Your browser does not fully support modern features. Please upgrade for a smoother experience.
Please note this is a comparison between Version 3 by Camila Xu and Version 6 by Catherine Yang.
Acinetobacter baumannii is an emerging pathogen, and over the last three decades it has proven to be particularly difficult to treat by healthcare services. It is now regarded as a formidable infectious agent with a genetic setup for prompt development of resistance to most of the available antimicrobial agents. we provide a comprehensive updated overview of the available data about A. baumannii, the multi-drug resistant (MDR) phenotype spread, carbapenem-resistance, and the associated genetic resistance determinants in low-income countries (LIICs) since the beginning of the 21st century.
- A. baumannii
- low-income
- surveillance
- developing countries
- carbapenem-resistance
- MDR
- Syria
- Ethiopia
- COVID-19
1. Introduction
The ceaseless emergence of new pathogens and the constant mutations into pathogenic resistant strains are today’s humanity’s most tremendous threats. Carbapenem-resistant Acinetobacter baumannii (CRAP) is currently ranked first on the World Health Organization list of critical pathogens urgently in need of new antibiotic development. What is considered a low-virulence pathogen with reduced adhesion and invasion abilities has now surfaced as a hard to contain agent with the genetic setup for prompt development of resistance mechanisms to most of the available antimicrobial agents [1]. Acinetobacter was first isolated in 1911 from the soil [2]. It underwent taxonomic changes from its primary designated name Micrococcus calcoaceticus until it received the genus Acinetobacter due to its nonmotile nature. It is an aerobic Gram-negative, catalase-positive, oxidase-negative, nonfermenting coccobacillus belonging to the Moraxellacea family with a DNA G+C content ranging from 39% to 47% [2][3][2,3]. Being a non-lactose-fermenter with the ability to survive on low nutritional requirements enabled its germination in a wide range of environments. Moreover, its ability to form biofilms, resist dissection, and persist for extended periods of time on surfaces, especially in hospital settings, led to multiple outbreaks [4][5][6][4,5,6].
A. baumannii causes too many types of infections including ventilator-associated pneumonia, meningitis, urinary tract infection, bone/wound infections, and the most serious type is the bacteremia. What makes A. baumannii more critical is that the mortality rates associated with it could reach up to 35% and escalate to 43% in intensive care units (ICUs) [7]. The global prevalence of A. baumannii, its widespread antibiotic resistance, and most recently, the unforeseen emergence and spread of clinical isolates resistant to colistin, the last resort in our current antimicrobial arsenal [8], all this necessitates a more prominent understanding of this challenging organism. The emphasis of the current review will be on the economically disadvantaged populations residing in low-income countries (LICs). It is challenging to portray an accurate picture of the spread of A. baumannii and its antibiotic susceptibility profile in different settings in this part of the world. Notably, there is a great disparity in the economic resources available in low-income countries and the rest of the world. Accordingly, this could be reflected on the detection capabilities and the level of details and sophistication of the reports about A. baumannii outbreaks.
2. Methodology
The scientific literature in the three databases PubMed, Scopus, and Web of Science was screened in this order. The search terms included “baumannii” and the name of each of the twenty-nine countries included in the 2020/2021 World Bank classification and categorized as LICs [9]. The date limit was set to start from January 2000 to December 2020. In the case of a country name with any known variations, the different versions of the name were included in separate searches. No limit was imposed on the language with which the article was published. Accordingly, the studies included English, French, and Spanish articles. Both original reports and case reports were included.
A total of 493 hits were obtained from the three databases, then the following inclusion criteria were applied to the retrieved studies to select the relevant ones. The study had to include the detection of A. baumannii isolate or its DNA either in a medical or an environmental setting. The isolate or the DNA should have been detected in a setting located within one of the 29 LICs included in the study. Furthermore, studies of isolates confirmed to be isolated from individuals who are nationals of the targeted LICs but were transferred to be treated in other countries, either for war conflicts or political instabilities, were also included. Studies focused on performing research on standard strains to test the efficacy of medications or natural products collected from the targeted LICs were excluded. Finally, studies with just one, or more, of the authors who have an affiliation associated with one of the 29 LICs, but without including isolates or nucleic acids that fit the above criteria, were excluded. Repeated studies among the database were excluded considering first the ones detected in PubMed, then Scopus, and finally Web of Science. A summary of the search methodology adopted in the current study is presented in Figure 1.
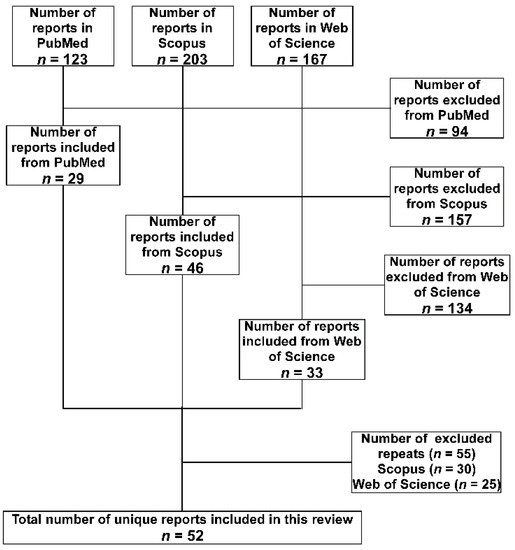
Figure 1. Flow diagram of the search strategy and selection of articles adopted in the current review. The databases were searched in the order (PubMed, Scopus, and Web of Science). Accordingly, articles found in the subsequent database(s) were considered a repeat and excluded.
The total number of included studies after applying the exclusion criteria was 52. They were analyzed for the method of identification and any phenotypic or genotypic characterizations. Special attention was paid to the antibiotics susceptibility profiles and the detection of the multi-drug-resistance (MDR) which was defined as being resistant to at least three antibiotics belonging to different classes. If the study included testing of resistance to carbapenems, the percentage of resistance was also noted.
3. A. baumannii in the LICs Distributed over the Different Geographical Regions
The 29 LICs included in the World Bank classification are distributed over six geographical regions; Sub-Saharan Africa (n = 23), the Middle East and North Africa (n = 2), South Asia (n = 1), Latin America and The Caribbean (n = 1), Europe and Central Asia (n = 1), and East Asia and Pacific (n = 1). Accordingly, this review will address the studies that originated from each of these six regions. In particular, an overview of the incidences of A. baumannii infections, the MDR phenotype spread, the presence of carbapenem-resistance, the associated genetic resistance determinants if available, and any phenotypic or genotypic characterization performed.
3.1. Sub-Saharan Africa
The Sub-Saharan Africa region included a total of 23 countries of the LICs. The reports about A. baumannii in these countries in the searched databases are summarized below. They are arranged in a descending order according to the number of reports related to each country.
Ethiopia is ranked the first among this group in the number of reports about A. baumannii infections with a total of 10 studies. The first report of the detection of A. baumannii in this country came in 2012, and it was isolated from human head and body lice collected from healthy individuals [10]. The genotypic analysis of these isolates showed that these particularly sensitive strains harbored several hundred insertion sequence elements that served in their genome reduction (disruptions of genes and simple loss of DNA). Additionally, it was pointed out that they have low catabolic capacities compared to the human MDR A. baumannii isolates. This signifies the adaptation of this bacterium to the louse environment [10]. In the same year, A. baumannii was reported in human infections for the first time from the ulcers of leprosy patients [11]. Yet, only 20% of these isolates showed MDR phenotype. It took almost five years to detect MDR A. baumannii that are carbapenem-resistant harboring the blaNDM-1 resistance determinant. However, susceptibility testing indicated that they remain susceptible to both polymyxin and amikacin [12]. Genome sequencing confirmed that the isolates carried the blaNDM-1; however, they are distinct from the outbreak strains reported from neighboring countries like Kenya. This argued against the regional spread of the blaNDM-1-positive organism but rather implied the independent environmental dependent generation of the resistance [12]. Moreover, MDR A. baumannii was isolated from a hospital’s air setup implicating this troublesome pathogen in the widespread colonization of the hospital’s setting [13]. The prevalence of MDR and Extended Spectrum Beta-Lactamase (ESBL) producers are reaching alarming rates from the isolated specimens in Ethiopian hospitals [14][15][16][14,15,16]. In another study, 33% of the isolates from nosocomial infections were meropenem-resistant [17]. The situation got more complicated in other instances where the isolates turned out to be extensively drug resistant (XDR); resistant to at least one agent in all but two or fewer antimicrobial categories, pan-drug-resistant (PDR); resistant to all antibiotic classes [18][19][18,19].
With only five studies dealing with A. baumannii infections, Madagascar came next in the list of LIC nations in this geographical sub-region. The first reports about A. baumannii outbreaks in Madagascar hospitals came in 2010 [20][21][20,21]. The isolated strains showed up to 44% carbapenem-resistance, and harboring genes encoding the β-lactamases AmpC, OXA23, and OXA51 [21]. Interestingly, the emergence of these strains in Madagascar preceded the availability of carbapenems in this country’s hospitals. Nosocomial infections between 2006 and 2013 were caused by carbapenem-resistant A. baumannii carrying only the blaOXA23 and blaOXA51 genes as the only carbapenemase-producing genes [21]. During the period between 2011 and 2013, MDR A. baumannii started to show up as one of the causative agents of urinary tract infections (UTI) in Madagascar; however, they were still carbapenem-sensitive [22]. On the other hand, reports about clinical samples collected between 2013 and 2016 indicated the predominance of Sequence Type 2 (ST2) and the acquisition of carbapenemases blaOXA24 and blaOXA58 for the first time since the first outbreak of A. baumannii in Madagascar [23]. In 2019, A. baumannii were detected in human head lice, yet they were negative for blaOXA23, blaOXA24, blaIMP, and blaVIM [24].
Four reports have come from Uganda in the last 21 years reporting the detection of A. baumannii in the hospitals of this country. A recent report regarded A. baumannii as a minor cause of sepsis (~1%) in a multisite study [25]. An earlier report, examining the prevalence of A. baumannii and its rates of resistance between 2007 and 2009, showed that it was responsible for about 3% of the infections with 31% of these isolates carbapenem-resistant [26]. What was alarming in this report is that A. baumannii represented 14% of the environmental samples in the hospital and 55% were carbapenem-resistant. The β-lactamases encoding genes detected in this study included; blaOXA23-like, blaOXA24-like, blaOXA58-like, and blaVIM-like [26]. Focusing on the isolates from the same hospital, cluster analysis using repetitive element sequence-based polymerase chain reaction (Rep-PCR) fingerprinting indicated a high level of genetic diversity among the isolates [27]. Yet, certain MDR isolates from the environment and patients were clustered, indicating possible environmental transmission of these strains to the patients in the hospital. Ten years later and in a follow-up study published in 2019, the same hospital reported a significant decrease in carbapenem resistance prevalence in the isolated Acinetobacter reaching only 2.7% [28]. However, all the carbapenemase-producing isolates were MDR and the blaVIM was the most prevalent carbapenemase-encoding gene. An explanation for this decrease was not discussed by the authors, however a possible one could be the very limited number of the A. baumannii strains investigated in the earlier study as compared to the very large number of isolates in the later study (more than 1000). Accordingly, the high rates in the earlier study could be an over-estimation of the situation then.
Only two reports about A. baumannii infections in each of Burkina Faso, Democratic Republic of the Congo, Malawi, Mozambique, and Sudan were published in the last 21 years. For Burkina Faso, the first report came in 2016 about oral infections and they detected 3 A. baumannii strains out of 125 clinical samples [29]. All three were MDR and ESBL-positive and the β-lactamases encoding genes identified were blaTEM and blaCTX-M. In a more recent study, few A. baumannii strains were isolated from a hospital there and they belonged to sequence type 2, or what is known as international clone II which has a high distribution worldwide [30][31][30,31]. On the other hand, in the Democratic Republic of the Congo (formally known as Zaire), a single report about nosocomial infections caused by A. baumannii came out in 2017 [32]. The isolates were not MDR and remained sensitive to both cefotaxime and gentamycin. Another report came from the same country in 2019; however, this time A. baumannii was isolated from human body lice but without determination of their antibiotic susceptibility patterns [33]. Regarding Malawi, a study published in 2012 reported the isolation of only one A. baumannii strain from a blood-stream infection of an HIV patient [34]. Following the incidence of bloodstream infections in children ≤5 years in a central hospital in Malawi in 20 years from 1998 to 2017, A. baumannii increased 10-fold from 0.2% (2003–2007) to 2.2% (2013–2017), and the isolates showed MDR phenotypes [35]. In Mozambique, a case of a hospitalized fatal pneumonia caused by an MDR A. baumannii strain belonging to the international clone II took place in 2014, yet it was not reported until 2018 as a case study [36]. The same case was reported earlier by another study in 2016 dealing with evaluating minimally invasive autopsy procedures to identify the infectious cause of death in the same hospital [37]. Finally, for Sudan, which has moved recently to the LIC group, the first report of A. baumannii was published in 2019 about the whole genome sequencing of an A. baumannii strain isolated from a hospitalized patient in the capital, Khartoum [38]. The sequence analyses indicated that the strain belonged to sequence type 164. In 2020, a survey of antimicrobial resistance in a hospital in Khartoum indicated that A. baumannii was the most frequent carbapenemase-producing organism (89% of the isolates) and they demonstrated higher resistance rates (100%) for cephalosporins and trimethoprim/sulfamethoxazole [39].
Only two studies about A. baumannii in Rwanda were published with the first one in 2004 and reported ten strains, from three different genotypes, detected in human body louse [40]. However, a very recent report came in 2020 and indicated the detection of A. baumannii in flies from a tertiary hospital [41]. Only one strain of A. baumannii was detected in this setting, yet its genome carried genes encoding virulence factors that are known to be associated with serum survival and invasion [41]. Only one study about A. baumannii in Burundi was published in 2004, which is the same report as the one mentioned above from the neighboring Rwanda. However, it only reported three strains detected in human body louse from the same genotype [40]. On the other hand, only one study reported A. baumannii infection in Mali, where it represented only 10% of the studied cases of pregnant women with cervicofacial cellulitis [42]. The isolated strain remained sensitive to amoxicillin/clavulanic acid. While in Sierra Leone, A. baumannii was responsible for 14 out of 164 nosocomial infections in an urban tertiary hospital [43]. The isolated strains were MDR and positive for ESBL, yet the carbapenem-resistance was as low as 10%.
There is only one report that originated from Somalia and was published online in November 2020 that describes the detection of A. baumannii in a hospital in Somalia [44]. In this report, A. baumannii was the most prevalent pathogen that belonged to MDR and XDR patterns in 69.1% of the samples with isolates showing 100% resistance rates against beta-lactam, cephalosporins, fluoroquinolones, and carbapenems. Finally, only one report about A. baumannii in head lice and Niger was published in 2018 [45]. In this case, A. baumannii were not detected in head lice from people residing in Niger; however, they were refugees from this country in neighboring Algeria.
No reports were published in the searched three databases for A. baumannii infections in the following countries since the beginning of the 21st century; Central African Republic, Chad, Eritrea, Gambia, Guinea, Guinea-Bissau, Liberia, South Sudan, and Togo. A graphical representation of the distribution of the included studies in the LICs in the Sub-Saharan region is presented in Figure 2.
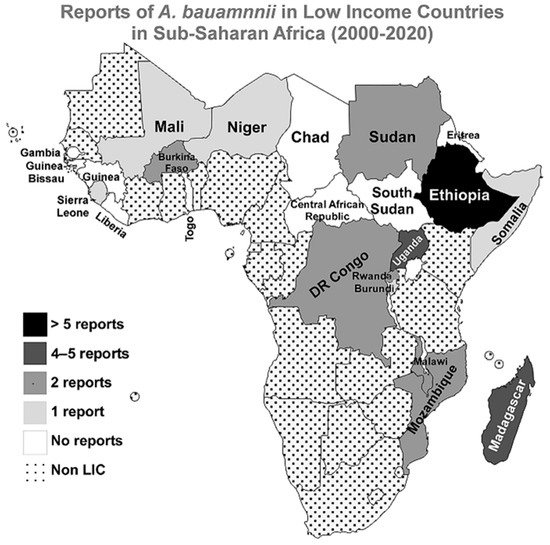
Figure 2. Reports of A. baumannii infections in LICs in Sub-Saharan Africa from 2000 to 2020. A geographical map of the Sub-Saharan Africa region highlighting the numbers of reports about A. baumannii infections in the 21 first years of the 21st century. The map was generated using the MapChart web tool (https://mapchart.net/, accessed on 21 June 2021).
3.2. Middle East and North Africa
Only two countries belonging to the LIC group are located within this geographical region: Syria and Yemen. Despite being ravaged by wars for many years within the last two decades, the linkage between Syria and A. baumannii infections has a very good share in the reports covered in this review; 10 studies. However, most of these reports were including Syrian patients who were treated in hospitals outside Syria especially in neighboring countries. The first report of A. baumannii in Syria was in 2012 and the study was conducted in a Syrian city showing high rates of MDR strains and the carbapenem-resistance was around 65% among them [46]. Yet, these strains remained susceptible to colistin. All the later reports dealt with Syrians treated in other countries. Wounded Syrians treated in Jordan showed up to 80% carbapenem-resistance among MDR A. baumannii infections between August 2011 and March 2013 [47]. Syrian bone-wounded civilians treated in Jordan and infected with A. baumannii were also among those reported in another study in 2017 together with wounded civilians from two other wars [48]. In another study analyzing data in Jordanian hospitals over a decade (2006–2016), 140 Syrians who had bone war wounds were included. This study showed that MDR A. baumannii was among the causative agents of these infections and the carbapenem-resistance level among these isolates was as high as 66.7% [49]. Moreover, Syrians treated in Israeli hospitals showed carriage of MDR A. baumannii [50]. Concerns about the exporting of NDM-1-producing A. baumannii strains and/or their genetic determinants from Syria was highlighted in a report from Turkey, where a Syrian refugee was linked to the introduction of the MDR and carbapenem-resistant strains to Turkey, and the strain belonged to sequence type 85 [51]. More recently in 2019, a Turkish study reported the detection of A. baumannii (~5%) among the causative agents of infection in Syrian war-injured patients [52]. As in Turkey, Lebanon too linked the introduction of MDR NDM-1 producing A. baumannii strains to Syrians injured in the war and transferred to Lebanese hospitals for treatment [53]. Similar to the Turkish case, the isolates belonged to sequence type 85 showing that Syria could constitute a reservoir for these strains. Two more studies from Lebanon reported the detection of MDR A. baumannii strains originating in Syria with carbapenem-resistance rates ranging from 74% to 100% [54][55][54,55].
The second country of the LIC group in the Middle East and North Africa is Yemen, where the situation is not greatly different from Syria as the country was inflicted with war for many years during the last 21 years. Only one study was reported from inside a hospital in Yemen, where three MDR carbapenem-resistant isolates were investigated [56]. Genetic analyses indicated that they harbor the blaOXA23-like, 16S rRNA methylase armA, and the acetyltransferase aac(6′)-Ib genes. The three strains belonged to sequence type 2 (ST2) and they were still susceptible to colistin. Yemen was linked to A. baumannii infections in another study but not for people receiving healthcare inside the country. In these cases, they were bones war-wounded patients hospitalized in Jordan and when detected, A. baumannii was carbapenem-resistant [49].
3.3. South Asia
Only Afghanistan is located in this geographical area and belongs to the LIC group. A single report came out in 2011 about Afghani patients treated in a deployed American Military Hospital that reported the recovery of Acinetobacter spp., including A. baumannii, from both patients and the hospital environment. Many of the isolated strains had high rates of antimicrobial resistance [57].
3.4. Latin America and the Caribbean
Only Haiti is included in this sub-category and the literature contained few reports about A. baumannii infection among Haitian people. The first report dealt with A. baumannii isolates collected from patients who had been injured in the Haiti earthquake in 2010 and they were blaCTX-M-15-positive [58]. Another report came in 2012 dealt also with the 2010 earthquake disaster and reported one case infected with MDR A. baumannii among the 14 wound-infection patients who had purulent discharges included in this study [59]. A third report dealt with infections burn units in Haiti. In this unit, A. baumannii was responsible for 15% of the reported infections, the A. baumannii associated with blood infection among patients who did not survive to discharge was resistant to aminoglycosides, fluoroquinolones, third-generation cephalosporins, and also carbapenems [60]. In a surveillance study in an obstetrics emergency hospital and neonatal care unit conducted during 2016, A. baumannii was isolated from two women’s rectal swabs [61]. In a recent metagenomic study to assess the quality of surface water in Haiti, A. baumannii was estimated to represent 5% of Acinetobacter species and 1% of total relative abundance [62]. These findings indicate that this pathogen is present in the water and could potentially be responsible for a proportion of the human infections in this country.
3.5. Europe and Central Asia
Only Tajikistan is located in this area; however, no reports about A. baumannii in this country were found in our search of the three respective databases.
3.6. East Asia and Pacific
Democratic People’s Republic of Korea (also known as North Korea) is the only low-income country that belongs to this geographical area. There are no records in PubMed directly reporting the isolation of A. baumannii in North Korea. Although, in the review by Zarrilli and co-workers [63], they citied another study by Yum and colleagues [64]. However, careful examination of the latter study indicated that this was a report about isolates from South Korea rather than its northern counterpart. Accordingly, it is safe to say that there are no reported studies about the detection of the A. baumannii in North Korea in the included databases in the last 21 years.
A summary of all the analyzed studies is presented in Table 1.
Table 1. A summary of the studies about A. baumannii in LICs from 2000 to 2020.
| Country | Study | Isolates ( | n | ) | MDR % * | CRAB% | Isolates Characterization | References | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sub-Saharan Africa | |||||||||||||||||||||||
| Ethiopia | Kempf et al., 2012 | 40 | NA | NA | rpoB | and | recA | sequencing for genotyping | [10] | ||||||||||||||
| Lema et al., 2012 | 5 | ≥20% | NA | AST with KB | [11] | ||||||||||||||||||
| Pritsch et al., 2017 | 3 | 100% | 100% | AST with KB and VITEK 2, CT102 Micro-Array, real-time PCR, WGS, MLST, and detection of the | blaNDM-1 | [12] | |||||||||||||||||
| Solomon et al., 2017 | 43 | 81% | 37% | AST with KB and phenotypic detection of ESBLs and MBLs | [13] | ||||||||||||||||||
| Bitew et al., 2017 | 2 | 100% | NA | Identification and AST with VITEK 2 | [15] | ||||||||||||||||||
| Demoz et al., 2018 | 1 | 100% | 100% | AST with KB | [18] | ||||||||||||||||||
| Gashaw et al., 2018 | 2 | 50% XDR and 50% PDR | 100% | AST with KB and phenotypic detection of ESBLs and AmpC | [19] | ||||||||||||||||||
| Moges et al., 2019 | 15 | ≥63% | Yes | AST with KB and phenotypic detection of ESBLs and carbapenemases | [14] | ||||||||||||||||||
| Admas et al., 2020 | 6 | 100% | NA | Identification and AST with VITEK 2 | [16] | ||||||||||||||||||
| Motbainor et al., 2020 | 9 | 100% | 33% | Identification with VITEK 2 and AST with KB | [17] | ||||||||||||||||||
| Madagascar | Randrianirina et al., 2010 | 50 | ≥44% | 44% | AST with KB and phenotypic detection of ESBLs | [20] | |||||||||||||||||
| Andriamanantena et al., 2010 | 53 | 100% | 100% | AST with KB and MIC determination, phenotypic detection of carbapenemases, ReP-PCR for genotyping and PCR for detection of; | blaAmpC | , | blaoxa51 | , | blaoxa23 | , | blaoxa24 | , | blaVIM | , | blaIMP | , and | isAba-1 | [21] | |||||
| Rasamiravaka et al., 2015 | 10 | ≥50% | 0% | AST with KB | [22] | ||||||||||||||||||
| Tchuinte et al., 2019 | 15 | 100% | 100% | MALDI-TOF MS for identification, AST with KB and MIC determination, WGS, MLST for genotyping and WGS detecting; | blaoxa51 | , | blaoxa23 | , | blaoxa24 | , | blaoxa58 | , and | isAba-1 | [23] | |||||||||
| Eremeeva et al., 2019 | 14 | NA | NA | TaqMan PCR of the | rpoB | for identification, and PCR for detecting: | blaoxa51-like | , | blaoxa23 | , | blaoxa24 | , | blaVIM | , and | blaIMP | [24] | |||||||
| Uganda | Kateete et al., 2016 | 40 | 60% | 38% | AST with Phoenix Automated Microbiology System, PCR for: | blaoxa51-like | , | blaoxa51 | , | blaoxa23 | , | blaoxa24 | , | blaoxa58 | , | blaVIM | , | blaSPM | , and | blaIMP | [26] | ||
| Kateete et al., 2017 | 20 | 40% | 35% | AST with MIC determination, PAMS, Rep-PCR for genotyping and phenotypic detection of ESBLs and AmpC | [27] | ||||||||||||||||||
| Moore et al., 2019 | 3 | NA | NA | qPCR TAC | [25] | ||||||||||||||||||
| Aruhomukama et al., 2019 | 1077 | 3% | 3% | AST with KB, PCR for detecting: | blaoxa23 | , | blaoxa24 | , | blaoxa58 | , | blaVIM | , | blaSPM | , | blaKPC | , and | blaIMP | , phenotypic detection of carbapenemases, and conjugation to show transferability of | blaVIM | . | [28] | ||
| Burkina Faso | Kaboré et al., 2016 | 3 | 100% | NA | AST with KB and phenotypic detection of ESBLs | [29] | |||||||||||||||||
| Sanou et al., 2021 | # | 5 | 100% | 60% | MALDI-TOF MS for identification, AST with KB and MIC determination, phenotypic detection of ESBLs, PCR and sequencing of multiple resistance genes including; | blaoxa1-like | , | blaoxa48-like | , | blaNDM | , | blaVIM | , | blaSPM | , | blaKPC | , | blaCTX-M | , and | blaIMP | , and MLST for genotyping. | [30] | |
| DR of the Congo | Lukuke et al., 2017 | 2 | 0% | NA | API for identification and AST with KB | [32] | |||||||||||||||||
| Koyo et al., 2019 | 15 | NA | NA | qPCR and phylogenetic analysis using the | rpoB | gene | [33] | ||||||||||||||||
| Malawi | Bedell et al., 2012 | 1 | NA | NA | Identification with standard diagnostic techniques | [34] | |||||||||||||||||
| Iroh Tam et al., 2019 | 84 | ≥44% | NA | API for identification, AST with KB, and phenotypic detection of ESBLs | [35] | ||||||||||||||||||
| Mozambique | Martínez et al., 2016 | 1 | NA | NA | 16S rRNA PCR and MALDI-TOF MS for identification | [37] | |||||||||||||||||
| Hurtado et al., 2019 | 1 | 100% | 0% | 16S rRNA for identification and AST with KB | [36] | ||||||||||||||||||
| Sudan | Mohamed et al., 2019 | 1 | NA | NA | API for identification followed by WGS | [38] | |||||||||||||||||
| Dirar et al., 2020 | 12 | ≥83% | 89% | Identification with PAMS, AST with KB and phenotypic detection of ESBLs and carbapenemases. | [39] | ||||||||||||||||||
| Rwanda | La Scola and Raoult 2004 | 10 | NA | NA | API for identification and | recA | genotyping | [40] | |||||||||||||||
| Heiden et al., 2020 | 1 | 100% | 0% | MALDI-TOF MS for identification, AST with VITEK 2, phenotypic detection of ESBLs and carbapenemases, and WGS | [41] | ||||||||||||||||||
| Burundi | La Scola and Raoult 2004 | 3 | NA | NA | API for identification and | recA | genotyping | [40] | |||||||||||||||
| Mali | Doumbia-Singare et al., 2014 | 1 | NA | NA | Not mentioned | [42] | |||||||||||||||||
| Sierra Leone | Lakoh et al., 2020 | 14 | ≥40% | 10% | Identification and AST with VITEK 2 | [43] | |||||||||||||||||
| Somalia | Mohamed et al., 2020 | 7 | 100% | 100% | AST with KB | [44] | |||||||||||||||||
| Niger | Louni et al., 2018 | 29 | NA | NA | qPCR and | rpoB | PCR for identification and phylogenetic analysis | [45] | |||||||||||||||
| Central African Republic | No Reports | ||||||||||||||||||||||
| Chad | No Reports | ||||||||||||||||||||||
| Eritrea | No Reports | ||||||||||||||||||||||
| Gambia | No Reports | ||||||||||||||||||||||
| Guinea | No Reports | ||||||||||||||||||||||
| Guinea-Bissau | No Reports | ||||||||||||||||||||||
| Liberia | No Reports | ||||||||||||||||||||||
| South Sudan | No Reports | ||||||||||||||||||||||
| Togo | No Reports | ||||||||||||||||||||||
| Middle East and North Africa | |||||||||||||||||||||||
| Syria | Hamzeh et al., 2012 | 260 | ≥65% | 65% | Identification and AST with PAMS | [46] | |||||||||||||||||
| Teicher et al., 2014 | 6 | 100% | 80% | API for identification and AST with MicroScan Walk-Away System | [47] | ||||||||||||||||||
| Peretz et al., 2014 | 5 | 100% | NA | Not mentioned | [50] | ||||||||||||||||||
| Rafei et al., 2014 | 4 | 100% | 100% | Identification with | rpoB | sequencing and | blaoxa51 | , PCR, AST with KB and Etest, PCR for: | blaoxa23-like | , | blaoxa24-like | , | blaoxa58-like | , and | blaNDM | , and PFGE for genotyping | [53] | ||||||
| Heydari et al., 2015 | 1 | 100% | 100% | Identification and AST with VITEK 2, phenotypic detection of ESBLs and carbapenemases, PCR for the | blaNDM | , PFGE and MLST for typing | [51] | ||||||||||||||||
| Rafei et al., 2015 | 59 | Yes | 74% | Identification with MALDI-TOF MS, | rpoB | sequencing and | blaoxa51 | PCR, AST with KB and Etest, PCR for detecting: | blaoxa23 | , | blaoxa24 | , | blaoxa58 | , | blaNDM-1 | , | blaVIM | , | blaoxa143 | , and | blaIMP | , and MLST for typing | [54] |
| Herard and Fakhri 2017 | 38 | NA | NA | Not mentioned | [48] | ||||||||||||||||||
| Salloum et al., 2018 | 2 | 100% | 100% | AST with KB and Etest, PCR for | blaoxa58 | and | blaNDM | , plasmid typing with PBRT, MLST, and WGS | [55] | ||||||||||||||
| Fily et al., 2019 | 6 | NA | 67% | AST with KB | [49] | ||||||||||||||||||
| Hasde et al., 2019 | 5 | NA | NA | Not mentioned | [52] | ||||||||||||||||||
| Yemen | Bakour et al., 2014 | 3 | 100% | 100% | API and MALDI-TOF MS for identification, AST with KB and E-test, phenotypic detection of carbapenemases, PCR detection of: | blaoxa23 | , | blaoxa24 | , | blaoxa58 | , | blaNDM | , | blaVIM | , | blaSIM | , | blaoxa48-like | , | blaIMP | and others, and MLST | [56] | |
| Fily et al., 2019 | 1 | NA | 100% | AST with KB | [49] | ||||||||||||||||||
| South Asia | |||||||||||||||||||||||
| Afghanistan | Sutter et al., 2011 | 57 | ¥ | ≥75% | 76% | Identification and AST with MicroScan autoSCAN-4 | [57] | ||||||||||||||||
| Latin America and The Caribbean | |||||||||||||||||||||||
| Haiti | Potron et al., 2011 | 3 | 66.7% | 0% | API and 16sRNA for identification, AST with KB and E-test, phenotypic detection of ESBLs, PCR for detection of: | blaTEM | , | blaSHV | , | blaPER-1 | , | blaVEB-1 | , | blaGES-1 | , and | blaCTX-M | [58] | ||||||
| Marra et al., 2012 | 1 | 100% | 0% | Identification and AST with VITEK 2 | [59] | ||||||||||||||||||
| Murphy et al., 2016 | 4 | ≥25% | 25% | AST but the method was not indicated | [60] | ||||||||||||||||||
| Chaintarli et al., 2018 | 2 | 0% | 0% | Identification and AST with VITEK 2 and phenotypic detection of ESBLs | [61] | ||||||||||||||||||
| Roy et al., 2018 | 0 | ϕ | NA | NA | Metagenomic analyses of water samples | [62] | |||||||||||||||||
| Europe and Central Asia | |||||||||||||||||||||||
| Tajikistan | No Reports | ||||||||||||||||||||||
| East Asia and Pacific | |||||||||||||||||||||||
| Democratic People’s Republic of Korea | No Reports | ||||||||||||||||||||||
Abbreviations: API: Analytical Profile Index, AST: Antibiotic Susceptibility testing, CRAB: Carbapenem-resistant A. baumannii, KB: Kirby-Bauer disc diffusion method, MALDI-TOF MS: Matrix-Assisted Laser Desorption/Ionization Time-of-Flight Mass Spectrometry, MDR: Multi-drug resistant, MLST: Multilocus sequence typing, NA: not available, PAMS; Phoenix Automated Microbiology System, PDR: Pan-drug-resistant, PBRT: PCR-based replicon typing, PCR: Polymerase chain reaction, PFGE: Pulse Field Gel Electrophoresis, rep-PCR: Repetitive element sequence-based PCR, WGS: Whole Genome Shotgun, TAC: TaqMan Array Card, XDR: Extensively drug-resistant. * When the MDR% is not directly mentioned, it was presented as ≥lowest % of resistance among the tested antibiotics classes. All %s were approximated to the nearest whole number. # The study was originally published in Jun 2020, then appeared in its final version in Jan 2021. ¥ Acinetobacter spp. Including A. baumannii. ϕ No isolates were obtained but A. baumannii DNA was detected in the samples.
Abbreviations: API: Analytical Profile Index, AST: Antibiotic Susceptibility testing, CRAB: Carbapenem-resistant A. baumannii, KB: Kirby-Bauer disc diffusion method, MALDI-TOF MS: Matrix-Assisted Laser Desorption/Ionization Time-of-Flight Mass Spectrometry, MDR: Multi-drug resistant, MLST: Multilocus sequence typing, NA: not available, PAMS; Phoenix Automated Microbiology System, PDR: Pan-drug-resistant, PBRT: PCR-based replicon typing, PCR: Polymerase chain reaction, PFGE: Pulse Field Gel Electrophoresis, rep-PCR: Repetitive element sequence-based PCR, WGS: Whole Genome Shotgun, TAC: TaqMan Array Card, XDR: Extensively drug-resistant. * When the MDR% is not directly mentioned, it was presented as ≥lowest % of resistance among the tested antibiotics classes. All %s were approximated to the nearest whole number. # The study was originally published in Jun 2020, then appeared in its final version in Jan 2021. ¥ Acinetobacter spp. Including A. baumannii. ϕ No isolates were obtained but A. baumannii DNA was detected in the samples.
