Secreted a disintegrin-like and metalloprotease with thrombospondin type 1 motif (ADAMTS) proteases play crucial roles in tissue development and homeostasis. The biological and pathological functions of ADAMTS proteases are determined broadly by their respective substrates and their interactions with proteins in the pericellular and extracellular matrix. For some ADAMTS proteases, substrates have been identified and substrate cleavage has been implicated in tissue development and in disease. For other ADAMTS proteases, substrates were discovered in vitro, but the role of these proteases and the consequences of substrate cleavage in vivo remains to be established. Mutations in ADAMTS10 and ADAMTS17 cause Weill–Marchesani syndrome (WMS), a congenital syndromic disorder that affects the musculoskeletal system (short stature, pseudomuscular build, tight skin), the eyes (lens dislocation), and the heart (heart valve abnormalities). WMS can also be caused by mutations in fibrillin-1 (FBN1), which suggests that ADAMTS10 and ADAMTS17 cooperate with fibrillin-1 in a common biological pathway during tissue development and homeostasis.
- extracellular matrix
- ADAMTS proteases
- fibrillin
- microfibrils
- Weill-Marchesani syndrome
- short stature
- lens dislocation
1. The ADAMTS Protease Family
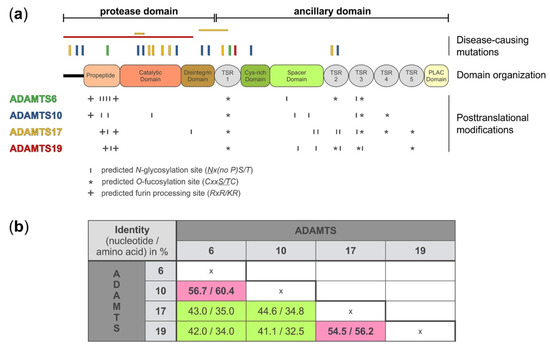
2. Domain Organization and Posttranslational Modifications of ADAMTS6, 10, 17, and 19
On the protein level, ADAMTS6, 10, 17, and 19 share the same domain organization (Figure 1a). However, each protease pair arose from distinct gene duplication events [30]. When comparing the nucleotide and amino acid sequences between the four proteases, it is evident that ADAMTS10 sequences are more similar to ADAMTS6, where 60% of the amino acid residues are identical, and ADAMTS17 sequences are more similar to ADAMTS19, with 56% of the amino acid residues being identical (Figure 1b) [1][31]. Despite the evolutionary homology, the identical domain organization, and the relatively high amino acid sequence identity, the ADAMTS proteases that form the individual protease pairs, ADAMTS6/ADAMTS10 and ADAMTS17/ADAMTS19, appear to have distinct biological functions, based on their involvement in different human disorders (see below). One possible explanation for the diversification in the function of these proteases could be differences in posttranscriptional and posttranslational modifications. Alternative splicing is a posttranscriptional mechanism that can expand the diversity and thus function of ADAMTS proteases by generating different isoforms. ADAMTS proteases have several splice variants based on the NCBI protein database. There are 13 isoforms listed for ADAMTS6, 4 for ADAMTS10, 12 for ADAMTS17, and 5 for ADAMTS19. For most of these isoforms, tissue-specific expression or functional data are not available. However, by homology mapping with ADAMTS10 as a template and analysis of expressed sequence tags in the GenBank™ database the existence of at least two splice variants for ADAMTS6 were predicted and subsequently shown experimentally in epithelial cells [32][33]. In addition, northern blot analysis of total mRNA isolated from adult human tissue demonstrated two ADAMTS10 mRNA species that differed in size, suggesting alternative splicing of ADAMTS10 mRNA as well [33]. Two isoforms of ADAMTS17 with distinct expression patterns have been described previously [10]. Our own unpublished data show expression of at least three additional isoforms of ADAMTS17 that differ in the sequence of the spacer domain (Balic, et al., manuscript in preparation). In addition to alternative splicing, ADAMTS6, 10, 17, and 19 show differences in the number and location of predicted and experimentally verified posttranslational modifications, such as furin/PACE-processing, autocatalysis, N-glycosylation, or O-fucosylation. Based on western blot analysis, ADAMTS6 and ADAMTS19 are furin-processed but do not undergo apparent autocatalysis (Karoulias et al., unpublished data for ADAMTS19) [34]. However, a direct comparison between active ADAMTS6 and an inactive mutant form was not shown to completely rule out the possibility of ADAMTS6 autocatalysis. The propeptide of ADAMTS17 is also processed by furin, but in contrast to ADAMTS6 and ADAMTS19, ADAMTS17 undergoes extensive autoproteolysis at the cell surface or in the ECM [22]. Furin-processing of the ADAMTS17 propeptide is not required to activate ADAMTS17. Instead, the ADAMTS17 propeptide may act as a chaperone to facilitate ADAMTS17 secretion, since removal of the propeptide did abolish ADAMTS17 secretion or its release from the cell surface [22]. A similar role was previously described for the propeptide of ADAMTS9 [35]. ADAMTS10 on the other hand has a degenerated consensus sequence for furin-processing (GLKR instead of RLKR) and thus the propeptide remains covalently associated with the ADAMTS10 protease after secretion. Wild type ADAMTS10 has weak protease activity against fibrillin-1 [36]. However, upon restoration of the consensus furin-processing site in recombinant ADAMTS10, the propeptide was efficiently excised and proteolytic activity of ADAMTS10 against fibrillin-1 was enhanced [36]. ADAMTS proteases can be N-glycosylated and O-fucosylated. ADAMTS6 and ADAMTS10 contain six predicted N-glycosylation sites but their location is different (Figure 1a and Table 1). For example, four of the six predicted N-glycosylation sites in ADAMTS6 are located in the propeptide domain while ADAMTS10 harbors only two of the six predicted N-glycosylation sites in the propeptide. Interestingly, one of the predicted N-glycosylation sites in ADAMTS10 is located in the catalytic domain and it is tempting to speculate that this site may modulate proteolytic activity if differentially glycosylated. ADAMTS17 has seven predicted N-glycosylation sites with five of them being located in the ancillary domain. In contrast, ADAMTS19 contains only five predicted N-glycosylation sites. In addition to N-glycosylation, TSR domains of ADAMTS proteases can undergo O-fucosylation at the Cxx(S/T)C consensus motif as part of a non-canonical quality control pathway in the endoplasmic reticulum [37][38]. Three of the five TSR domains of ADAMTS6 and ADAMTS10 and four of the five TSR domains of ADAMTS17 and ADAMTS19 contain the consensus sequence for O-fucosylation [39]. For ADAMTS17, it was shown that TSR1, 3, and 5 were fully modified [22]. TSR4 was not O-fucosylated, despite the presence of the consensus motif. Functionally, the absence of O-fucosylation on TSR3 and TSR5, and to a lesser extent on TSR1, resulted in the lack of secretion of recombinant ADAMTS17 [22]. It is anticipated that O-fucosylation of TSRs in ADAMTS6, 10, and 19 would play a similar role in protein quality control and protein secretion. However, experimental data is lacking and the degree of O-fucosylation needs to be determined for each of these proteases.| ADAMTS6 | ADAMTS10 | ADAMTS17 | ADAMTS19 | |
|---|---|---|---|---|
| N-Glycosylation sites 1 | 6 | 6 | 7 | 5 |
| O-Fucosylation sites 2 | 3 | 3 | 4 | 4 |
| Furin consensus sequences 3 | 2 | 1 | 2 | 2 |
| Furin processing 4 | yes | no | yes | yes |
| Autocatalysis | no | no | yes | no |
| Substrates | LTBP1, SDC4 | FBN1, FBN2 | ADAMTS17 | n.d. |
| Alternative splicing | yes | yes | yes | likely |
| Human disorders | Prolonged QRS syndrome | WMS 1 (MIM #277600) |
WMS 4 (MIM #613195) |
Non-syndromic heart valve disease |
| Disease-causing human mutations | 2 | 9 | 9 | 2 |
| Gene knockout phenotype in mice | Prenatal/neo-natal lethality, double outlet right ventricle, ventricular hypertrophy, atrial and ventricular septal defects |
Some prenatal/ neonatal lethality, abnormal ciliary zonule, shorter long bones due to growth plate abnormalities, skeletal muscle abnormalities | Some prenatal/ neonatal lethality, skeletal growth impairment due to growth plate abnormalities, brachydactyly by 8 mos. of age | Aortic valve dysfunction in ~40% of knockout mice |
| Fibrillin-1 binding | n.d. | yes | yes | n.d. |
| Fibrillin microfibril formation 5 | n.d. | Promotes FBN1 deposition | No effect | n.d. |
3. Gene Expression Patterns of ADAMTS6, 10, 17, and 19
The biological functions of ADAMTS6, 10, 17, and 19 are not only determined by differential posttranscriptional and posttranslational modifications, which may guide ECM localization and substrate binding and cleavage, but also by the cell types and tissues that express these proteases. Adamts10 for example is widely expressed in most adult tissues in mice and almost universally expressed during mouse embryonic development with the exception of the ectoderm [20][33]. RNA in situ hybridization demonstrated dynamic Adamts10 expression, which was generally low during early embryonic development [embryonic day (E) 9.5 to E12.5] and induced in later stages of development (E14.5 and E17.5) [33]. At E14.5, Adamts10 was expressed in several craniofacial tissues, notably in the perichondrium and periosteum, but not in cartilage, of newly forming bones of the mandible, and in the tongue muscle. In developing lungs, Adamts10 was expressed in the connective tissue between the bronchi and the accompanying blood vessels. Adamts10 was also expressed in the stomach, duodenum, pancreas, dorsal root ganglia, and primary ossification centers of vertebrae, but was absent in the liver. In the developing musculoskeletal system, Adamts10 mRNA was detected between the cartilaginous metacarpals and metatarsals of the hand and feet and in dense connective tissues such as the joint capsule and developing tendons and ligaments. At E17.5 Adamts10 expression was strong in the cartilage of developing bones and in the wall of large arteries. Adamts10 was expressed in several ocular tissues during development, including non-pigmented ciliary epithelium, lens fiber cells, and parts of the retina [20]. Expression of ADAMTS10 in the eye is relevant for the characteristic ectopia lentis phenotype observed in individuals with Weill–Marchesani syndrome (WMS). All these data support a role for ADAMTS10 in the early steps of tissue and organ formation. Adamts10 continued to be expressed in adult chondrocytes, tendon, and skeletal muscle in mice [20]. In human adult tissue, ADAMTS10 mRNA was expressed in the heart, brain, lung, pancreas, but ADAMTS10 expression was notably low or absent in skeletal muscle [33]. Given the broad expression of ADAMTS10 during embryonic development, it was somewhat surprising, that most Adamts10 knockout mice apparently undergo normal development, with limited embryonic lethality (see below) [19][20]. Similar to ADAMTS10, ADAMTS17 is widely expressed in many fetal and adult tissues in humans [10]. Two isoforms of ADAMTS17, isoform a (22 exons) and isoform b (16 exons), with distinct expression patterns, were identified. Both ADAMTS17 isoforms were expressed in lung, brain, and the eye, including the retina. Immunostaining in human normal and cancer tissues showed widespread expression in multiple tissues to varying degrees [10]. However, no images were shown and the reported localization of ADAMTS17 in the cytoplasm raises the possibility that the antibody used in this study was non-specific in tissue staining, even though it recognizes recombinant ADAMTS17 protein [22]. In-situ hybridization studies in mouse embryonic tissue at E16.5 and in neonates demonstrated Adamts17 expression in the eye, especially at the lens equator and in the trabecular region [22]. Apart from the eye, Adamts17 mRNA was detected in the perichondrium, the intervertebral disc, and in the smooth muscle cell layer of blood vessel walls. The most pronounced expression of Adamts17 was observed in lung parenchyma, skin epidermal basal cell layer, and in developing hair follicles. Adamts17 mRNA was notably absent from the growth plate of long bones, the endothelium of blood vessels, and the bronchial epithelium in lung tissue. Less detailed information about the expression patterns of ADAMTS6 and ADAMTS19 is currently available. ADAMTS6 mRNA was detected in the mouse heart, specifically in the outflow tract, the heart valves, the atria, and the ventricular myocardium [34]. In northern blot analysis, ADAMTS6 mRNA was weakly expressed in placental tissue, but was not detected in mouse embryonic development or in adult human tissues [40]. ADAMTS19 mRNA was detected in fetal lung and osteosarcoma tissue [41]. More recently, Adamts19 was shown to be expressed in the heart valve at all stages of heart valve formation and elongation and in bone and cartilage anlagen at E10.5 [42]. Taken together, ADAMTS10 and ADAMTS17 show broad, both overlapping and distinct gene expression patterns in tissues. Therefore, it is possible that ADAMTS10 and ADAMTS17 may compensate for each other in mouse knockout models of the individual proteases at least in some tissues (see below). If these proteases can indeed compensate for each other in murine loss-of-function models, similar to what has been observed for the ADAMTS7/ADAMTS12 and ADAMTS9/ADAMTS20 pairs, needs to be explored [27][28]. No compensation of ADAMTS10 with ADAMTS17 or vice versa appears to occur in tissues affected in WMS, specifically the eye and the musculoskeletal system. However, a lack of cardiac involvement and joint contractures in WMS4 due to ADAMTS17 mutations leaves the possibility that compensation with ADAMTS10, but not vice versa, causes the lack of cardiac and joint presentations in WMS4 (see below). ADAMTS6 and ADAMTS19 appear to have more tissue-specific expression patterns, which are reflected in the specific diseases associated with mutations in ADAMTS6 and ADAMTS19, as described in the next section.References
- Apte, S.S. A disintegrin-like and metalloprotease (reprolysin-type) with thrombospondin type 1 motif (ADAMTS) superfamily: Functions and mechanisms. J. Biol. Chem. 2009, 284, 31493–31497.
- Akiyama, M.; Takeda, S.; Kokame, K.; Takagi, J.; Miyata, T. Crystal structures of the noncatalytic domains of ADAMTS13 reveal multiple discontinuous exosites for von Willebrand factor. Proc. Natl. Acad. Sci. USA 2009, 106, 19274–19279.
- Ai, J.; Smith, P.; Wang, S.; Zhang, P.; Zheng, X.L. The proximal carboxyl-terminal domains of ADAMTS13 determine substrate specificity and are all required for cleavage of von Willebrand factor. J. Biol. Chem. 2005, 280, 29428–29434.
- Tortorella, M.; Pratta, M.; Liu, R.Q.; Abbaszade, I.; Ross, H.; Burn, T.; Arner, E. The thrombospondin motif of aggrecanase-1 (ADAMTS-4) is critical for aggrecan substrate recognition and cleavage. J. Biol. Chem. 2000, 275, 25791–25797.
- Takeda, S. ADAM and ADAMTS Family Proteins and Snake Venom Metalloproteinases: A Structural Overview. Toxins 2016, 8, 155.
- Karoulias, S.Z.; Beyens, A.; Balic, Z.; Symoens, S.; Vandersteen, A.; Rideout, A.L.; Dickinson, J.; Callewaert, B.; Hubmacher, D. A novel ADAMTS17 variant that causes Weill-Marchesani syndrome 4 alters fibrillin-1 and collagen type I deposition in the extracellular matrix. Matrix Biol. 2019, 11, 11.
- Yi, H.; Zha, X.; Zhu, Y.; Lv, J.; Hu, S.; Kong, Y.; Wu, G.; Yang, Y.; He, Y. A novel nonsense mutation in ADAMTS17 caused autosomal recessive inheritance Weill-Marchesani syndrome from a Chinese family. J. Hum. Genet. 2019, 64, 681–687.
- Shah, M.H.; Bhat, V.; Shetty, J.S.; Kumar, A. Whole exome sequencing identifies a novel splice-site mutation in ADAMTS17 in an Indian family with Weill-Marchesani syndrome. Mol. Vis. 2014, 20, 790–796.
- Khan, A.O.; Aldahmesh, M.A.; Al-Ghadeer, H.; Mohamed, J.Y.; Alkuraya, F.S. Familial spherophakia with short stature caused by a novel homozygous ADAMTS17 mutation. Ophthalmic Genet. 2012, 33, 235–239.
- Morales, J.; Al-Sharif, L.; Khalil, D.S.; Shinwari, J.M.; Bavi, P.; Al-Mahrouqi, R.A.; Al-Rajhi, A.; Alkuraya, F.S.; Meyer, B.F.; Al Tassan, N. Homozygous mutations in ADAMTS10 and ADAMTS17 cause lenticular myopia, ectopia lentis, glaucoma, spherophakia, and short stature. Am. J. Hum. Genet. 2009, 85, 558–568.
- Kutz, W.E.; Wang, L.W.; Dagoneau, N.; Odrcic, K.J.; Cormier-Daire, V.; Traboulsi, E.I.; Apte, S.S. Functional analysis of an ADAMTS10 signal peptide mutation in Weill-Marchesani syndrome demonstrates a long-range effect on secretion of the full-length enzyme. Hum. Mutat. 2008, 29, 1425–1434.
- Dagoneau, N.; Benoist-Lasselin, C.; Huber, C.; Faivre, L.; Megarbane, A.; Alswaid, A.; Dollfus, H.; Alembik, Y.; Munnich, A.; Legeai-Mallet, L.; et al. ADAMTS10 mutations in autosomal recessive Weill-Marchesani syndrome. Am. J. Hum. Genet. 2004, 75, 801–806.
- Steinkellner, H.; Etzler, J.; Gogoll, L.; Neesen, J.; Stifter, E.; Brandau, O.; Laccone, F. Identification and molecular characterisation of a homozygous missense mutation in the ADAMTS10 gene in a patient with Weill-Marchesani syndrome. Eur. J. Hum. Genet. 2015, 23, 1186–1191.
- Li, J.; Jia, X.; Li, S.; Fang, S.; Guo, X. Mutation survey of candidate genes in 40 Chinese patients with congenital ectopia lentis. Mol. Vis. 2014, 20, 1017–1024.
- Stanton, H.; Melrose, J.; Little, C.B.; Fosang, A.J. Proteoglycan degradation by the ADAMTS family of proteinases. Biochim. Biophys. Acta 2011, 1812, 1616–1629.
- Mead, T.J.; Apte, S.S. ADAMTS proteins in human disorders. Matrix Biol. 2018, 71, 225–239.
- Apte, S.S. ADAMTS Proteins: Concepts, Challenges, and Prospects. Methods Mol. Biol. 2020, 2043, 1–12.
- Schnellmann, R.; Sack, R.; Hess, D.; Annis, D.S.; Mosher, D.F.; Apte, S.S.; Chiquet-Ehrismann, R. A Selective Extracellular Matrix Proteomics Approach Identifies Fibronectin Proteolysis by A Disintegrin-like and Metalloprotease Domain with Thrombospondin Type 1 Motifs (ADAMTS16) and Its Impact on Spheroid Morphogenesis. Mol. Cell. Proteom. 2018, 17, 1410–1425.
- Mularczyk, E.J.; Singh, M.; Godwin, A.R.F.; Galli, F.; Humphreys, N.; Adamson, A.D.; Mironov, A.; Cain, S.A.; Sengle, G.; Boot-Handford, R.P.; et al. ADAMTS10-mediated tissue disruption in Weill-Marchesani syndrome. Hum. Mol. Genet. 2018, 27, 3675–3687.
- Wang, L.W.; Kutz, W.E.; Mead, T.J.; Beene, L.C.; Singh, S.; Jenkins, M.W.; Reinhardt, D.P.; Apte, S.S. Adamts10 inactivation in mice leads to persistence of ocular microfibrils subsequent to reduced fibrillin-2 cleavage. Matrix Biol. 2019, 77, 117–128.
- Colige, A.; Monseur, C.; Crawley, J.T.B.; Santamaria, S.; de Groot, R. Proteomic discovery of substrates of the cardiovascular protease ADAMTS7. J. Biol. Chem. 2019, 294, 8037–8045.
- Hubmacher, D.; Schneider, M.; Berardinelli, S.J.; Takeuchi, H.; Willard, B.; Reinhardt, D.P.; Haltiwanger, R.S.; Apte, S.S. Unusual life cycle and impact on microfibril assembly of ADAMTS17, a secreted metalloprotease mutated in genetic eye disease. Sci. Rep. 2017, 7, 41871.
- Hubmacher, D.; Apte, S.S. ADAMTS proteins as modulators of microfibril formation and function. Matrix Biol. 2015, 47, 34–43.
- Ramirez, F.; Caescu, C.; Wondimu, E.; Galatioto, J. Marfan syndrome; A connective tissue disease at the crossroads of mechanotransduction, TGFbeta signaling and cell stemness. Matrix Biol. 2018, 71, 82–89.
- Sakai, L.Y.; Keene, D.R. Fibrillin protein pleiotropy: Acromelic dysplasias. Matrix Biol. 2019, 80, 6–13.
- Le Goff, C.; Cormier-Daire, V. The ADAMTS(L) family and human genetic disorders. Hum. Mol. Genet. 2011, 20, 163–167.
- Mead, T.J.; McCulloch, D.R.; Ho, J.C.; Du, Y.; Adams, S.M.; Birk, D.E.; Apte, S.S. The metalloproteinase-proteoglycans ADAMTS7 and ADAMTS12 provide an innate, tendon-specific protective mechanism against heterotopic ossification. JCI Insight 2018, 3, 92941.
- Enomoto, H.; Nelson, C.M.; Somerville, R.P.; Mielke, K.; Dixon, L.J.; Powell, K.; Apte, S.S. Cooperation of two ADAMTS metalloproteases in closure of the mouse palate identifies a requirement for versican proteolysis in regulating palatal mesenchyme proliferation. Development 2010, 137, 4029–4038.
- McCulloch, D.R.; Nelson, C.M.; Dixon, L.J.; Silver, D.L.; Wylie, J.D.; Lindner, V.; Sasaki, T.; Cooley, M.A.; Argraves, W.S.; Apte, S.S. ADAMTS metalloproteases generate active versican fragments that regulate interdigital web regression. Dev. Cell 2009, 17, 687–698.
- Brunet, F.G.; Fraser, F.W.; Binder, M.J.; Smith, A.D.; Kintakas, C.; Dancevic, C.M.; Ward, A.C.; McCulloch, D.R. The evolutionary conservation of the A Disintegrin-like and Metalloproteinase domain with Thrombospondin-1 motif metzincins across vertebrate species and their expression in teleost zebrafish. BMC Evol. Biol. 2015, 15, 22.
- Nicholson, A.C.; Malik, S.B.; Logsdon, J.M., Jr.; Van Meir, E.G. Functional evolution of ADAMTS genes: Evidence from analyses of phylogeny and gene organization. BMC Evol. Biol. 2005, 5, 11.
- Bevitt, D.J.; Li, Z.; Lindrop, J.L.; Barker, M.D.; Clarke, M.P.; McKie, N. Analysis of full length ADAMTS6 transcript reveals alternative splicing and a role for the 5′ untranslated region in translational control. Gene 2005, 359, 99–110.
- Somerville, R.P.; Jungers, K.A.; Apte, S.S. Discovery and characterization of a novel, widely expressed metalloprotease, ADAMTS10, and its proteolytic activation. J. Biol. Chem. 2004, 279, 51208–51217.
- Prins, B.P.; Mead, T.J.; Brody, J.A.; Sveinbjornsson, G.; Ntalla, I.; Bihlmeyer, N.A.; van den Berg, M.; Bork-Jensen, J.; Cappellani, S.; Van Duijvenboden, S.; et al. Exome-chip meta-analysis identifies novel loci associated with cardiac conduction, including ADAMTS6. Genome Biol. 2018, 19, 87.
- Koo, B.H.; Longpre, J.M.; Somerville, R.P.; Alexander, J.P.; Leduc, R.; Apte, S.S. Regulation of ADAMTS9 secretion and enzymatic activity by its propeptide. J. Biol. Chem. 2007, 282, 16146–16154.
- Kutz, W.E.; Wang, L.W.; Bader, H.L.; Majors, A.K.; Iwata, K.; Traboulsi, E.I.; Sakai, L.Y.; Keene, D.R.; Apte, S.S. ADAMTS10 protein interacts with fibrillin-1 and promotes its deposition in extracellular matrix of cultured fibroblasts. J. Biol. Chem. 2011, 286, 17156–17167.
- Holdener, B.C.; Haltiwanger, R.S. Protein O-fucosylation: Structure and function. Curr. Opin. Struct. Biol. 2019, 56, 78–86.
- Vasudevan, D.; Takeuchi, H.; Johar, S.S.; Majerus, E.; Haltiwanger, R.S. Peters plus syndrome mutations disrupt a noncanonical ER quality-control mechanism. Curr. Biol. 2015, 25, 286–295.
- Schneider, M.; Al-Shareffi, E.; Haltiwanger, R.S. Biological functions of fucose in mammals. Glycobiology 2017, 27, 601–618.
- Hurskainen, T.L.; Hirohata, S.; Seldin, M.F.; Apte, S.S. ADAM-TS5, ADAM-TS6, and ADAM-TS7, novel members of a new family of zinc metalloproteases. General features and genomic distribution of the ADAM-TS family. J. Biol. Chem. 1999, 274, 25555–25563.
- Cal, S.; Obaya, A.J.; Llamazares, M.; Garabaya, C.; Quesada, V.; Lopez-Otin, C. Cloning, expression analysis, and structural characterization of seven novel human ADAMTSs, a family of metalloproteinases with disintegrin and thrombospondin-1 domains. Gene 2002, 283, 49–62.
- Wunnemann, F.; Ta-Shma, A.; Preuss, C.; Leclerc, S.; van Vliet, P.P.; Oneglia, A.; Thibeault, M.; Nordquist, E.; Lincoln, J.; Scharfenberg, F.; et al. Loss of ADAMTS19 causes progressive non-syndromic heart valve disease. Nat. Genet. 2020, 52, 40–47.
