gram-Negative Targets |
Source | SourceEnrichment ɸ | Characterization Method | Enrichment Phage Strains Name | ɸ | Characterization MethodFamily * | Genome Length | References | |||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Phage Strains Name | Family * | Genome Length | References | ||||||||||
| Aeromonas hydrophila | River water | No | TEM | ɸ2 and ɸ5 | Myoviridae | ~20 kb | [52] | ||||||
| Fishponds; Polluted rivers | Single | TEM | N21, W3, G65, Y71 and Y81 | Myoviridae; Podoviridae | n.d. | [ | |||||||
| Lactococcus garvieae | L. garvieae isolated from diseased yellowtail | No | TEM, dsDNA | PLgY(16) | Siphoviridae | n.d. | [134]53] | ||||||
| Yellowtail (Y) Water (W) Sediments (S) |
Single | TEM, dsDNA | PLgW1-6 PLgY16 PLgY30 PLgY886 PLgS1 |
Siphoviridae | >20 kbp | [135][136][137] | Stream water | Single | TEM, dsDNA | pAh-1 | Myoviridae | ~64 kb | [54] |
| Domestic compost | Single | GE1 | Siphoviridae | 24,847 bp | [138] | Sea water | Single | TEM, DNA sequencing | Akh-2 | Siphoviridae | 114,901 bp | [55] | |
| L. garvieae host | No | TEM, DNA sequencing | PLgT-1 | Siphoviridae | 29,284 bp | [139][140][141] | Carp tissues | Single | TEM | AHP-1 | Myoviridae | n.d. | [56] |
| Lake water | Single | TEM, dsDNA, DNA sequencing | AhyVDH1 | Myxoviridae | 39,175 bp | [57] | |||||||
| River water | No | TEM, dsDNA, DNA sequencing | MJG | Podoviridae | 45,057 bp | [58] | |||||||
| Sewage water | Single | TEM | AH1 | n.d. | n.d. | [59] | |||||||
| Striped catfish pond water | Single | TEM, dsDNA, DNA sequencing | PVN02 | Myoviridae | 51,668 bp | [60][61] | |||||||
| River water | TEM, dsDNA | pAh1-C pAh6-C |
Myoviridae | 55 kb 58 kb |
[62] | ||||||||
| Wastewater | No | TEM, dsDNA, DNA sequencing | Ahp1 | Podoviridae | ~42 kb | [63] | |||||||
| Aeromonas punctata | Stream water | Single | TEM, dsDNA | IHQ1 | Myoviridae | 25–28 kb | [64] | ||||||
| Aeromonas salmonicida | River waters, two passing through fish farms | Single | TEM, DNA sequencing | SW69-9 L9-6 Riv-10 |
Myoviridae | 173,097 bp, 173,578 bp and 174,311 bp | [65] | ||||||
| River water | Single | TEM, DNA sequencing | phiAS5 | Myoviridae | 225,268 bp | [66] | |||||||
| Sediment of a Rainbow trout culture farm | Single | TEM, dsDNA, DNA sequencing | PAS-1 | Myoviridae | ~48 kb | [67] | |||||||
| Rainbow trout farm water | Wastewater from a seafood market | No | TEM, DNA sequencing | AsXd-1 | Siphoviridae | 39,014 bp | [68] | ||||||
| Sewage network water from a lift station | Single | TEM | AS-A AS-D AS-E |
Myoviridae | n.d. | [40][41] | |||||||
| River water | No | TEM | HER 110 | Myoviridae | n.d. | [69][70] | |||||||
| Aeromonas spp. | Gastrointestinal content of variated fish species | No | TEM, DNA sequencing | phiA8-29 | Myoviridae | 144,974 bp | [71][72] | ||||||
| Citrobacter freundii | Sewage water | No | TEM, DNA sequencing | IME-JL8 | Siphoviridae | 49,838 bp | [73] | ||||||
| Edwardsiella ictaluri | Water from catfish ponds | Single | TEM, dsDNA, DNA sequencing | eiAU eiDWF eiMSLS |
Siphoviridae | 42.80 kbp 42.12 kbp 42.69 kbp |
[74][75] | ||||||
| Single | River water | Multiple | DNA Sequencing | PEi21 | Myoviridae | 43,378 bp | [76][77] | ||||||
| Striped catfish kidney and liver | Single | TEM, dsDNA | MK7 | Myoviridae | ~34 kb | [78] | |||||||
| Edwardsiella tarda | Seawater | Single | TEM, dsDNA | ETP-1 | Podoviridae | ~40 kb | [23] | ||||||
| River water | No | TEM, DNA sequencing | pEt-SU | Myoviridae | 276,734 bp | [79] | |||||||
| Wastewater | Single | DNA sequencing | PETp9 | Myoviridae | 89,762 bp | [80] | |||||||
| Fish tissues and rearing seawater | No | TEM, DNA sequencing | GF-2 | Myoviridae | 43,129 bp | [81] | |||||||
| Flavobacterium columnare | River water | Single | TEM, DNA sequencing | FCL-2 | Myoviridae | 47,142 bp | [82][83][84] | ||||||
| TEM, DNA sequencing | WP-2 | Fishpond’s water and bottom sediments | No | TEM, dsDNA | FCP1-FCP9 | Podoviridae | n.d. | [42] | |||||
| Flavobacterium psychrophilum | Rainbow trout farm water | Single/double | TEM, dsDNA | ø (FpV-1 to FpV-22) | Podoviridae Siphoviridae Myoviridae |
(~8 to ~90 kb) | [85][86] | ||||||
| Ayu kidneys and pondwater collected from ayu farms | Multiple | TEM, dsDNA | PFpW-3, PFpC-Y PFpW-6, PFpW-7 PFpW-8 |
Myoviridae; Podoviridae; Siphoviridae | n.d. | [87] | |||||||
| Picovirinae | 18,899 bp | [ | 142 | Photobacterium damselae subsp. damselae | Raw oysters | Single | TEM, dsDNA | Phda1 | Myoviridae | 35.2–39.5 kb | [88] | ||
| Gastrointestinal tract of lollipop catshark | Single | TEM, DNA sequencing | vB_Pd_PDCC-1 | Myoviridae | 237,509 bp | [89] | |||||||
| Pseudomonas plecoglossicida | Ayu pond water and diseased fish | No | TEM, DNA sequencing | PPpW-3 PPpW-4 |
Myoviridae Podoviridae | 43,564 bp 41,386 bp |
[90][91] | ||||||
| ] | |||||||||||||
| Streptococcus agalactiae | Tilapia pond | No | TEM | HN48 | Caudoviridae | Pseudomonas aeruginosa | Wastewater | No | TEM, DNA sequencing | MBL | n.d. | 42,519 bp | [92] |
| Shewanella spp. | Wastewater from a marketplace |
Single | TEM, DNA sequencing | SppYZU01 to SppYZU10 | Myoviridae; Siphoviridae. | SppYZU01 (43.567 bp) SppYZU5 (54.319 bp) |
[93] | ||||||
| Tenacibaculum maritimum | Seawater | Multiple | TEM, DNA sequencing | PTm1 PTm5 |
Myoviridae | 224,680 bp 226,876 bp |
[94] | ||||||
| Vibrio alginolyticus | Aquaculture tank water | Single | TEM, DNA sequencing | VEN | Podoviridae | 44,603 bp | [95] | ||||||
| Marine sediment | No | TEM, DNA sequencing | ValKK3 | Myoviridae | 248,088 bp | [96] | |||||||
| Marine water | Single | TEM, dsDNA | St2 Grn1 |
Myoviridae | 250,485 bp 248,605 bp | [97] | |||||||
| n.d. | [ | 143 | ] | ||||||||||
| S. iniae | S. iniae host | No | Vibrio anguillarum | Soft tissues from clams and mussels | No | TEM, dsDNA | 309 ALMED CHOED ALME CHOD CHOB |
Several shapes | ~47–48 kb | [98] | |||
| Sewage water | Double | dsDNA | VP-2 VA-1 |
n.d. | n.d. | [51] | |||||||
| Water samples from fish farms | Multiple | TEM, DNA sequencing | ø H1, H7, S4-7, H4, H5 H8, H20 S4-18, 2E-1, H2 |
Myoviridae Siphoviridae Podoviridae | ~194–195 kb ~50 kb ~45–51 kb |
[99] | |||||||
| Vibrio campbellii | Host strain (V. campbellii) isolated form a dead shrimp | No | TEM, DNA sequencing | HY01 | Siphoviridae | 41.772 bp | [100] | ||||||
| Hepatopancreas of Pacific white shrimp |
Single | dsDNA, DNA sequencing | vB_Vc_SrVc9 | Autographiviridae | ~43.15 kb | [101] | |||||||
| Vibrio harveyi | Shrimp farm, hatcheries and marine water | Multiple | TEM, dsDNA | A | Siphoviridae | n.d. | [102] | ||||||
| Vibrio harveyi | No | TEM, dsDNA | VHML | Myovirus-like | n.d. | [103] | |||||||
| Shrimp pond water | Single | TEM, dsDNA | PW2 | Siphoviridae | ~46 kb | [104] | |||||||
| Water and sediment samples | Single | TEM, dsDNA | VHM1, VHM2 VHS1 |
Myoviridae, Siphoviridae |
~55 kb, ~66 kb ~69 kb |
[105] | |||||||
| Hatchery water and oyster tissues | Single | TEM, dsDNA | vB_VhaS-a vB_VhaS-tm |
Siphoviridae | ~82 kb ~59 kb |
[106] | |||||||
| Commercial clam samples | Multiple | Genomic analysis, dsDNA | ø VhCCS-01 VhCCS-02 VhCCS-04 VhCCS-06 VhCCS-17 VhCCS-20 VhCCS-19 VhCCS-21 |
Siphoviridae, Myoviridae |
n.d. | [107] | |||||||
| Oyster, clam, shrimp, and seawater samples | No | TEM, DNA sequencing | VHP6b | Siphoviridae | 78,081 bp | [108] | |||||||
| shrimp hatchery and farm water, oysters from estuaries, coastal sea water |
Multiple | TEM, dsDNA | Viha10 Viha8 Viha9 Viha11 Viha1 to Viha7 |
Siphoviridae - Siphoviridae Myoviridae (Viha4) |
n.d. ~44–94 kb ~85 kb (Viha4) |
[109][110] | |||||||
| Seawater sample | Single | TEM | VhKM4 | Myoviridae | n.d. | [111] | |||||||
| Vibrio ordalii | Macerated specimens of mussels | No | TEM, DNA sequencing | B_VorS-PVo5 | Siphoviridae | 80,578 bp | [112] | ||||||
| Vibrio parahaemolyticus | Sewage sample | No | TEM, dsDNA | VPp1 | Tectiviridae | ~15 kb | [113] | ||||||
| Polluted seawater | No | TEM, dsDNA | KVP40 KVP41 |
Myoviridae | n.d. | [114][115] | |||||||
| Seawater or mussels | Single | dsDNA | SPA2 SPA3 |
n.d. | ~21 kb | [116] | |||||||
| Coastal water | Single | TEM, DNA sequencing | pVP-1 | Siphoviridae | 111,506 bp | [117][118] | |||||||
| V. parahaemolyticus isolated from sewage samples collected from an aquatic product market | No | TEM, DNA sequencing | vB_VpS_BA3 vB_VpS_CA8 | Siphoviridae | 58,648 bp 58,480 bp |
[119] | |||||||
| Shrimp pond water | Single | TEM, DNA sequencing | VP-1 | Myoviridae | Yersinia ruckeri | Wastewater containing suspended trout feces from a settling pond at a trout farm | Single | TEM | NC10 | Podoviridae | n.d. | [49] | |
| TEM, DNA sequencing | Sewage | No | TEM | YerA41 (several phages) | icosahedral head, contractile tail | n.d. | [131] | ||||||
| Sewage | No | TEM, DNA sequencing, dsDNA | R1-37 | Myoviridae | ~270 kb | [132][133] |
| Gram-Positive Targets | |||||||
|---|---|---|---|---|---|---|---|
| TEM, dsDNA | |||||||
| vB_SinS-44 vB_SinS-45 vB_SinS-46 vB_SinS-48 | |||||||
| Siphoviridae | |||||||
| ~51.7 kb | ~28.4 kb | ~66.3 kb | ~27.5 kb |
[144] | |||
| Weissella ceti | W. ceti host strain | No | TEM | PWc | Siphoviridae | 38,783 bp | [145] |
| 150,764 bp | |||||||
| [ | |||||||
| 120 | |||||||
| ] | |||||||
| Coastal sand sediment | |||||||
| double | |||||||
| TEM, DNA sequencing | |||||||
| VpKK5 | |||||||
| Siphoviridae | |||||||
| 56,637 bp | |||||||
| [ | |||||||
| 121 | |||||||
| ] | |||||||
| [ | |||||||
| 122 | |||||||
| ] | |||||||
| Vibrio splendidus | |||||||
| Raw sewage obtained from local hatcheries | |||||||
| Single | |||||||
| TEM | |||||||
| PVS-1, PVS-2 | |||||||
| PVS-3 | |||||||
| Myoviridae; Siphoviridae | |||||||
| n.d. | |||||||
| [ | |||||||
| 123 | |||||||
| ] | |||||||
| Seawater near a fish farm cage | |||||||
| Single | |||||||
| TEM, DNA sequencing | vB_VspP_pVa5 | Podoviridae | 78,145 bp | [124] | |||
| Vibrio coralliilyticus | sewage in oyster hatchery | Single | TEM | pVco-14 | Siphoviridae | n.d. | [125] |
| Vibrio vulnificus | Seawater sample | Single | TEM, DNA sequencing | SSP002 | Siphoviridae | 76,350 bp | [126][127] |
| Abalone samples | No | TEM, sequencing | VVPoo1 | Siphoviridae | 76,423 bp | [128] | |
| Initial host strain (V. vulnificus) | No | TEM | VV1 VV2 VV3 VV4 |
Tectiviridae | n.d. | [129] | |
| Vibrio sp. | Sewage draining exits | Single | TEM, DNA sequencing | VspDsh-1 VpaJT-1 ValLY-3 ValSw4-1 VspSw-1 |
Siphoviridae | 46,692 bp 60,177 bp 76,310 bp 79,545 bp 113,778 bp |
[130] |
