Bilaterian animals operate the clusters of Hox genes through a rich repertoire of diverse mechanisms, including, due to a large set of various non-coding RNAs. Long non-coding RNAs (lncRNAs), which are transcribed from the sense (coding) DNA strands of Hox clusters, control the work of Hox genes in the cis and trans position, are involved in the establishment and maintenance of the epigenetic code of Hox loci, and can even serve as a source of regulatory peptides. which switch cellular energy metabolism. All antisense lncRNAs in human Hox clusters are therapeutic targets for malignant tumors, and their careful study has profound practical meaning.
- long noncoding RNAs
- lncRNAs
- antisense ncRNAs
- Hox genes
- Hox clusters
1. Introduction
-
Their fundamental role in the ground plan formation (this excludes the loss of Hox genes in most bilaterian animals);
-
The simplicity of DNA-consensus for the binding of Hox homeodomain and the ability of Hox genes to form the dimers with cofactors to make this consensus more complicated;
-
The complex regulation of their transcription.
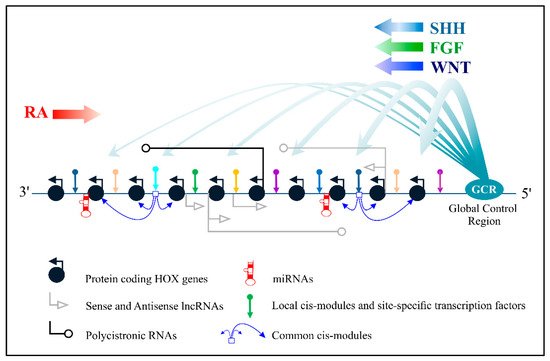
2. Antisense LncRNAs in Mammalian Hox Clusters
| LncRNA | Length (nt) | Type of LncRNA | Position in the Hox Cluster | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Targets | Orthologs | Discovered in | Refs | ||||||||
| HOXA Cluster | |||||||||||
| HOTAIRM1 | Control of the cell cycle in the myeloid cell lineage; control of the differentiation of granulocytes; control of neuronal differentiation timing; control of osteogenesis in dental follicle stem cells | Serve as protein scaffolds; Enhancer; Sponges big set miRNAs |
Nucleus Cytoplasm | HOXA cluster; NEUROGENIN 2; miR-196b; miR-125b | Chordata | 2009 | [36][39][40][41][42][43][44] | HOTAIRM1 | 4000; 1052; 783 | Linc | |
| HOXA-AS2 | Promotion of proliferation, migration and invasion in many types of tumors; regulation EMT; negative regulates endothelium inflammation | Serve as protein scaffolds; HOXA1→HOXA2 |
|||||||||
| Sponges big set miRNAs | Nucleus | Cytoplasm |
c-MYC; EGFR; Bax/TRAIL; EZH2/LSD1; PBX3; NF-kB; miR-373 | Primates | 2013 | [45][46][47][48][49] | HOXA-AS2 | 1048 | Linc or NAT | HOXA3→H | |
| HOXA-AS3 | OXA4 | ||||||||||
| Control of cell cycle, proliferation, migration and apoptosis in many types of cancer cells; positive regulation of endothelium inflammation; activation the MEK/ERK Signaling Pathway | Stabilization of HOXA6 mRNA, sponges miR-29c and mir-455-5p | Nucleus Cytoplasm |
HOXA6; NF-kB; miR-29c | Homo sapiens | 2017 | [50][51][52][53][54] | HOXA-AS3 | 3918; 3992 | Linc or NAT | HOXA4→A5,→A6→HOXA7 | |
| HOXA10-AS | |||||||||||
| HOXA10-AS | 1161 | Linc or NAT | HOXA9→HOXA10 | ||||||||
| Cell cycle and apoptosis control in glioma, lung adenocarcinoma (LAD), oral cancer and acute myeloid leukemia (AML) cells | ? | Cytoplasm | HOXA10; Wnt pathway; NF-kB |
Birds Mammals |
2018 | [55][56][57][58] | |||||
| HOXA11-AS | Control of the menstrual cycle; cell cycle, proliferation, migration and apoptosis control in many types of cancer cells | Serve as protein scaffolds; Sponges big set miRNAs |
Nucleus Cytoplasm |
HOXA11; TGF-b pathway, LATS1; CyclinD1; CyclinE; CDK4; CDK2 | Rodents Primates Bamboo shark |
2002 | [59] | HOXA11-AS | 1628 | Linc or NAT | HOXA11→HOXA13 |
| HOTTIP | Control of 5′HOXA genes’ transcription during development. Participation in pathogenesis of almost all types of cancer. | Activates HOXA genes through recruiting of WDR5 и MLL | Nucleus | HOXA7-HOXA13; LSD1; EZH2; IL-6; miR-30b | Rodents Primates Bamboo shark |
2011 | [60][61] | HOTTIP | 4665 | Linc or NAT | HOXA13→EVX1 |
| HOXB-AS1 | Glioblastoma and endometrial carcinoma and multiple myeloma (MM) promotion | ILF3-mediated activation of HOXB3 and HOXB3 transcription; stabilization of their mRNAs; stabilization of FUT4 mRNA | Nucleus Cytoplasm |
HOXB2; HOXB3; Wnt pathway; FUT4; miR-186-5p; miR-149-3p | Homo sapiens | 2019 | [62][63][64] | HOXB Cluster | |||
| HOXB-AS2 | Potentially participate in the development of atrial fibrillation | ? | ? | ? | Homo sapiens | 2020 | [65] | HOXB-AS1 | 797 | Linc or NAT | HOXB2→H |
| HOXB-AS3 | OXB3 | ||||||||||
| Control of energetic metabolism in the cell through alternating the isoforms of pyruvate kinase M (PKM); promoting of the cancer processes through repression of p53 transcription; activation of PI3/AKT pathway | Codes the conservative peptide of 53 amino acids long, which is important for PKM splicing; Sponges miRNAs | Nucleus Cytoplasm |
PKM; DNMT1; p53; I3K-AKT-mTOR pathway; miR-378a-3p |
Homo sapiens Rodents |
2017 | [66][67][68][69][70] | HOXB-AS2 | 3594 | NAT RNA | HO | |
| HOXB-AS4 | XB3 | ||||||||||
| The sequence is differentially methylated in normal and pancreatic cancer cells | ? | ? | ? | Homo sapiens | 2018 | [71] | HOXB-AS3 | 785; 611; 549; 545; 514; 452; 446; 336 | Linc or NAT * | HOXB4→B5→HOXB6 | |
| HOXB-AS5 or PRAC2 |
Associated with breast cancer (lncRNA) and protstate cancer (protein) | Encodes 140 aa nuclear protein |
Nucleus (protein) | I3K-AKT-mTOR pathway (lncRNA) | Artiodactyla Bats Colugo Primates |
2003 (protein) 2017 (lncRNA) |
[72][73 | HOXB-AS4 | 543; 513 | Linc | HOXB9→HOXA13 |
| ] | |||||||||||
| HOXC-AS1 | Cholesterol homeostasis participation, inhibition of atherosclerosis; promotion of growth and metastatic formation in several types of malignant tumors | Promotes the transcription and translation of HOXC6; boosts c-MYC mRNA | Nucleus Cytoplasm |
HOXC6; miR-590-3p; c-MYC; Wnt pathway; miR-590-3p |
Homo sapiens | 2016 | [74][75][76][77][78] | PRAC2 | 1193; 560; 518; 503; 448 | Linc * | HOXB9→HOXA13 |
| HOXC-AS2 | Promotion of growth and metastatic formation in several types of malignant tumors | Promote HOXC13 transcription; can sponge miR-876-5p to affect ZEB1 expression | Nucleus Cytoplasm |
HOXC13; ZEB1; miR-876-5p |
Homo sapiens | 2019 | [79][80][81][82] | HOXC Cluster | |||
| HOXC-AS3 | Functions under the direct control of HOXB1. Promotion of growth and metastatic formation in several types of malignant tumors |
Promote the transcription of 5’HOXC genes; stabilizes HOXC10 mRNA; can sponge miR-3922-5p, impairs the maturation of miR-96 | Nucleus | HOXC8; HOXC9; HOXC10; HOXC11; HOXC12; HOXC13; YBX1; thymidine kinase 1 (TK1); FOXM1; miR-96; miR-3922-5p | Homo sapiens | 2018 | [37][81][82][83][84][85][86][87][88][89] | HOXC-AS1 | 548 | ||
| HOTAIR | Reprogramming of chromatin state to promote cancer metastasis; PRC2 and PRC2-independent induction of transcriptional repression. Promotion of growth and metastatic formation in several types of malignant tumorsIntronic | Scaffold: A 5′ domain of HOTAIR binds PRC2, whereas a 3′ domain of HOTAIR binds the LSD1/CoREST/REST complex; HOXC9 |
|||||||||
| Sponging big set microRNA | Nucleus | Cytoplasm |
HOXD cluster (40 Kb in 5′area) HOXA1; HOXA5; HOXC11; p53; p27; E-cadherin; NOTCH1/JAGGED1; SNAIL; GLI2; Protocadherin 10; Wnt pathway; Dozens of miRNAs, critical for proliferation and differentiation control | Rodents Carnivores Primates Marsupials |
2007 | [28][35][90][91][92][93][94][95][96][97][98][99][100][101][102][103][104][105][106][107][108][109][110][111][112][113][114][115][116][117][118][119] | HOXC-AS2 | 504 | Linc or NAT | HOXC9→H | |
| HOXC13-AS | OXC10 | ||||||||||
| Promotion of proliferation, migration and invasion of cells of several cancer types | Sponging big set microRNA | Cytoplasm | HOXC13; c-MYC; miR-383-3p; miR-497-5p |
Homo sapiens | 2019 | [120][121][122][123][124] | HOXC-AS3 | 368 | Linc or NAT | HOXC10→HOXC11 | |
| HAGLR or HOXD-AS1 | Impairment of HAGLR regulation (up or down) to promote growth and metastatic formation of several types of malignant tumors | Binds to WDR5 and EZH2 for activation and repression of target genes | Nucleus | HOTAIR | 2370; 2364; 2337 | Linc | HOXC11→HOXC12 | ||||
| HOXC13-AS | 1408 | Linc or NAT | HOXC9→ | ||||||||
| HOXD Cluster | |||||||||||
| HAGLR | 4086; 4037; 4007; 3942; 3923; 3905; 3893; 3891; 3821; 3812; 3794; 3782 | Linc or NAT | HOXD1→HOXD3 | ||||||||
| HOXD-AS2 | 692 | Linc or NAT | HOXD4→D8→D9→HOXD10 | ||||||||
| As ncRNA | Functions | Mechanism of Work | Localization | ||||
|---|---|---|---|---|---|---|---|
| Cytoplasm | |||||||
| HOXD3; | |||||||
| JAK2/STAT3 pathway; Wnt pathway; Ras/ERK pathway; TGF-β pathway; p57; miR-133a-3p; miR-133b; miR-130a; miR-185-5p | |||||||
| Rodents | |||||||
| Primates | |||||||
| 2014 | |||||||
| [ | |||||||
| 125 | |||||||
| ] | |||||||
| [ | 126 | ] | [ | 127 | ] | [ | 128][129][130][131][132][133][134][135][136][137][138] |
| HOXD-AS2 | Downregulation of HOXD-AS2 significantly promotes the progression of gastric cancer | ? | Cytoplasm | HOXD8; PI3K/Akt pathway |
Homo sapiens | 2018 | [139][140][141][142][143][144][145][146][147][148][149] |
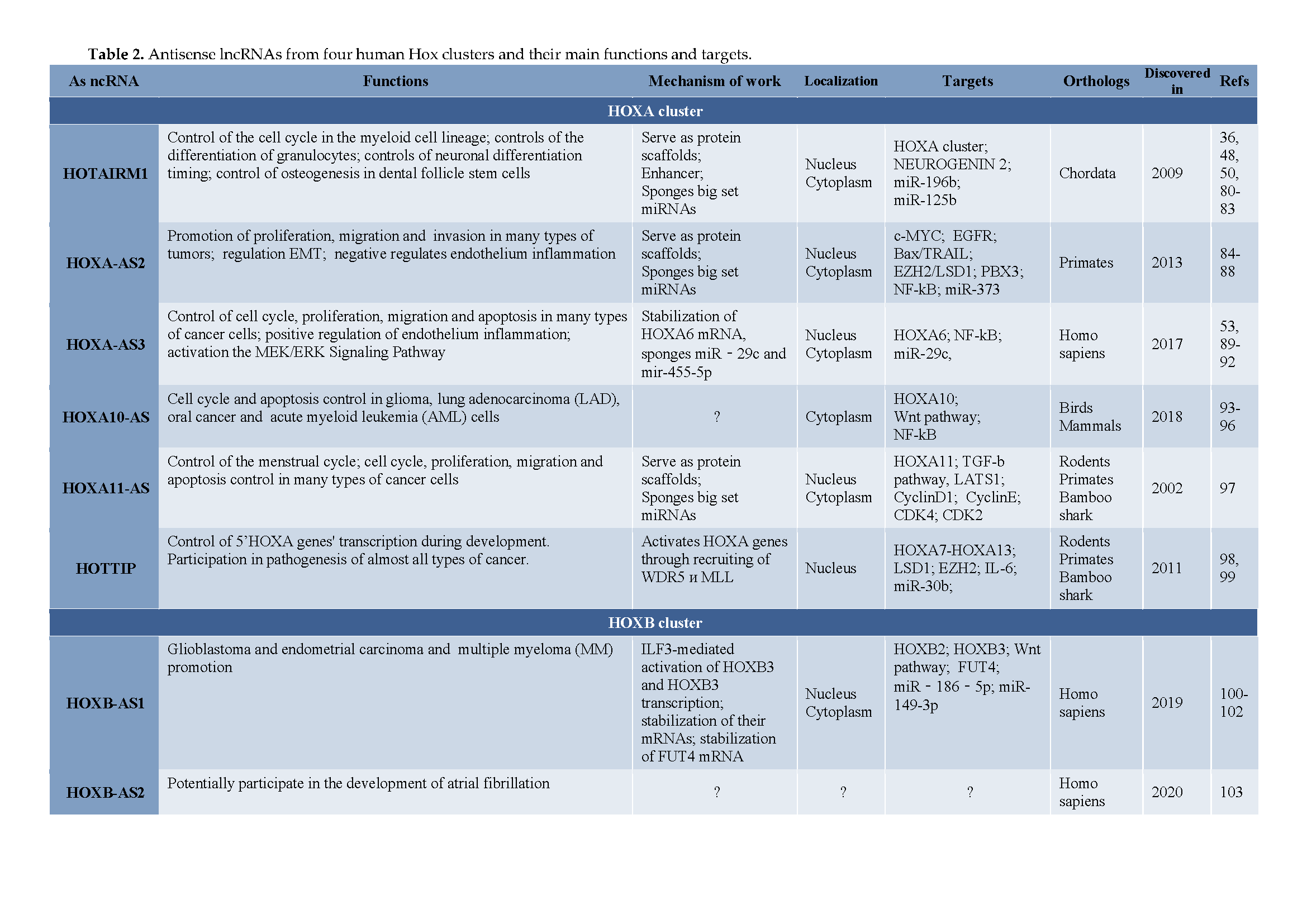
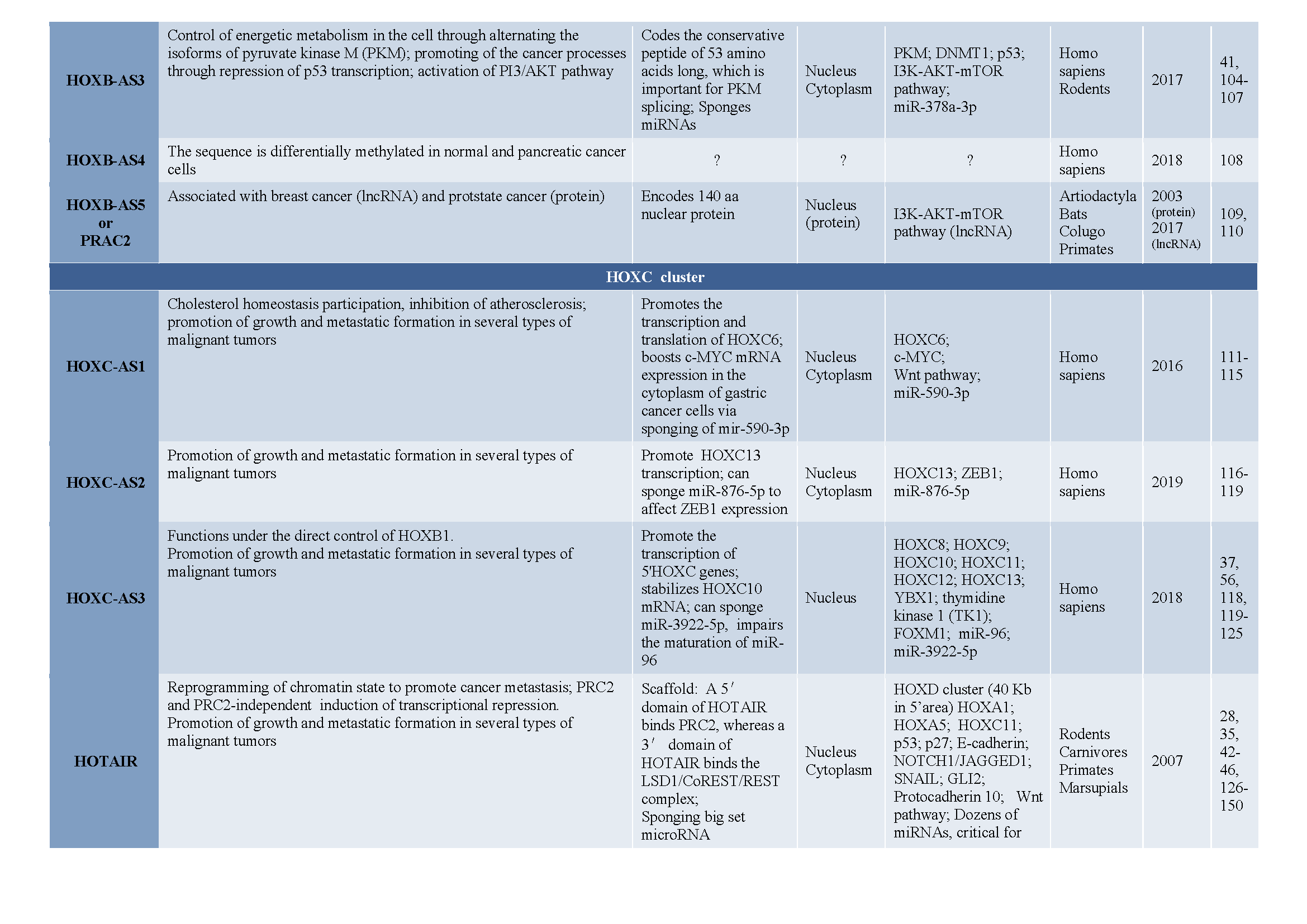
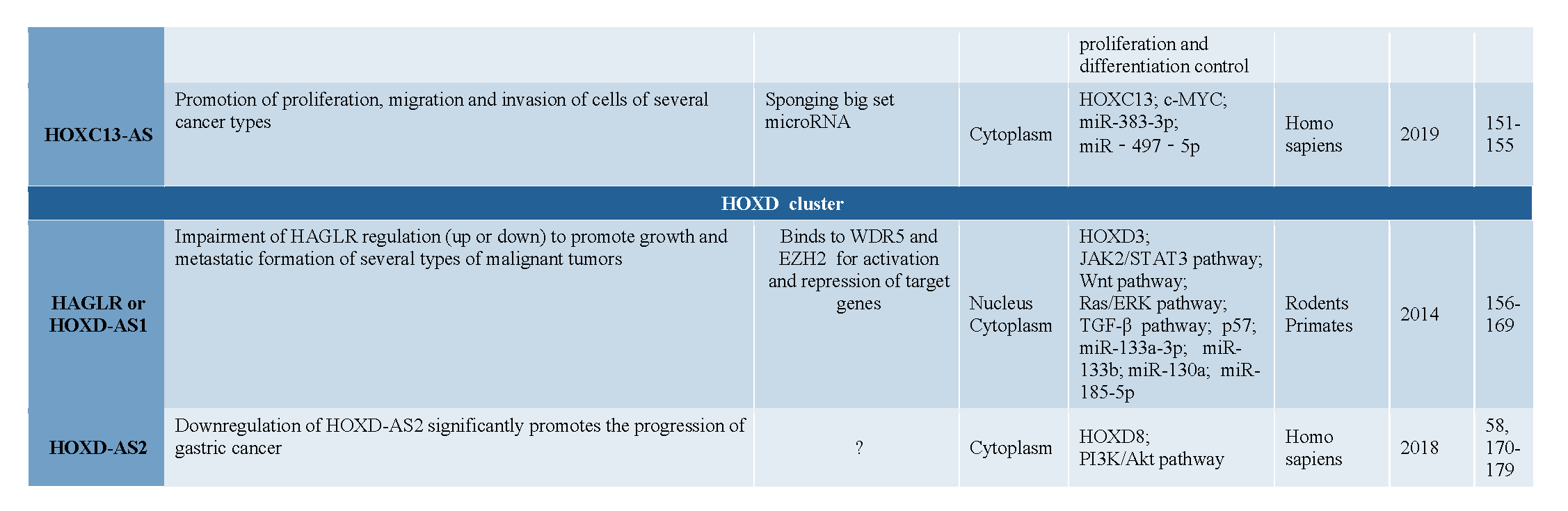
3. Antisense LncRNAs in Hox Clusters of Protostomian Animals
4. The Implications for the Uprise of Antisense LncRNAs in Hox Clusters and the Reasons for Their Evolutionary Maintenance
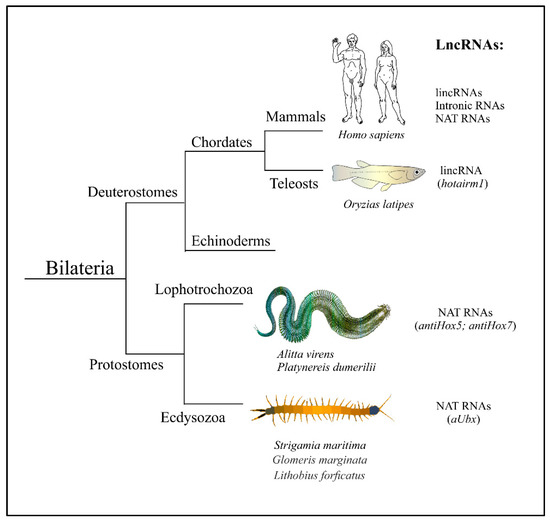
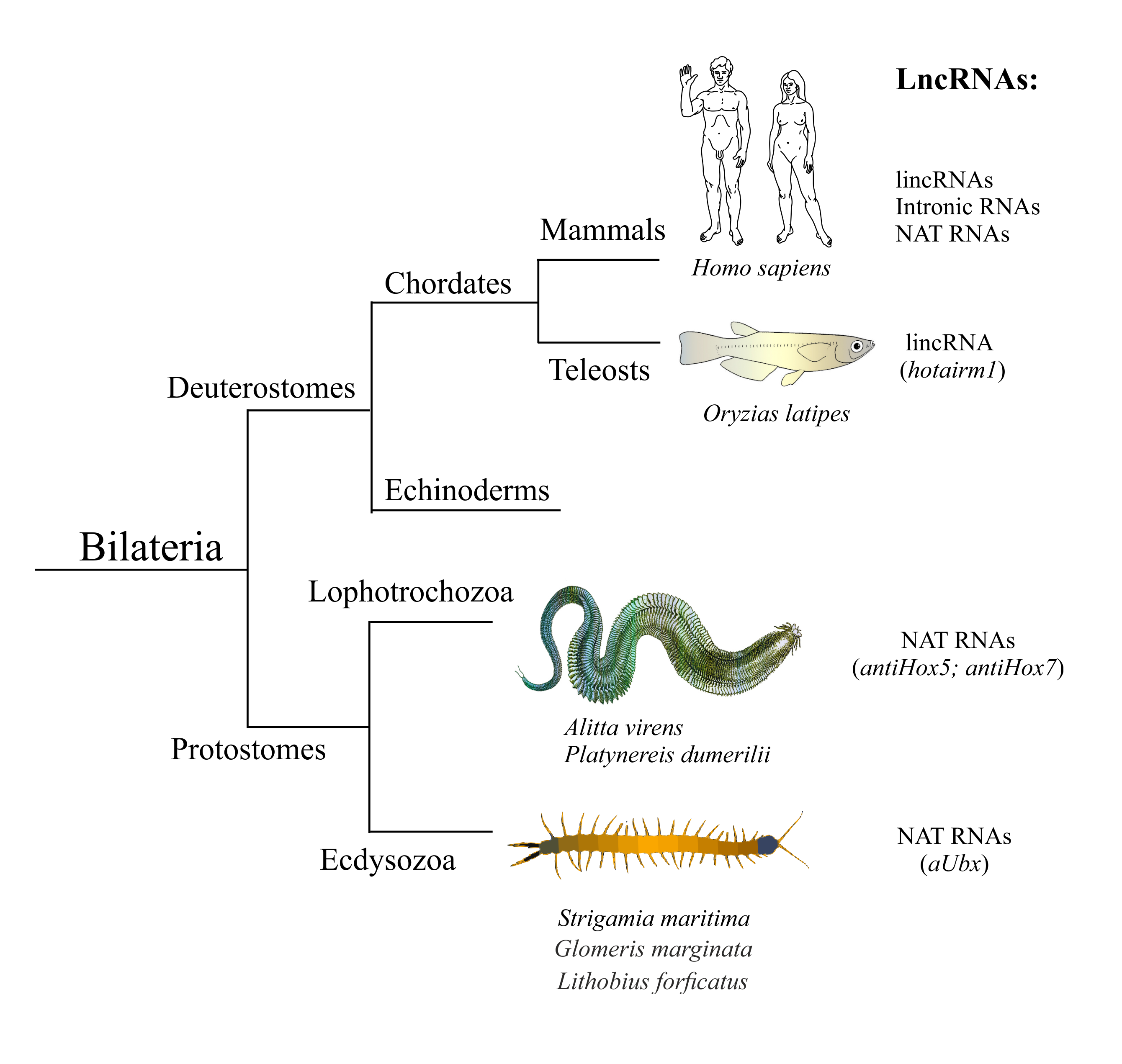
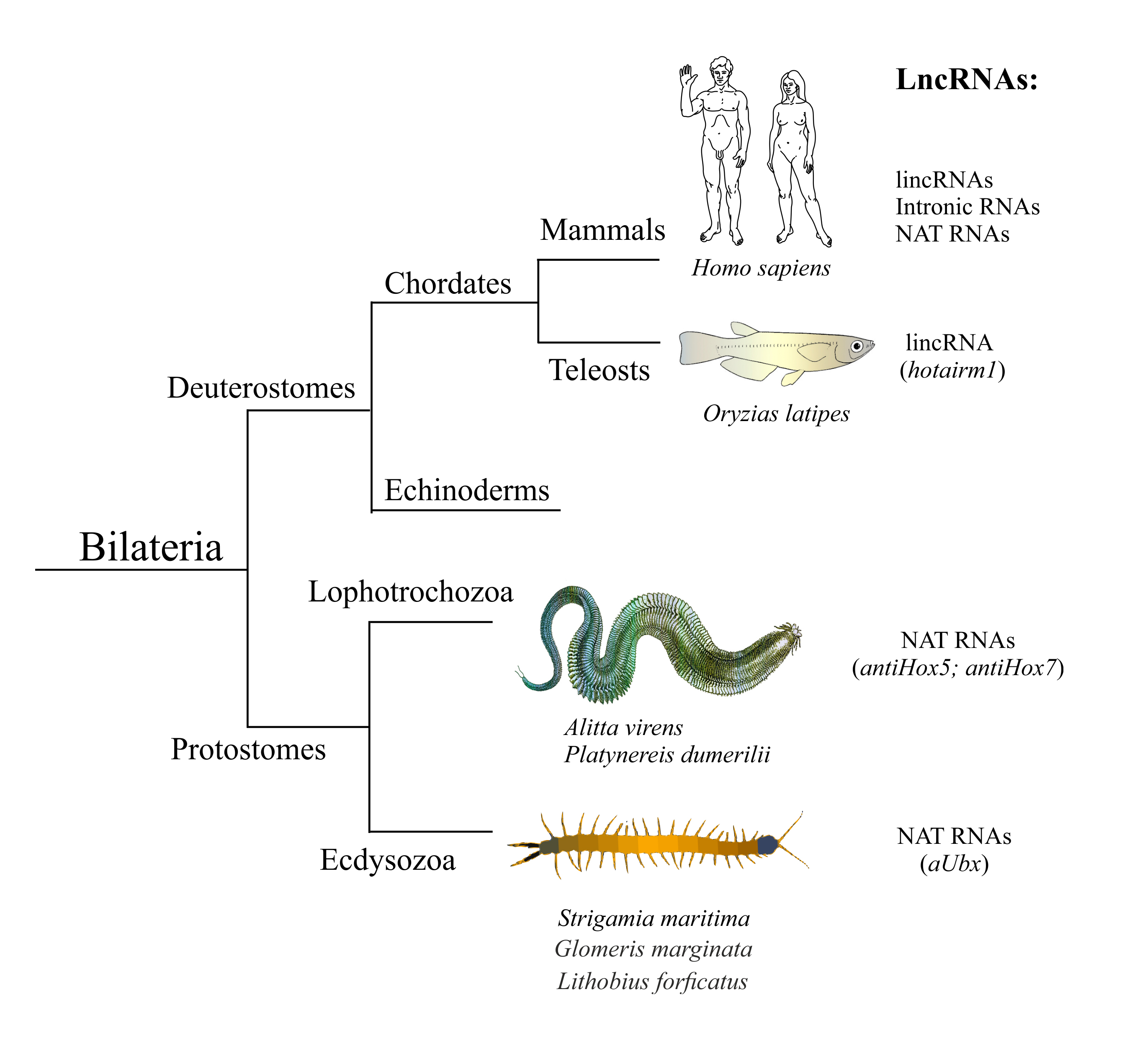
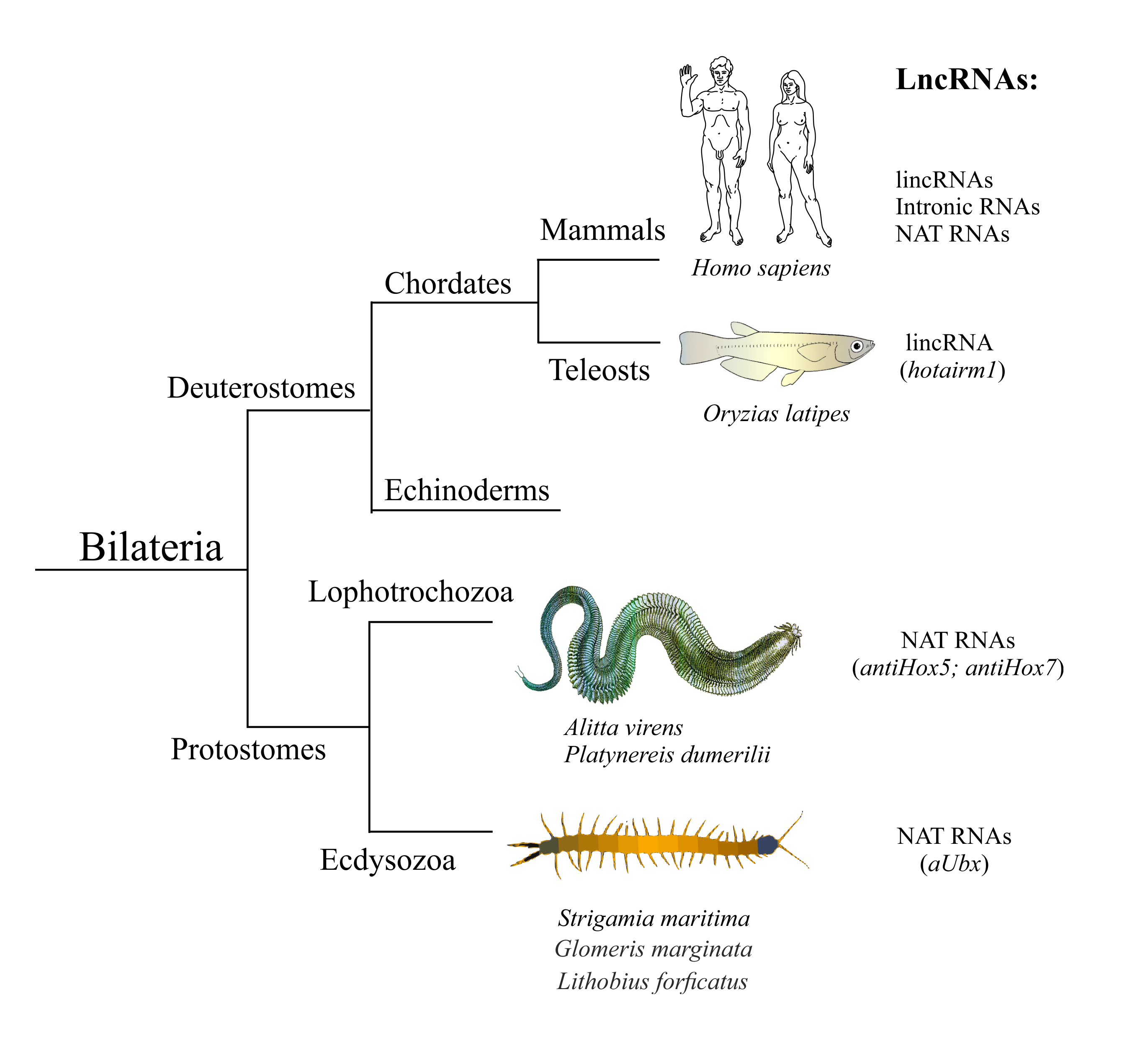
ineages - Deuterostomia, Lophotrochozoa, and Ecdysozoa. Individual human lncRNAs are listed in Tab1.
References
- Lewis, E.B. A gene complex controlling segmentation in Drosophila. Nature 1978, 7, 565–570.Lewis, E.B. A gene complex controlling segmentation in Drosophila. Nature 1978, 7, 565–570. [CrossRef]
- Maslakov, G.P.; Kulishkin, N.S.; Surkova, A.A.; Kulakova, M.A. Maternal Transcripts of Hox Genes are Found in Oocytes of Platynereis dumerilii (Annelida, Nereididae). J. Dev. Biol. 2021. Unpublished work.
- Awgulewitsch, A. Hox in hair growth and development. Naturwissenschaften 2003, 90, 193–211.Awgulewitsch, A. Hox in hair growth and development. Naturwissenschaften 2003, 90, 193–211. [CrossRef] [PubMed]
- Brun, A.C.; Björnsson, J.M.; Magnusson, M.; Larsson, N.; Leveén, P.; Ehinger, M.; Nilsson, E.; Karlsson, S. Hoxb4-deficient mice undergo normal hematopoietic development but exhibit a mild proliferation defect in hematopoietic stem cells. Blood 2004, 103, 4126–4133.Brun, A.C.; Björnsson, J.M.; Magnusson, M.; Larsson, N.; Leveén, P.; Ehinger, M.; Nilsson, E.; Karlsson, S. Hoxb4-deficient mice undergo normal hematopoietic development but exhibit a mild proliferation defect in hematopoietic stem cells. Blood 2004, 103, 4126–4133. [CrossRef]
- Argiropoulos, B.; Humphries, R.K. Hox genes in hematopoiesis and leukemogenesis. Oncogene 2007, 26, 6766–6776.Argiropoulos, B.; Humphries, R.K. Hox genes in hematopoiesis and leukemogenesis. Oncogene 2007, 26, 6766–6776. [CrossRef] [PubMed]
- Morgan, R.; Whiting, K. Differential expression of HOX genes upon activation of leukocyte sub-populations. Int. J. Hematol. 2008, 87, 246–249.Morgan, R.; Whiting, K. Differential expression of HOX genes upon activation of leukocyte sub-populations. Int. J. Hematol. 2008, 87, 246–249. [CrossRef] [PubMed]
- Stansbury, M.S.; Moczek, A.P. The function of Hox and appendage-patterning genes in the development of an evolutionary novelty, the Photuris firefly lantern. Proc. Biol. Sci. 2014, 281, 20133333.Stansbury, M.S.; Moczek, A.P. The function of Hox and appendage-patterning genes in the development of an evolutionary novelty, the Photuris firefly lantern. Proc. Biol. Sci. 2014, 281, 20133333. [CrossRef]
- Horabin, J.I. Long noncoding RNAs as metazoan developmental regulators. Chromosome Res. 2013, 21, 673–684.Horabin, J.I. Long noncoding RNAs as metazoan developmental regulators. Chromosome Res. 2013, 21, 673–684. [CrossRef]
- Brosnan, C.A.; Voinnet, O. The long and the short of noncoding RNAs. Curr. Opin. Cell Biol. 2009, 21, 416–425.Brosnan, C.A.; Voinnet, O. The long and the short of noncoding RNAs. Curr. Opin. Cell Biol. 2009, 21, 416–425. [CrossRef]
- Ponting, C.P.; Belgard, T.G. Transcribed dark matter: Meaning or myth? Hum. Mol. Genet. 2010, 19, R162–R168.Ponting, C.P.; Belgard, T.G. Transcribed dark matter: Meaning or myth? Hum. Mol. Genet. 2010, 19, R162–R168. [CrossRef] [PubMed]
- Fernandez-Valverde, S.L.; Calcino, A.D.; Degnan, B.M. Deep developmental transcriptome sequencing uncovers numerous new genes and enhances gene annotation in the sponge Amphimedon queenslandica. BMC Genom. 2015, 16, 387.Fernandez-Valverde, S.L.; Calcino, A.D.; Degnan, B.M. Deep developmental transcriptome sequencing uncovers numerous new genes and enhances gene annotation in the sponge Amphimedon queenslandica. BMC Genom. 2015, 16, 387. [CrossRef]
- Gaiti, F.; Fernandez-Valverde, S.L.; Nakanishi, N.; Calcino, A.D.; Yanai, I.; Tanurdzic, M.; Degnan, B.M. Dynamic and Widespread lncRNA Expression in a Sponge and the Origin of Animal Complexity. Mol. Biol. Evol. 2015, 32, 2367–2382.Gaiti, F.; Fernandez-Valverde, S.L.; Nakanishi, N.; Calcino, A.D.; Yanai, I.; Tanurdzic, M.; Degnan, B.M. Dynamic and Widespread lncRNA Expression in a Sponge and the Origin of Animal Complexity. Mol. Biol. Evol. 2015, 32, 2367–2382. [CrossRef] [PubMed]
- Vickaryous, M.K.; Hall, B.K. Human cell type diversity, evolution, development, and classification with special reference to cells derived from the neural crest. Biol. Rev. Camb. Philos. Soc. 2006, 81, 425–455.Vickaryous, M.K.; Hall, B.K. Human cell type diversity, evolution, development, and classification with special reference to cells derived from the neural crest. Biol. Rev. Camb. Philos. Soc. 2006, 81, 425–455. [CrossRef]
- Pertea, M.; Shumate, A.; Pertea, G.; Varabyou, A.; Breitwieser, F.P.; Chang, Y.-C.; Madugundu, A.K.; Pandey, A.; Salzberg, S.L. CHESS: A new human gene catalog curated from thousands of large-scale RNA sequencing experiments reveals extensive transcriptional noise. Genome Biol. 2018, 19, 208.Pertea, M.; Shumate, A.; Pertea, G.; Varabyou, A.; Breitwieser, F.P.; Chang, Y.-C.; Madugundu, A.K.; Pandey, A.; Salzberg, S.L. CHESS: A new human gene catalog curated from thousands of large-scale RNA sequencing experiments reveals extensive transcriptional noise. Genome Biol. 2018, 19, 208. [CrossRef] [PubMed]
- Cabili, M.N.; Trapnell, C.; Goff, L.; Koziol, M.; Tazon-Vega, B.; Regev, A.; Rinn, J.L. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev. 2011, 25, 1915–1927.Cabili, M.N.; Trapnell, C.; Goff, L.; Koziol, M.; Tazon-Vega, B.; Regev, A.; Rinn, J.L. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev. 2011, 25, 1915–1927. [CrossRef]
- Sarropoulos, I.; Marin, R.; Cardoso-Moreira, M.; Kaessmann, H. Developmental dynamics of lncRNAs across mammalian organs and species. Nature 2019, 571, 510–514.Sarropoulos, I.; Marin, R.; Cardoso-Moreira, M.; Kaessmann, H. Developmental dynamics of lncRNAs across mammalian organs and species. Nature 2019, 571, 510–514. [CrossRef]
- Katayama, S.; Tomaru, Y.; Kasukawa, T.; Waki, K.; Nakanishi, M.; Nakamura, M.; Nishida, H.; RIKEN Genome Exploration Research Group; Genome Science Group (Genome Network Project Core Group); FANTOM Consortium; et al. Antisense transcription in the mammalian transcriptome. Science 2005, 309, 1564–1566.Katayama, S.; Tomaru, Y.; Kasukawa, T.; Waki, K.; Nakanishi, M.; Nakamura, M.; Nishida, H.; RIKEN Genome Exploration Research Group; Genome Science Group (Genome Network Project Core Group); FANTOM Consortium; et al. Antisense transcription in the mammalian transcriptome. Science 2005, 309, 1564–1566. [PubMed]
- Lipshitz, H.D.; Peattie, D.A.; Hogness, D.S. Novel transcripts from the Ultrabithorax domain of the bithorax complex. Genes Dev. 1987, 1, 307–322.Lipshitz, H.D.; Peattie, D.A.; Hogness, D.S. Novel transcripts from the Ultrabithorax domain of the bithorax complex. Genes Dev. 1987, 1, 307–322. [CrossRef] [PubMed]
- Zhang, X.; Lian, Z.; Padden, C.; Gerstein, M.B.; Rozowsky, J.; Snyder, M.; Gingeras, T.R.; Kapranov, P.; Weissman, S.M.; Newburger, P.E. A myelopoiesis-associated regulatory intergenic noncoding RNA transcript within the human HOXA cluster. Blood 2009, 113, 2526–2534.Zhang, X.; Lian, Z.; Padden, C.; Gerstein, M.B.; Rozowsky, J.; Snyder, M.; Gingeras, T.R.; Kapranov, P.;Weissman, S.M.; Newburger, P.E. A myelopoiesis-associated regulatory intergenic noncoding RNA transcript within the human HOXA cluster. Blood 2009, 113, 2526–2534. [CrossRef]
- Li, L.; Liu, B.; Wapinski, O.L.; Tsai, M.C.; Qu, K.; Zhang, J.; Carlson, J.C.; Lin, M.; Fang, F.; Gupta, R.A.; et al. Targeted disruption of Hotair leads to homeotic transformation and gene derepression. Cell Rep. 2013, 5, 3–12.Li, L.; Liu, B.; Wapinski, O.L.; Tsai, M.C.; Qu, K.; Zhang, J.; Carlson, J.C.; Lin, M.; Fang, F.; Gupta, R.A.; et al. Targeted disruption of Hotair leads to homeotic transformation and gene derepression. Cell Rep. 2013, 5, 3–12. [CrossRef]
- Zhao, H.; Zhang, X.; Frazão, J.B.; Condino-Neto, A.; Newburger, P.E. HOX antisense lincRNA HOXA-AS2 is an apoptosis repressor in all trans retinoic acid treated NB4 promyelocytic leukemia cells. J. Cell Biochem. 2013, 114, 2375–2383.Zhao, H.; Zhang, X.; Frazão, J.B.; Condino-Neto, A.; Newburger, P.E. HOX antisense lincRNA HOXA-AS2 is an apoptosis repressor in all trans retinoic acid treated NB4 promyelocytic leukemia cells. J. Cell Biochem. 2013, 114, 2375–2383. [CrossRef]
- Pradeepa, M.M.; McKenna, F.; Taylor, G.C.A.; Bengani, H.; Grimes, G.R.; Wood, A.J.; Bhatia, B.; Bickmore, W.A. Psip1/p52 regulates posterior Hoxa genes through activation of lncRNA Hottip. PLoS Genet. 2017, 13, e1006677.Pradeepa, M.M.; McKenna, F.; Taylor, G.C.A.; Bengani, H.; Grimes, G.R.; Wood, A.J.; Bhatia, B.; Bickmore, W.A. Psip1/p52 regulates posterior Hoxa genes through activation of lncRNA Hottip. PLoS Genet. 2017, 13, e1006677. [CrossRef]
- Duboule, D. The rise and fall of Hox gene clusters. Development 2007, 134, 2549–2560.Duboule, D. The rise and fall of Hox gene clusters. Development 2007, 134, 2549–2560. [CrossRef]
- Engström, P.G.; Suzuki, H.; Ninomiya, N.; Akalin, A.; Sessa, L.; Lavorgna, G.; Brozzi, A.; Luzi, L.; Tan, S.L.; Yang, L.; et al. Complex Loci in human and mouse genomes. PLoS Genet. 2006, 2, e47.Engström, P.G.; Suzuki, H.; Ninomiya, N.; Akalin, A.; Sessa, L.; Lavorgna, G.; Brozzi, A.; Luzi, L.; Tan, S.L.; Yang, L.; et al. Complex Loci in human and mouse genomes. PLoS Genet. 2006, 2, e47. [CrossRef] [PubMed]
- Bedford, M.; Arman, E.; Orr-Urtreger, A.; Lonai, P. Analysis of the Hoxd-3 gene: Structure and localization of its sense and natural antisense transcripts. DNA Cell Biol. 1995, 14, 295–304.Bedford, M.; Arman, E.; Orr-Urtreger, A.; Lonai, P. Analysis of the Hoxd-3 gene: Structure and localization of its sense and natural antisense transcripts. DNA Cell Biol. 1995, 14, 295–304. [CrossRef]
- Hsieh-Li, H.M.; Witte, D.P.; Weinstein, M.; Branford, W.; Li, H.; Small, K.; Potter, S.S. Hoxa 11 structure, extensive antisense transcription, and function in male and female fertility. Development 1995, 121, 1373–1385.Hsieh-Li, H.M.; Witte, D.P.; Weinstein, M.; Branford, W.; Li, H.; Small, K.; Potter, S.S. Hoxa 11 structure, extensive antisense transcription, and function in male and female fertility. Development 1995, 121, 1373–1385. [CrossRef]
- Sessa, L.; Breiling, A.; Lavorgna, G.; Silvestri, L.; Casari, G.; Orlando, V. Noncoding RNA synthesis and loss of Polycomb group repression accompanies the colinear activation of the human HOXA cluster. RNA 2007, 13, 223–239.Sessa, L.; Breiling, A.; Lavorgna, G.; Silvestri, L.; Casari, G.; Orlando, V. Noncoding RNA synthesis and loss of Polycomb group repression accompanies the colinear activation of the human HOXA cluster. RNA 2007, 13, 223–239. [CrossRef] [PubMed]
- Rinn, J.L.; Kertesz, M.; Wang, J.K.; Squazzo, S.L.; Xu, X.; Brugmann, S.A.; Goodnough, L.H.; Helms, J.A.; Farnham, P.J.; Segal, E.; et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 2007, 129, 1311–1323.Rinn, J.L.; Kertesz, M.; Wang, J.K.; Squazzo, S.L.; Xu, X.; Brugmann, S.A.; Goodnough, L.H.; Helms, J.A.; Farnham, P.J.; Segal, E.; et al. Functional demarcation of active and silent chromatin domains in human HOX loci by noncoding RNAs. Cell 2007, 129, 1311–1323. [CrossRef] [PubMed]
- Mainguy, G.; Koster, J.; Woltering, J.; Jansen, H.; Durston, A. Extensive polycistronism and antisense transcription in the mammalian Hox clusters. PLoS ONE 2007, 2, e356.Mainguy, G.; Koster, J.; Woltering, J.; Jansen, H.; Durston, A. Extensive polycistronism and antisense transcription in the mammalian Hox clusters. PLoS ONE 2007, 2, e356. [CrossRef]
- Simeone, A.; Pannese, M.; Acampora, D.; D’Esposito, M.; Boncinelli, E. At least three human homeoboxes on chromosome 12 belong to the same transcription unit. Nucleic Acids Res. 1988, 16, 5379–5390.Simeone, A.; Pannese, M.; Acampora, D.; D’Esposito, M.; Boncinelli, E. At least three human homeoboxes on chromosome 12 belong to the same transcription unit. Nucleic Acids Res. 1988, 16, 5379–5390. [CrossRef]
- Shiga, Y.; Sagawa, K.; Takai, R.; Sakaguchi, H.; Yamagata, H.; Hayashi, S. Transcriptional readthrough of Hox genes Ubx and Antp and their divergent post-transcriptional control during crustacean evolution. Evol. Dev. 2006, 8, 407–414.Shiga, Y.; Sagawa, K.; Takai, R.; Sakaguchi, H.; Yamagata, H.; Hayashi, S. Transcriptional readthrough of Hox genes Ubx and Antp and their divergent post-transcriptional control during crustacean evolution. Evol. Dev. 2006, 8, 407–414. [CrossRef] [PubMed]
- Brena, C.; Chipman, A.D.; Minelli, A.; Akam, M. Expression of trunk Hox genes in the centipede Strigamia maritima: Sense and anti-sense transcripts. Evol. Dev. 2006, 8, 252–265.Brena, C.; Chipman, A.D.; Minelli, A.; Akam, M. Expression of trunk Hox genes in the centipede Strigamia maritima: Sense and anti-sense transcripts. Evol. Dev. 2006, 8, 252–265. [CrossRef] [PubMed]
- Janssen, R.; Budd, G.E. Gene expression suggests conserved aspects of Hox gene regulation in arthropods and provides additional support for monophyletic Myriapoda. Evodevo 2010, 1, 4.Janssen, R.; Budd, G.E. Gene expression suggests conserved aspects of Hox gene regulation in arthropods and provides additional support for monophyletic Myriapoda. Evodevo 2010, 1, 4. [CrossRef]
- Degani, N.; Ainbinder, E.; Ulitsky, I. Highly conserved and cis-acting lncRNAs produced from paralogous regions in the center of HOXA and HOXB clusters in the endoderm lineage. bioRxiv 2020.Degani, N.; Ainbinder, E.; Ulitsky, I. Highly conserved and cis-acting lncRNAs produced from paralogous regions in the center of HOXA and HOXB clusters in the endoderm lineage. bioRxiv 2020. [CrossRef]
- Diederichs, S. The four dimensions of noncoding RNA conservation. Trends Genet. 2014, 30, 121–123.Diederichs, S. The four dimensions of noncoding RNA conservation. Trends Genet. 2014, 30, 121–123. [CrossRef]
- Herrera-Úbeda, C.; Marín-Barba, M.; Navas-Pérez, E.; Gravemeyer, J.; Albuixech-Crespo, B.; Wheeler, G.N.; Garcia-Fernàndez, J. Microsyntenic Clusters Reveal Conservation of lncRNAs in Chordates Despite Absence of Sequence Conservation. Biology 2019, 8, 61.Herrera-Úbeda, C.; Marín-Barba, M.; Navas-Pérez, E.; Gravemeyer, J.; Albuixech-Crespo, B.; Wheeler, G.N.; Garcia-Fernàndez, J. Microsyntenic Clusters Reveal Conservation of lncRNAs in Chordates Despite Absence of Sequence Conservation. Biology 2019, 8, 61. [CrossRef]
- Wang, X.; Sun, Y.; Xu, T.; Qian, K.; Huang, B.; Zhang, K.; Song, Z.; Qian, T.; Shi, J.; Li, L. HOXB13 promotes proliferation, migration, and invasion of glioblastoma through transcriptional upregulation of lncRNA HOXC-AS3. J. Cell Biochem. 2019, 120, 15527–15537.Wang, X.; Sun, Y.; Xu, T.; Qian, K.; Huang, B.; Zhang, K.; Song, Z.; Qian, T.; Shi, J.; Li, L. HOXB13 promotes proliferation, migration, and invasion of glioblastoma through transcriptional upregulation of lncRNA HOXC-AS3. J. Cell Biochem. 2019, 120, 15527–15537. [CrossRef]
- Latgé, G.; Poulet, C.; Bours, V.; Josse, C.; Jerusalem, G. Natural Antisense Transcripts: Molecular Mechanisms and Implications in Breast Cancers. Int. J. Mol. Sci. 2018, 19, 123.Latgé, G.; Poulet, C.; Bours, V.; Josse, C.; Jerusalem, G. Natural Antisense Transcripts: Molecular Mechanisms and Implications in Breast Cancers. Int. J. Mol. Sci. 2018, 19, 123. [CrossRef]
- Chen, Z.; Zheng, J.; Hong, H.; Chen, D.; Deng, L.; Zhang, X.; Ling, J.; Wu, L. lncRNA HOTAIRM1 promotes osteogenesis of hDFSCs by epigenetically regulating HOXA2 via DNMT1 in vitro. J. Cell Physiol. 2020, 235, 8507–8519.Chen, Z.; Zheng, J.; Hong, H.; Chen, D.; Deng, L.; Zhang, X.; Ling, J.; Wu, L. lncRNA HOTAIRM1 promotes osteogenesis of hDFSCs by epigenetically regulating HOXA2 via DNMT1 in vitro. J. Cell Physiol. 2020, 235, 8507–8519. [CrossRef] [PubMed]
- Rea, J.; Menci, V.; Tollis, P.; Santini, T.; Armaos, A.; Garone, M.G.; Iberite, F.; Cipriano, A.; Tartaglia, G.G.; Rosa, A.; et al. HOTAIRM1 regulates neuronal differentiation by modulating NEUROGENIN 2 and the downstream neurogenic cascade. Cell Death Dis. 2020, 11, 527.Rea, J.; Menci, V.; Tollis, P.; Santini, T.; Armaos, A.; Garone, M.G.; Iberite, F.; Cipriano, A.; Tartaglia, G.G.; Rosa, A.; et al. HOTAIRM1 regulates neuronal differentiation by modulating NEUROGENIN 2 and the downstream neurogenic cascade. Cell Death Dis. 2020, 11, 527. [CrossRef] [PubMed]
- Zhang, X.; Weissman, S.M.; Newburger, P.E. Long intergenic non-coding RNA HOTAIRM1 regulates cell cycle progression during myeloid maturation in NB4 human promyelocytic leukemia cells. RNA Biol. 2014, 11, 777–787.Zhang, X.;Weissman, S.M.; Newburger, P.E. Long intergenic non-coding RNA HOTAIRM1 regulates cell cycle progression during myeloid maturation in NB4 human promyelocytic leukemia cells. RNA Biol. 2014, 11, 777–787. [CrossRef] [PubMed]
- Díaz-Beyá, M.; Brunet, S.; Nomdedéu, J.; Pratcorona, M.; Cordeiro, A.; Gallardo, D.; Escoda, L.; Tormo, M.; Heras, I.; Ribera, J.M.; et al. The lincRNA HOTAIRM1, located in the HOXA genomic region, is expressed in acute myeloid leukemia, impacts prognosis in patients in the intermediate-risk cytogenetic category, and is associated with a distinctive microRNA signature. Oncotarget 2015, 6, 31613–31627.Díaz-Beyá, M.; Brunet, S.; Nomdedéu, J.; Pratcorona, M.; Cordeiro, A.; Gallardo, D.; Escoda, L.; Tormo, M.; Heras, I.; Ribera, J.M.; et al. The lincRNA HOTAIRM1, located in the HOXA genomic region, is expressed in acute myeloid leukemia, impacts prognosis in patients in the intermediate-risk cytogenetic category, and is associated with a distinctive microRNA signature. Oncotarget, 2015, 6, 31613–31627. [PubMed]
- Shi, T.; Guo, D.; Xu, H.; Su, G.; Chen, J.; Zhao, Z.; Shi, J.; Wedemeyer, M.; Attenello, F.; Zhang, L.; et al. HOTAIRM1, an enhancer lncRNA, promotes glioma proliferation by regulating long-range chromatin interactions within HOXA cluster genes. Mol. Biol. Rep. 2020, 47, 2723–2733.Shi, T.; Guo, D.; Xu, H.; Su, G.; Chen, J.; Zhao, Z.; Shi, J.; Wedemeyer, M.; Attenello, F.; Zhang, L.; et al. HOTAIRM1, an enhancer lncRNA, promotes glioma proliferation by regulating long-range chromatin interactions within HOXA cluster genes. Mol. Biol. Rep. 2020, 47, 2723–2733. [CrossRef] [PubMed]
- Li, Y.Q.; Sun, N.; Zhang, C.S.; Li, N.; Wu, B.; Zhang, J.L. Inactivation of lncRNA HOTAIRM1 caused by histone methyltransferase RIZ1 accelerated the proliferation and invasion of liver cancer. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 8767–8777.
- Fang, Y.; Wang, J.; Wu, F.; Song, Y.; Zhao, S.; Zhang, Q. Long non-coding RNA HOXA-AS2 promotes proliferation and invasion of breast cancer by acting as a miR-520c-3p sponge. Oncotarget 2017, 8, 46090–46103.Fang, Y.; Wang, J.; Wu, F.; Song, Y.; Zhao, S.; Zhang, Q. Long non-coding RNA HOXA-AS2 promotes proliferation and invasion of breast cancer by acting as a miR-520c-3p sponge. Oncotarget 2017, 8, 46090–46103. [CrossRef]
- Zhang, Y.; Xu, J.; Zhang, S.; An, J.; Zhang, J.; Huang, J.; Jin, Y. HOXA-AS2 Promotes Proliferation and Induces Epithelial-Mesenchymal Transition via the miR-520c-3p/GPC3 Axis in Hepatocellular Carcinoma. Cell Physiol. Biochem. 2018, 50, 2124–2138.Zhang, Y.; Xu, J.; Zhang, S.; An, J.; Zhang, J.; Huang, J.; Jin, Y. HOXA-AS2 Promotes Proliferation and Induces Epithelial-Mesenchymal Transition via the miR-520c-3p/GPC3 Axis in Hepatocellular Carcinoma. Cell Physiol. Biochem. 2018, 50, 2124 2138. [CrossRef]
- Zhu, X.; Liu, Y.; Yu, J.; Du, J.; Guo, R.; Feng, Y.; Zhong, G.; Jiang, Y.; Lin, J. LncRNA HOXA-AS2 represses endothelium inflammation by regulating the activity of NF-κB signaling. Atherosclerosis 2019, 281, 38–46.Zhu, X.; Liu, Y.; Yu, J.; Du, J.; Guo, R.; Feng, Y.; Zhong, G.; Jiang, Y.; Lin, J. LncRNA HOXA-AS2 represses endothelium inflammation by regulating the activity of NF-_B signaling. Atherosclerosis 2019, 281, 38–46. [CrossRef]
- Xiao, S.; Song, B. LncRNA HOXA-AS2 promotes the progression of prostate cancer via targeting miR-509-3p/PBX3 axis. Biosci. Rep. 2020, 40, BSR20193287.Xiao, S.; Song, B. LncRNA HOXA-AS2 promotes the progression of prostate cancer via targeting miR-509-3p/PBX3 axis. Biosci. Rep. 2020, 40, BSR20193287. [CrossRef]
- Wang, J.; Su, Z.; Lu, S.; Fu, W.; Liu, Z.; Jiang, X.; Tai, S. LncRNA HOXA-AS2 and its molecular mechanisms in human cancer. Clin. Chim. Acta 2018, 485, 229–233.Wang, J.; Su, Z.; Lu, S.; Fu, W.; Liu, Z.; Jiang, X.; Tai, S. LncRNA HOXA-AS2 and its molecular mechanisms in human cancer. Clin. Chim. Acta 2018, 485, 229–233. [CrossRef] [PubMed]
- Wu, F.; Zhang, C.; Cai, J.; Yang, F.; Liang, T.; Yan, X.; Wang, H.; Wang, W.; Chen, J.; Jiang, T. Upregulation of long noncoding RNA HOXA-AS3 promotes tumor progression and predicts poor prognosis in glioma. Oncotarget 2017, 8, 53110–53123.Wu, F.; Zhang, C.; Cai, J.; Yang, F.; Liang, T.; Yan, X.; Wang, H.; Wang, W.; Chen, J.; Jiang, T. Upregulation of long noncoding RNA HOXA-AS3 promotes tumor progression and predicts poor prognosis in glioma. Oncotarget 2017, 8, 53110–53123. [CrossRef]
- Zhang, H.; Liu, Y.; Yan, L.; Zhang, M.; Yu, X.; Du, W.; Wang, S.; Li, Q.; Chen, H.; Zhang, Y.; et al. Increased levels of the long noncoding RNA, HOXA-AS3, promote proliferation of A549 cells. Cell Death Dis. 2018, 9, 707.Zhang, H.; Liu, Y.; Yan, L.; Zhang, M.; Yu, X.; Du, W.; Wang, S.; Li, Q.; Chen, H.; Zhang, Y.; et al. Increased levels of the long noncoding RNA, HOXA-AS3, promote proliferation of A549 cells. Cell Death Dis. 2018, 9, 707. [CrossRef]
- Tong, Y.; Wang, M.; Dai, Y.; Bao, D.; Zhang, J.; Pan, H. LncRNA HOXA-AS3 Sponges miR-29c to Facilitate Cell Proliferation, Metastasis, and EMT Process and Activate the MEK/ERK Signaling Pathway in Hepatocellular Carcinoma. Hum. Gene Ther. Clin. Dev. 2019, 30, 129–141.Tong, Y.; Wang, M.; Dai, Y.; Bao, D.; Zhang, J.; Pan, H. LncRNA HOXA-AS3 Sponges miR-29c to Facilitate Cell Proliferation, Metastasis, and EMT Process and Activate the MEK/ERK Signaling Pathway in Hepatocellular Carcinoma. Hum. Gene Ther. Clin. Dev. 2019, 30, 129–141. [CrossRef] [PubMed]
- Zhu, X.; Chen, D.; Liu, Y.; Yu, J.; Qiao, L.; Lin, S.; Chen, D.; Zhong, G.; Lu, X.; Wang, Y.; et al. Long Noncoding RNA HOXA-AS3 Integrates NF-κB Signaling to Regulate Endothelium Inflammation. Mol. Cell Biol. 2019, 39, e00139-19.Zhu, X.; Chen, D.; Liu, Y.; Yu, J.; Qiao, L.; Lin, S.; Chen, D.; Zhong, G.; Lu, X.;Wang, Y.; et al. Long Noncoding RNA HOXA-AS3 Integrates NF-_B Signaling to Regulate Endothelium Inflammation. Mol. Cell Biol. 2019, 39, e00139-19. [CrossRef] [PubMed]
- Chen, W.; Li, Q.; Zhang, G.; Wang, H.; Zhu, Z.; Chen, L. LncRNA HOXA-AS3 promotes the malignancy of glioblastoma through regulating miR-455-5p/USP3 axis. J. Cell Mol. Med. 2020, 24, 11755–11767.Chen, W.; Li, Q.; Zhang, G.; Wang, H.; Zhu, Z.; Chen, L. LncRNA HOXA-AS3 promotes the malignancy of glioblastoma through regulating miR-455-5p/USP3 axis. J. Cell Mol. Med. 2020, 24, 11755–11767. [CrossRef] [PubMed]
- Dong, C.Y.; Cui, J.; Li, D.H.; Li, Q.; Hong, X.Y. HOXA10-AS: A novel oncogenic long non-coding RNA in glioma. Oncol. Rep. 2018, 40, 2573–2583.Dong, C.Y.; Cui, J.; Li, D.H.; Li, Q.; Hong, X.Y. HOXA10-AS: A novel oncogenic long non-coding RNA in glioma. Oncol. Rep. 2018, 40, 2573–2583. [CrossRef] [PubMed]
- Sheng, K.; Lu, J.; Zhao, H. ELK1-induced upregulation of lncRNA HOXA10-AS promotes lung adenocarcinoma progression by increasing Wnt/β-catenin signaling. Biochem. Biophys. Res. Commun. 2018, 501, 612–618.Sheng, K.; Lu, J.; Zhao, H. ELK1-induced upregulation of lncRNA HOXA10-AS promotes lung adenocarcinoma progression by increasing Wnt/_-catenin signaling. Biochem. Biophys. Res. Commun. 2018, 501, 612–618. [CrossRef]
- Al-Kershi, S.; Bhayadia, R.; Ng, M.; Verboon, L.; Emmrich, S.; Gack, L.; Schwarzer, A.; Strowig, T.; Heckl, D.; Klusmann, J.H. The stem cell-specific long noncoding RNA HOXA10-AS in the pathogenesis of KMT2A-rearranged leukemia. Blood Adv. 2019, 3, 4252–4263.Al-Kershi, S.; Bhayadia, R.; Ng, M.; Verboon, L.; Emmrich, S.; Gack, L.; Schwarzer, A.; Strowig, T.; Heckl, D.; Klusmann, J.H. The stem cell-specific long noncoding RNA HOXA10-AS in the pathogenesis of KMT2A-rearranged leukemia. Blood Adv. 2019, 3, 4252–4263. [CrossRef] [PubMed]
- Yan, X.; Cong, B.; Chen, Q.; Liu, L.; Luan, X.; Du, J.; Cao, M. Silencing lncRNA HOXA10-AS decreases cell proliferation of oral cancer and HOXA10-antisense RNA can serve as a novel prognostic predictor. J. Int. Med. Res. 2020, 48.Yan, X.; Cong, B.; Chen, Q.; Liu, L.; Luan, X.; Du, J.; Cao, M. Silencing lncRNA HOXA10-AS decreases cell proliferation of oral cancer and HOXA10-antisense RNA can serve as a novel prognostic predictor. J. Int. Med. Res. 2020, 48. [CrossRef]
- Wei, C.; Zhao, L.; Liang, H.; Zhen, Y.; Han, L. Recent advances in unraveling the molecular mechanisms and functions of HOXA11-AS in human cancers and other diseases (Review). Oncol. Rep. 2020, 43, 1737–1754.Wei, C.; Zhao, L.; Liang, H.; Zhen, Y.; Han, L. Recent advances in unraveling the molecular mechanisms and functions of HOXA11-AS in human cancers and other diseases (Review). Oncol. Rep. 2020, 43, 1737–1754. [CrossRef]
- Wang, K.C.; Yang, Y.W.; Liu, B.; Sanyal, A.; Corces-Zimmerman, R.; Chen, Y.; Lajoie, B.R.; Protacio, A.; Flynn, R.A.; Gupta, R.A.; et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature 2011, 472, 120–124.Wang, K.C.; Yang, Y.W.; Liu, B.; Sanyal, A.; Corces-Zimmerman, R.; Chen, Y.; Lajoie, B.R.; Protacio, A.; Flynn, R.A.; Gupta, R.A.; et al. A long noncoding RNA maintains active chromatin to coordinate homeotic gene expression. Nature 2011, 472, 120–124. [CrossRef]
- Ghafouri-Fard, S.; Dashti, S.; Taheri, M. The HOTTIP (HOXA transcript at the distal tip) lncRNA: Review of oncogenic roles in human. Biomed. Pharmacother. 2020, 127, 110158.Ghafouri-Fard, S.; Dashti, S.; Taheri, M. The HOTTIP (HOXA transcript at the distal tip) lncRNA: Review of oncogenic roles in human. Biomed. Pharmacother. 2020, 127, 110158. [CrossRef] [PubMed]
- Chen, R.; Zhang, X.; Wang, C. LncRNA HOXB-AS1 promotes cell growth in multiple myeloma via FUT4 mRNA stability by ELAVL1. J. Cell Biochem. 2020, 121, 4043–4051.Chen, R.; Zhang, X.; Wang, C. LncRNA HOXB-AS1 promotes cell growth in multiple myeloma via FUT4 mRNA stability by ELAVL1. J. Cell Biochem. 2020, 121, 4043–4051. [CrossRef] [PubMed]
- Bi, Y.; Mao, Y.; Su, Z.; Du, J.; Ye, L.; Xu, F. HOXB-AS1 accelerates the tumorigenesis of glioblastoma via modulation of HOBX2 and HOBX3 at transcriptional and posttranscriptional levels. J. Cell Physiol. 2021, 236, 93–106.Bi, Y.; Mao, Y.; Su, Z.; Du, J.; Ye, L.; Xu, F. HOXB-AS1 accelerates the tumorigenesis of glioblastoma via modulation of HOBX2 and HOBX3 at transcriptional and posttranscriptional levels. J. Cell Physiol. 2021, 236, 93–106. [CrossRef] [PubMed]
- Liu, D.; Qiu, M.; Jiang, L.; Liu, K. Long Noncoding RNA HOXB-AS1 Is Upregulated in Endometrial Carcinoma and Sponged miR-149-3p to Upregulate Wnt10b. Technol. Cancer Res. Treat. 2020, 19.Liu, D.; Qiu, M.; Jiang, L.; Liu, K. Long Noncoding RNA HOXB-AS1 Is Upregulated in Endometrial Carcinoma and Sponged miR-149-3p to Upregulate Wnt10b. Technol. Cancer Res. Treat. 2020, 19. [CrossRef]
- Shi, X.; Shao, X.; Liu, B.; Lv, M.; Pandey, P.; Guo, C.; Zhang, R.; Zhang, Y. Genome-wide screening of functional long noncoding RNAs in the epicardial adipose tissues of atrial fibrillation. Biochim. Biophys. Acta Mol. Basis Dis. 2020, 1866, 165757.Shi, X.; Shao, X.; Liu, B.; Lv, M.; Pandey, P.; Guo, C.; Zhang, R.; Zhang, Y. Genome-wide screening of functional long noncoding RNAs in the epicardial adipose tissues of atrial fibrillation. Biochim. Biophys. Acta Mol. Basis Dis. 2020, 1866, 165757. [CrossRef]
- Huang, J.Z.; Chen, M.; Chen, D.; Gao, X.C.; Zhu, S.; Huang, H.; Hu, M.; Zhu, H.; Yan, G.R. A Peptide Encoded by a Putative lncRNA HOXB-AS3 Suppresses Colon Cancer Growth. Mol. Cell. 2017, 68, 171–184.Huang, J.Z.; Chen, M.; Chen, D.; Gao, X.C.; Zhu, S.; Huang, H.; Hu, M.; Zhu, H.; Yan, G.R. A Peptide Encoded by a Putative lncRNA HOXB-AS3 Suppresses Colon Cancer Growth. Mol. Cell. 2017, 68, 171–184. [CrossRef]
- Zhang, X.M.; Chen, H.; Zhou, B.; Zhang, Q.Y.; Liao, Y.; Wang, J.S.; Wang, Z.H. lncRNA HOXB-AS3 promotes hepatoma by inhibiting p53 expression. Eur. Rev. Med. Pharmacol. Sci. 2018, 22, 6784–6792.
- Jiang, W.; Kai, J.; Li, D.; Wei, Z.; Wang, Y.; Wang, W. lncRNA HOXB-AS3 exacerbates proliferation, migration, and invasion of lung cancer via activating the PI3K-AKT pathway. J. Cell Physiol. 2020, 235, 7194–7203.Jiang,W.; Kai, J.; Li, D.;Wei, Z.;Wang, Y.;Wang,W. lncRNA HOXB-AS3 exacerbates proliferation, migration, and invasion of lung cancer via activating the PI3K-AKT pathway. J. Cell Physiol. 2020, 235, 7194–7203. [CrossRef]
- Huang, H.H.; Chen, F.Y.; Chou, W.C.; Hou, H.A.; Ko, B.S.; Lin, C.T.; Tang, J.L.; Li, C.C.; Yao, M.; Tsay, W.; et al. Long non-coding RNA HOXB-AS3 promotes myeloid cell proliferation and its higher expression is an adverse prognostic marker in patients with acute myeloid leukemia and myelodysplastic syndrome. BMC Cancer 2019, 19, 617.Huang, H.H.; Chen, F.Y.; Chou, W.C.; Hou, H.A.; Ko, B.S.; Lin, C.T.; Tang, J.L.; Li, C.C.; Yao, M.; Tsay, W.; et al. Long non-coding RNA HOXB-AS3 promotes myeloid cell proliferation and its higher expression is an adverse prognostic marker in patients with acute myeloid leukemia and myelodysplastic syndrome. BMC Cancer 2019, 19, 617. [CrossRef]
- Xu, S.; Jia, G.; Zhang, H.; Wang, L.; Cong, Y.; Lv, M.; Xu, J.; Ruan, H.; Jia, X.; Xu, P.; et al. LncRNA HOXB-AS3 promotes growth, invasion and migration of epithelial ovarian cancer by altering glycolysis. Life Sci. 2021, 264, 118636.Xu, S.; Jia, G.; Zhang, H.;Wang, L.; Cong, Y.; Lv, M.; Xu, J.; Ruan, H.; Jia, X.; Xu, P.; et al. LncRNA HOXB-AS3 promotes growth, invasion and migration of epithelial ovarian cancer by altering glycolysis. Life Sci. 2021, 264, 118636. [CrossRef]
- Ishihara, H.; Yamashita, S.; Amano, R.; Kimura, K.; Hirakawa, K.; Ueda, T.; Murakami, Y.; Tamori, A.; Tanabe, K.; Kawada, N.; et al. Pancreatic Cancer Cell Fraction Estimation in a DNA Sample. Oncology 2018, 95, 370–379.Ishihara, H.; Yamashita, S.; Amano, R.; Kimura, K.; Hirakawa, K.; Ueda, T.; Murakami, Y.; Tamori, A.; Tanabe, K.; Kawada, N.; et al. Pancreatic Cancer Cell Fraction Estimation in a DNA Sample. Oncology 2018, 95, 370–379. [CrossRef]
- Olsson, P.; Motegi, A.; Bera, T.K.; Lee, B.; Pastan, I. PRAC2: A new gene expressed in human prostate and prostate cancer. Prostate 2003, 56, 123–130.Olsson, P.; Motegi, A.; Bera, T.K.; Lee, B.; Pastan, I. PRAC2: A new gene expressed in human prostate and prostate cancer. Prostate, 2003, 56, 123–130. [CrossRef]
- Rui, J.; Chunming, Z.; Binbin, G.; Na, S.; Shengxi, W.; Wei, S. IL-22 promotes the progression of breast cancer through regulating HOXB-AS5. Oncotarget 2017, 8, 103601–103612.Rui, J.; Chunming, Z.; Binbin, G.; Na, S.; Shengxi, W.; Wei, S. IL-22 promotes the progression of breast cancer through regulating HOXB-AS5. Oncotarget 2017, 8, 103601–103612. [CrossRef] [PubMed]
- Huang, C.; Hu, Y.W.; Zhao, J.J.; Ma, X.; Zhang, Y.; Guo, F.X.; Kang, C.M.; Lu, J.B.; Xiu, J.C.; Sha, Y.H.; et al. Long Noncoding RNA HOXC-AS1 Suppresses Ox-LDL-Induced Cholesterol Accumulation Through Promoting HOXC6 Expression in THP-1 Macrophages. DNA Cell Biol. 2016, 35, 722–729.Huang, C.; Hu, Y.W.; Zhao, J.J.; Ma, X.; Zhang, Y.; Guo, F.X.; Kang, C.M.; Lu, J.B.; Xiu, J.C.; Sha, Y.H.; et al. Long Noncoding RNA HOXC-AS1 Suppresses Ox-LDL-Induced Cholesterol Accumulation Through Promoting HOXC6 Expression in THP-1 Macrophages. DNA Cell Biol. 2016, 35, 722–729. [CrossRef] [PubMed]
- Dong, Y.; Li, X.; Lin, Z.; Zou, W.; Liu, Y.; Qian, H.; Jia, J. HOXC-AS1-MYC regulatory loop contributes to the growth and metastasis in gastric cancer. J. Exp. Clin. Cancer Res. 2019, 38, 502.Dong, Y.; Li, X.; Lin, Z.; Zou,W.; Liu, Y.; Qian, H.; Jia, J. HOXC-AS1-MYC regulatory loop contributes to the growth and metastasis in gastric cancer. J. Exp. Clin. Cancer Res. 2019, 38, 502. [CrossRef] [PubMed]
- Takayama, K.I.; Fujimura, T.; Suzuki, Y.; Inoue, S. Identification of long non-coding RNAs in advanced prostate cancer associated with androgen receptor splicing factors. Commun. Biol. 2020, 3, 393.Takayama, K.I.; Fujimura, T.; Suzuki, Y.; Inoue, S. Identification of long non-coding RNAs in advanced prostate cancer associated with androgen receptor splicing factors. Commun. Biol. 2020, 3, 393. [CrossRef]
- Zhou, C.; An, N.; Cao, C.; Wang, G. lncRNA HOXC-AS1 promotes gastric cancer via binding eIF4AIII by activating Wnt/β-catenin signaling. J. Gene Med. 2020, 22, e3202.Zhou, C.; An, N.; Cao, C.;Wang, G. lncRNA HOXC-AS1 promotes gastric cancer via binding eIF4AIII by activating Wnt/_-catenin signaling. J. Gene Med. 2020, 22, e3202. [CrossRef]
- Zhang, S.; Wang, L.; Gao, Y.; Fan, Y.; Zhang, G.; Zhang, Y. Molecular Mechanism of 73HOXC-AS1-Activated Wntβ-Catenin Signaling and eIF4AIII in Promoting Progression of Gastric Cancer. Biomed Res. Int. 2021, 2021, 8814843.Zhang, S.; Wang, L.; Gao, Y.; Fan, Y.; Zhang, G.; Zhang, Y. Molecular Mechanism of 73HOXC-AS1-Activated Wntb-Catenin Signaling and eIF4AIII in Promoting Progression of Gastric Cancer. Biomed Res. Int. 2021, 2021, 8814843.
- Dong, N.; Guo, J.; Han, S.; Bao, L.; Diao, Y.; Lin, Z. Positive feedback loop of lncRNA HOXC-AS2/miR-876-5p/ZEB1 to regulate EMT in glioma. Onco Targets Ther. 2019, 12, 7601–7609.Dong, N.; Guo, J.; Han, S.; Bao, L.; Diao, Y.; Lin, Z. Positive feedback loop of lncRNA HOXC-AS2/miR-876-5p/ZEB1 to regulate EMT in glioma. Onco Targets Ther. 2019, 12, 7601–7609. [CrossRef]
- Turner, D.C.; Gorski, P.P.; Maasar, M.F.; Seaborne, R.A.; Baumert, P.; Brown, A.D.; Kitchen, M.O.; Erskine, R.M.; Dos-Remedios, I.; Voisin, S.; et al. DNA methylation across the genome in aged human skeletal muscle tissue and muscle-derived cells: The role of HOX genes and physical activity. Sci. Rep. 2020, 10, 15360.Turner, D.C.; Gorski, P.P.; Maasar, M.F.; Seaborne, R.A.; Baumert, P.; Brown, A.D.; Kitchen, M.O.; Erskine, R.M.; Dos-Remedios, I.; Voisin, S.; et al. DNA methylation across the genome in aged human skeletal muscle tissue and muscle-derived cells: The role of HOX genes and physical activity. Sci. Rep. 2020, 10, 15360. [CrossRef]
- Fu, T.; Ji, X.; Bu, Z.; Zhang, J.; Wu, X.; Zong, X.; Fan, B.; Jia, Z.; Ji, J. Identification of key long non-coding RNAs in gastric adenocarcinoma. Cancer Biomark. 2020, 27, 541–553.Fu, T.; Ji, X.; Bu, Z.; Zhang, J.; Wu, X.; Zong, X.; Fan, B.; Jia, Z.; Ji, J. Identification of key long non-coding RNAs in gastric adenocarcinoma. Cancer Biomark. 2020, 27, 541–553. [CrossRef] [PubMed]
- Liu, B.; Li, J.; Li, J.M.; Liu, G.Y.; Wang, Y.S. HOXC-AS2 mediates the proliferation, apoptosis, and migration of non-small cell lung cancer by combining with HOXC13 gene. Cell Cycle 2021, 20, 236–246.Liu, B.; Li, J.; Li, J.M.; Liu, G.Y.; Wang, Y.S. HOXC-AS2 mediates the proliferation, apoptosis, and migration of non-small cell lung cancer by combining with HOXC13 gene. Cell Cycle 2021, 20, 236–246. [CrossRef]
- Yang, Z.; Hu, T. Long noncoding RNA HOXC-AS3 facilitates the progression of invasive mucinous adenocarcinomas of the lung via modulating FUS/FOXM1. Vitr. Cell Dev. Biol. Anim. 2020, 56, 15–23.Yang, Z.; Hu, T. Long noncoding RNA HOXC-AS3 facilitates the progression of invasive mucinous adenocarcinomas of the lung via modulating FUS/FOXM1. Vitr. Cell Dev. Biol. Anim. 2020, 56, 15–23. [CrossRef]
- Zhang, E.; He, X.; Zhang, C.; Su, J.; Lu, X.; Si, X.; Chen, J.; Yin, D.; Han, L.; De, W. A novel long noncoding RNA HOXC-AS3 mediates tumorigenesis of gastric cancer by binding to YBX1. Genome Biol. 2018, 19, 154.Zhang, E.; He, X.; Zhang, C.; Su, J.; Lu, X.; Si, X.; Chen, J.; Yin, D.; Han, L.; De, W. A novel long noncoding RNA HOXC-AS3 mediates tumorigenesis of gastric cancer by binding to YBX1. Genome Biol. 2018, 19, 154. [CrossRef] [PubMed]
- Li, B.; Han, H.; Song, S.; Fan, G.; Xu, H.; Zhou, W.; Qiu, Y.; Qian, C.; Wang, Y.; Yuan, Z.; et al. HOXC10 Regulates Osteogenesis of Mesenchymal Stromal Cells Through Interaction with Its Natural Antisense Transcript lncHOXC-AS3. Stem Cells 2019, 37, 247–256.Li, B.; Han, H.; Song, S.; Fan, G.; Xu, H.; Zhou,W.; Qiu, Y.; Qian, C.;Wang, Y.; Yuan, Z.; et al. HOXC10 Regulates Osteogenesis of Mesenchymal Stromal Cells Through Interaction with Its Natural Antisense Transcript lncHOXC-AS3. Stem Cells 2019, 37, 247–256. [CrossRef] [PubMed]
- Shi, S.H.; Jiang, J.; Zhang, W.; Sun, L.; Li, X.J.; Li, C.; Ge, Q.D.; Zhuang, Z.G. A Novel lncRNA HOXC-AS3 Acts as a miR-3922-5p Sponge to Promote Breast Cancer Metastasis. Cancer Investig. 2020, 38, 1–12.Shi, S.H.; Jiang, J.; Zhang, W.; Sun, L.; Li, X.J.; Li, C.; Ge, Q.D.; Zhuang, Z.G. A Novel lncRNA HOXC-AS3 Acts as a miR-3922-5p Sponge to Promote Breast Cancer Metastasis. Cancer Investig. 2020, 38, 1–12. [CrossRef]
- Enteghami, M.; Ghorbani, M.; Zamani, M.; Galehdari, H. HOXC10 is significantly overexpressed in colorectal cancer. Biomed. Rep. 2020, 13, 18.Enteghami, M.; Ghorbani, M.; Zamani, M.; Galehdari, H. HOXC10 is significantly overexpressed in colorectal cancer. Biomed. Rep. 2020, 13, 18. [CrossRef]
- Su, J.; Yu, B.; Zhang, C.; Yi, P.; Li, H.; Xu, C.; Cao, L.; Chen, P.; Li, M.; Shen, K.; et al. Long noncoding RNA HOXC-AS3 indicates a poor prognosis and regulates tumorigenesis by binding to YBX1 in breast cancer. Am. J. Transl. Res. 2020, 12, 6335–6350.
- Yang, B.; Sun, L.; Liang, L. LncRNA HOXC-AS3 Suppresses the Formation of Mature miR-96 in Ovarian Cancer Cells to Promote Cell Proliferation. Reprod. Sci. 2021.Yang, B.; Sun, L.; Liang, L. LncRNA HOXC-AS3 Suppresses the Formation of Mature miR-96 in Ovarian Cancer Cells to Promote Cell Proliferation. Reprod. Sci. 2021. [CrossRef] [PubMed]
- Spitale, R.C.; Tsai, M.C.; Chang, H.Y. RNA templating the epigenome: Long noncoding RNAs as molecular scaffolds. Epigenetics 2011, 6, 539–543.Spitale, R.C.; Tsai, M.C.; Chang, H.Y. RNA templating the epigenome: Long noncoding RNAs as molecular scaffolds. Epigenetics 2011, 6, 539–543. [CrossRef]
- Battistelli, C.; Cicchini, C.; Santangelo, L.; Tramontano, A.; Grassi, L.; Gonzalez, F.J.; de Nonno, V.; Grassi, G.; Amicone, L.; Tripodi, M. The Snail repressor recruits EZH2 to specific genomic sites through the enrollment of the lncRNA HOTAIR in epithelial-to-mesenchymal transition. Oncogene 2017, 36, 942–955.Battistelli, C.; Cicchini, C.; Santangelo, L.; Tramontano, A.; Grassi, L.; Gonzalez, F.J.; de Nonno, V.; Grassi, G.; Amicone, L.; Tripodi, M. The Snail repressor recruits EZH2 to specific genomic sites through the enrollment of the lncRNA HOTAIR in epithelial-to-mesenchymal transition. Oncogene 2017, 36, 942–955. [CrossRef]
- He, S.; Liu, S.; Zhu, H. The sequence, structure and evolutionary features of HOTAIR in mammals. BMC Evol. Biol. 2011, 11, 102.He, S.; Liu, S.; Zhu, H. The sequence, structure and evolutionary features of HOTAIR in mammals. BMC Evol. Biol. 2011, 11, 102.[CrossRef]
- Schorderet, P.; Duboule, D. Structural and functional differences in the long non-coding RNA hotair in mouse and human. PLoS Genet. 2011, 7, e1002071.Schorderet, P.; Duboule, D. Structural and functional differences in the long non-coding RNA hotair in mouse and human. PLoS Genet. 2011, 7, e1002071. [CrossRef]
- Amândio, A.R.; Necsulea, A.; Joye, E.; Mascrez, B.; Duboule, D. Hotair Is Dispensible for Mouse Development. PLoS Genet. 2016, 12, e1006232.Amândio, A.R.; Necsulea, A.; Joye, E.; Mascrez, B.; Duboule, D. Hotair Is Dispensible for Mouse Development. PLoS Genet. 2016, 12, e1006232. [CrossRef] [PubMed]
- Khalil, A.M.; Guttman, M.; Huarte, M.; Garber, M.; Raj, A.; Morales, D.R.; Thomas, K.; Presser, A.; Bernstein, B.E.; van Oudenaarden, A.; et al. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc. Natl. Acad. Sci. USA 2009, 106, 11667–11672.Khalil, A.M.; Guttman, M.; Huarte, M.; Garber, M.; Raj, A.; Morales, D.R.; Thomas, K.; Presser, A.; Bernstein, B.E.; van Oudenaarden, A.; et al. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc. Natl. Acad. Sci. USA 2009, 106, 11667–11672. [CrossRef] [PubMed]
- Tsai, M.C.; Manor, O.; Wan, Y.; Mosammaparast, N.; Wang, J.K.; Lan, F.; Shi, Y.; Segal, E.; Chang, H.Y. Long noncoding RNA as modular scaffold of histone modification complexes. Science 2010, 329, 689–693.Tsai, M.C.; Manor, O.;Wan, Y.; Mosammaparast, N.;Wang, J.K.; Lan, F.; Shi, Y.; Segal, E.; Chang, H.Y. Long noncoding RNA as modular scaffold of histone modification complexes. Science 2010, 329, 689–693. [CrossRef] [PubMed]
- Gupta, R.A.; Shah, N.; Wang, K.C.; Kim, J.; Horlings, H.M.; Wong, D.J.; Tsai, M.C.; Hung, T.; Argani, P.; Rinn, J.L.; et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 2010, 464, 1071–1076.Gupta, R.A.; Shah, N.; Wang, K.C.; Kim, J.; Horlings, H.M.; Wong, D.J.; Tsai, M.C.; Hung, T.; Argani, P.; Rinn, J.L.; et al. Long non-coding RNA HOTAIR reprograms chromatin state to promote cancer metastasis. Nature 2010, 464, 1071–1076. [CrossRef]
- Hung, T.; Chang, H.Y. Long noncoding RNA in genome regulation: Prospects and mechanisms. RNA Biol. 2010, 7, 582–585.Hung, T.; Chang, H.Y. Long noncoding RNA in genome regulation: Prospects and mechanisms. RNA Biol. 2010, 7, 582–585. [CrossRef]
- Schiavo, G.; D’Antò, V.; Cantile, M.; Procino, A.; Di Giovanni, S.; Valletta, R.; Terracciano, L.; Baumhoer, D.; Jundt, G.; Cillo, C. Deregulated HOX genes in ameloblastomas are located in physical contiguity to keratin genes. J. Cell Biochem. 2011, 112, 3206–3215.Schiavo, G.; D’Antò, V.; Cantile, M.; Procino, A.; Di Giovanni, S.; Valletta, R.; Terracciano, L.; Baumhoer, D.; Jundt, G.; Cillo, C. Deregulated HOX genes in ameloblastomas are located in physical contiguity to keratin genes. J. Cell Biochem. 2011, 112, 3206–3215. [CrossRef]
- Wutz, A. RNA-mediated silencing mechanisms in mammalian cells. Prog. Mol. Biol. Transl. Sci. 2011, 101, 351–376.
- Guil, S.; Soler, M.; Portela, A.; Carrère, J.; Fonalleras, E.; Gómez, A.; Villanueva, A.; Esteller, M. Intronic RNAs mediate EZH2 regulation of epigenetic targets. Nat. Struct. Mol. Biol. 2012, 19, 664–670.Guil, S.; Soler, M.; Portela, A.; Carrère, J.; Fonalleras, E.; Gómez, A.; Villanueva, A.; Esteller, M. Intronic RNAs mediate EZH2 regulation of epigenetic targets. Nat. Struct. Mol. Biol. 2012, 19, 664–670. [CrossRef]
- Wu, Y.; Zhang, L.; Wang, Y.; Li, H.; Ren, X.; Wei, F.; Yu, W.; Wang, X.; Zhang, L.; Yu, J.; et al. Long noncoding RNA HOTAIR involvement in cancer. Tumour Biol. 2014, 35, 9531–9538.Wu, Y.; Zhang, L.; Wang, Y.; Li, H.; Ren, X.; Wei, F.; Yu, W.; Wang, X.; Zhang, L.; Yu, J.; et al. Long noncoding RNA HOTAIR involvement in cancer. Tumour Biol. 2014, 35, 9531–9538. [CrossRef] [PubMed]
- Deng, Q.; Sun, H.; He, B.; Pan, Y.; Gao, T.; Chen, J.; Ying, H.; Liu, X.; Wang, F.; Xu, Y.; et al. Prognostic value of long non-coding RNA HOTAIR in various cancers. PLoS ONE 2014, 9, e110059.Deng, Q.; Sun, H.; He, B.; Pan, Y.; Gao, T.; Chen, J.; Ying, H.; Liu, X.;Wang, F.; Xu, Y.; et al. Prognostic value of long non-coding RNA HOTAIR in various cancers. PLoS ONE 2014, 9, e110059. [CrossRef] [PubMed]
- Zhou, X.; Chen, J.; Tang, W. The molecular mechanism of HOTAIR in tumorigenesis, metastasis, and drug resistance. Acta Biochim. Biophys. Sin. 2014, 46, 1011–1015.Zhou, X.; Chen, J.; Tang,W. The molecular mechanism of HOTAIR in tumorigenesis, metastasis, and drug resistance. Acta Biochim. Biophys. Sin. 2014, 46, 1011–1015. [CrossRef]
- Dong, C.; Liu, S.; Lv, Y.; Zhang, C.; Gao, H.; Tan, L.; Wang, H. Long non-coding RNA HOTAIR regulates proliferation and invasion via activating Notch signalling pathway in retinoblastoma. J. Biosci. 2016, 41, 677–687.Dong, C.; Liu, S.; Lv, Y.; Zhang, C.; Gao, H.; Tan, L.;Wang, H. Long non-coding RNA HOTAIR regulates proliferation and invasion via activating Notch signalling pathway in retinoblastoma. J. Biosci. 2016, 41, 677–687. [CrossRef]
- Kalwa, M.; Hänzelmann, S.; Otto, S.; Kuo, C.C.; Franzen, J.; Joussen, S.; Fernandez-Rebollo, E.; Rath, B.; Koch, C.; Hofmann, A.; et al. The lncRNA HOTAIR impacts on mesenchymal stem cells via triple helix formation. Nucleic Acids Res. 2016, 44, 10631–10643.Kalwa, M.; Hänzelmann, S.; Otto, S.; Kuo, C.C.; Franzen, J.; Joussen, S.; Fernandez-Rebollo, E.; Rath, B.; Koch, C.; Hofmann, A.; et al. The lncRNA HOTAIR impacts on mesenchymal stem cells via triple helix formation. Nucleic Acids Res. 2016, 44, 10631–10643. [CrossRef] [PubMed]
- Pawłowska, E.; Szczepanska, J.; Blasiak, J. The Long Noncoding RNA HOTAIR in Breast Cancer: Does Autophagy Play a Role? Int. J. Mol. Sci. 2017, 18, 2317.Pawłowska, E.; Szczepanska, J.; Blasiak, J. The Long Noncoding RNA HOTAIR in Breast Cancer: Does Autophagy Play a Role? Int. J. Mol. Sci. 2017, 18, 2317. [CrossRef] [PubMed]
- Zhang, Z.; Zhu, H.; Liu, Y.; Quan, F.; Zhang, X.; Yu, L. LncRNA HOTAIR mediates TGF-β2-induced cell growth and epithelial-mesenchymal transition in human lens epithelial cells. Acta Biochim. Biophys. Sin. 2018, 50, 1028–1037.Zhang, Z.; Zhu, H.; Liu, Y.; Quan, F.; Zhang, X.; Yu, L. LncRNA HOTAIR mediates TGF-_2-induced cell growth and epithelialmesenchymal transition in human lens epithelial cells. Acta Biochim. Biophys. Sin. 2018, 50, 1028–1037. [CrossRef]
- Botti, G.; Scognamiglio, G.; Aquino, G.; Liguori, G.; Cantile, M. LncRNA HOTAIR in Tumor Microenvironment: What Role? Int. J. Mol. Sci. 2019, 20, 2279.Botti, G.; Scognamiglio, G.; Aquino, G.; Liguori, G.; Cantile, M. LncRNA HOTAIR in Tumor Microenvironment: What Role? Int. J. Mol. Sci. 2019, 20, 2279. [CrossRef] [PubMed]
- Shen, J.J.; Zhang, C.H.; Chen, Z.W.; Wang, Z.X.; Yang, D.C.; Zhang, F.L.; Feng, K.H. LncRNA HOTAIR inhibited osteogenic differentiation of BMSCs by regulating Wnt/β-catenin pathway. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 7232–7246.Shen, J.J.; Zhang, C.H.; Chen, Z.W.; Wang, Z.X.; Yang, D.C.; Zhang, F.L.; Feng, K.H. LncRNA HOTAIR inhibited osteogenic differentiation of BMSCs by regulating Wnt/_-catenin pathway. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 7232–7246. [PubMed]
- Bai, J.Y.; Jin, B.; Ma, J.B.; Liu, T.J.; Yang, C.; Chong, Y.; Wang, X.; He, D.; Guo, P. HOTAIR and androgen receptor synergistically increase GLI2 transcription to promote tumor angiogenesis and cancer stemness in renal cell carcinoma. Cancer Lett. 2021, 498, 70–79.Bai, J.Y.; Jin, B.; Ma, J.B.; Liu, T.J.; Yang, C.; Chong, Y.;Wang, X.; He, D.; Guo, P. HOTAIR and androgen receptor synergistically increase GLI2 transcription to promote tumor angiogenesis and cancer stemness in renal cell carcinoma. Cancer Lett. 2021, 498, 70–79. [CrossRef]
- Zhang, J.; Qiu, W.Q.; Zhu, H.; Liu, H.; Sun, J.H.; Chen, Y.; Shen, H.; Qian, C.L.; Shen, Z.Y. HOTAIR contributes to the carcinogenesis of gastric cancer via modulating cellular and exosomal miRNAs level. Cell Death Dis. 2020, 11, 780.Zhang, J.; Qiu,W.Q.; Zhu, H.; Liu, H.; Sun, J.H.; Chen, Y.; Shen, H.; Qian, C.L.; Shen, Z.Y. HOTAIR contributes to the carcinogenesis of gastric cancer via modulating cellular and exosomal miRNAs level. Cell Death Dis. 2020, 11, 780. [CrossRef]
- Ghafouri-Fard, S.; Dashti, S.; Farsi, M.; Taheri, M. HOX transcript antisense RNA: An oncogenic lncRNA in diverse malignancies. Exp. Mol. Pathol. 2021, 118, 104578.Ghafouri-Fard, S.; Dashti, S.; Farsi, M.; Taheri, M. HOX transcript antisense RNA: An oncogenic lncRNA in diverse malignancies. Exp. Mol. Pathol. 2021, 118, 104578. [CrossRef]
- Zhou, Y.H.; Cui, Y.H.; Wang, T.; Luo, Y. Long non-coding RNA HOTAIR in cervical cancer: Molecular marker, mechanistic insight, and therapeutic target. Adv. Clin. Chem. 2020, 97, 117–140.Zhou, Y.H.; Cui, Y.H.;Wang, T.; Luo, Y. Long non-coding RNA HOTAIR in cervical cancer: Molecular marker, mechanistic insight, and therapeutic target. Adv. Clin. Chem. 2020, 97, 117–140.
- Ren, M.M.; Xu, S.; Wei, Y.B.; Yang, J.J.; Yang, Y.N.; Sun, S.S.; Li, Y.J.; Wang, P.Y.; Xie, S.Y. Roles of HOTAIR in lung cancer susceptibility and prognosis. Mol. Genet. Genomic Med. 2020, 8, e1299.Ren, M.M.; Xu, S.; Wei, Y.B.; Yang, J.J.; Yang, Y.N.; Sun, S.S.; Li, Y.J.; Wang, P.Y.; Xie, S.Y. Roles of HOTAIR in lung cancer susceptibility and prognosis. Mol. Genet. Genomic Med. 2020, 8, e1299. [CrossRef]
- Mozdarani, H.; Ezzatizadeh, V.; Rahbar Parvaneh, R. The emerging role of the long non-coding RNA HOTAIR in breast cancer development and treatment. J. Transl. Med. 2020, 18, 152.Mozdarani, H.; Ezzatizadeh, V.; Rahbar Parvaneh, R. The emerging role of the long non-coding RNA HOTAIR in breast cancer development and treatment. J. Transl. Med. 2020, 18, 152. [CrossRef]
- Battistelli, C.; Garbo, S.; Riccioni, V.; Montaldo, C.; Santangelo, L.; Vandelli, A.; Strippoli, R.; Tartaglia, G.G.; Tripodi, M.; Cicchini, C. Design and Functional Validation of a Mutant Variant of the LncRNA HOTAIR to Counteract Snail Function in Epithelial-to-Mesenchymal Transition. Cancer Res. 2021, 81, 103–113.
- Cantile, M.; Di Bonito, M.; De Bellis, M.T.; Botti, G. Functional Interaction among lncRNA HOTAIR and MicroRNAs in Cancer and Other Human Diseases. Cancers 2021, 13, 570.Cantile, M.; Di Bonito, M.; De Bellis, M.T.; Botti, G. Functional Interaction among lncRNA HOTAIR and MicroRNAs in Cancer and Other Human Diseases. Cancers 2021, 13, 570. [CrossRef] [PubMed]
- Liang, Q.; Asila, A.; Deng, Y.; Liao, J.; Liu, Z.; Fang, R. Osteopontin-induced lncRNA HOTAIR expression is involved in osteoarthritis by regulating cell proliferation. BMC Geriatr. 2021, 21, 57.Liang, Q.; Asila, A.; Deng, Y.; Liao, J.; Liu, Z.; Fang, R. Osteopontin-induced lncRNA HOTAIR expression is involved in osteoarthritis by regulating cell proliferation. BMC Geriatr. 2021, 21, 57. [CrossRef] [PubMed]
- Gao, C.; Lu, W.; Lou, W.; Wang, L.; Xu, Q. Long noncoding RNA HOXC13-AS positively affects cell proliferation and invasion in nasopharyngeal carcinoma via modulating miR-383-3p/HMGA2 axis. J. Cell Physiol. 2019, 234, 12809–12820.Gao, C.; Lu,W.; Lou,W.;Wang, L.; Xu, Q. Long noncoding RNA HOXC13-AS positively affects cell proliferation and invasion in nasopharyngeal carcinoma via modulating miR-383-3p/HMGA2 axis. J. Cell Physiol. 2019, 234, 12809–12820. [CrossRef] [PubMed]
- Li, X.; Wang, Q.; Rui, Y.; Zhang, C.; Wang, W.; Gu, J.; Tang, J.; Ding, Y. HOXC13-AS promotes breast cancer cell growth through regulating miR-497-5p/PTEN axis. J. Cell Physiol. 2019, 234, 22343–22351.Li, X.;Wang, Q.; Rui, Y.; Zhang, C.;Wang,W.; Gu, J.; Tang, J.; Ding, Y. HOXC13-AS promotes breast cancer cell growth through regulating miR-497-5p/PTEN axis. J. Cell Physiol. 2019, 234, 22343–22351. [CrossRef]
- Liu, N.; Wang, Z.; Liu, D.; Xie, P. HOXC13-AS-miR-122-5p-SATB1-C-Myc feedback loop promotes migration, invasion and EMT process in glioma. Onco Targets Ther. 2019, 12, 7165–7173.Liu, N.; Wang, Z.; Liu, D.; Xie, P. HOXC13-AS-miR-122-5p-SATB1-C-Myc feedback loop promotes migration, invasion and EMT process in glioma. Onco Targets Ther. 2019, 12, 7165–7173. [CrossRef]
- Zhou, J.F.; Shi, Y.T.; Wang, H.G.; Yang, X.Z.; Wu, S.N. Overexpression of long noncoding RNA HOXC13-AS and its prognostic significance in hepatocellular carcinoma. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 7369–7374.Zhou, J.F.; Shi, Y.T.;Wang, H.G.; Yang, X.Z.;Wu, S.N. Overexpression of long noncoding RNA HOXC13-AS and its prognostic significance in hepatocellular carcinoma. Eur. Rev. Med. Pharmacol. Sci. 2019, 23, 7369–7374. [PubMed]
- Li, W.; Zhu, Q.; Zhang, S.; Liu, L.; Zhang, H.; Zhu, D. HOXC13-AS accelerates cell proliferation and migration in oral squamous cell carcinoma via miR-378g/HOXC13 axis. Oral Oncol. 2020, 111, 104946.Li, W.; Zhu, Q.; Zhang, S.; Liu, L.; Zhang, H.; Zhu, D. HOXC13-AS accelerates cell proliferation and migration in oral squamous cell carcinoma via miR-378g/HOXC13 axis. Oral Oncol. 2020, 111, 104946. [CrossRef] [PubMed]
- Yarmishyn, A.A.; Batagov, A.O.; Tan, J.Z.; Sundaram, G.M.; Sampath, P.; Kuznetsov, V.A.; Kurochkin, I.V. HOXD-AS1 is a novel lncRNA encoded in HOXD cluster and a marker of neuroblastoma progression revealed via integrative analysis of noncoding transcriptome. BMC Genom. 2014, 15, S7.Yarmishyn, A.A.; Batagov, A.O.; Tan, J.Z.; Sundaram, G.M.; Sampath, P.; Kuznetsov, V.A.; Kurochkin, I.V. HOXD-AS1 is a novel lncRNA encoded in HOXD cluster and a marker of neuroblastoma progression revealed via integrative analysis of noncoding transcriptome. BMC Genom. 2014, 15, S7. [CrossRef] [PubMed]
- Li, J.; Zhuang, C.; Liu, Y.; Chen, M.; Chen, Y.; Chen, Z.; He, A.; Lin, J.; Zhan, Y.; Liu, L.; et al. Synthetic tetracycline-controllable shRNA targeting long non-coding RNA HOXD-AS1 inhibits the progression of bladder cancer. J. Exp. Clin. Cancer Res. 2016, 35, 99.Li, J.; Zhuang, C.; Liu, Y.; Chen, M.; Chen, Y.; Chen, Z.; He, A.; Lin, J.; Zhan, Y.; Liu, L.; et al. Synthetic tetracycline-controllable shRNA targeting long non-coding RNA HOXD-AS1 inhibits the progression of bladder cancer. J. Exp. Clin. Cancer Res. 2016, 35, 99. [CrossRef] [PubMed]
- Lu, C.; Ma, J.; Cai, D. Increased HAGLR expression promotes non-small cell lung cancer proliferation and invasion via enhanced de novo lipogenesis. Tumour Biol. 2017, 39.Lu, C.; Ma, J.; Cai, D. Increased HAGLR expression promotes non-small cell lung cancer proliferation and invasion via enhanced de novo lipogenesis. Tumour Biol. 2017, 39. [CrossRef]
- Zheng, L.; Chen, J.; Zhou, Z.; He, Z. Knockdown of long non-coding RNA HOXD-AS1 inhibits gastric cancer cell growth via inactivating the JAK2/STAT3 pathway. Tumour Biol. 2017, 39.Zheng, L.; Chen, J.; Zhou, Z.; He, Z. Knockdown of long non-coding RNA HOXD-AS1 inhibits gastric cancer cell growth via inactivating the JAK2/STAT3 pathway. Tumour Biol. 2017, 39. [CrossRef]
- Zhang, Y.; Dun, Y.; Zhou, S.; Huang, X.H. LncRNA HOXD-AS1 promotes epithelial ovarian cancer cells proliferation and invasion by targeting miR-133a-3p and activating Wnt/β-catenin signaling pathway. Biomed. Pharmacother. 2017, 96, 1216–1221.Zhang, Y.; Dun, Y.; Zhou, S.; Huang, X.H. LncRNA HOXD-AS1 promotes epithelial ovarian cancer cells proliferation and invasion by targeting miR-133a-3p and activating Wnt/_-catenin signaling pathway. Biomed. Pharmacother. 2017, 96, 1216–1221. [CrossRef]
- Gu, P.; Chen, X.; Xie, R.; Han, J.; Xie, W.; Wang, B.; Dong, W.; Chen, C.; Yang, M.; Jiang, J.; et al. lncRNA HOXD-AS1 Regulates Proliferation and Chemo-Resistance of Castration-Resistant Prostate Cancer via Recruiting WDR5. Mol. Ther. 2017, 25, 1959–1973.Gu, P.; Chen, X.; Xie, R.; Han, J.; Xie,W.;Wang, B.; Dong,W.; Chen, C.; Yang, M.; Jiang, J.; et al. lncRNA HOXD-AS1 Regulates Proliferation and Chemo-Resistance of Castration-Resistant Prostate Cancer via Recruiting WDR5. Mol. Ther. 2017, 25, 1959–1973. [CrossRef]
- Hu, Y.C.; Wang, A.M.; Lu, J.K.; Cen, R.; Liu, L.L. Long noncoding RNA HOXD-AS1 regulates proliferation of cervical cancer cells by activating Ras/ERK signaling pathway. Eur. Rev. Med. Pharmacol. Sci. 2017, 21, 5049–5055.
- Chen, Y.; Zhao, F.; Cui, D.; Jiang, R.; Chen, J.; Huang, Q.; Shi, J. HOXD-AS1/miR-130a sponge regulates glioma development by targeting E2F8. Int. J. Cancer 2018, 142, 2313–2322.Chen, Y.; Zhao, F.; Cui, D.; Jiang, R.; Chen, J.; Huang, Q.; Shi, J. HOXD-AS1/miR-130a sponge regulates glioma development by targeting E2F8. Int. J. Cancer 2018, 142, 2313–2322. [CrossRef]
- Gu, W.; Zhang, E.; Song, L.; Tu, L.; Wang, Z.; Tian, F.; Aikenmu, K.; Chu, G.; Zhao, J. Long noncoding RNA HOXD-AS1 aggravates osteosarcoma carcinogenesis through epigenetically inhibiting p57 via EZH2. Biomed. Pharmacother. 2018, 106, 890–895.Gu,W.; Zhang, E.; Song, L.; Tu, L.;Wang, Z.; Tian, F.; Aikenmu, K.; Chu, G.; Zhao, J. Long noncoding RNA HOXD-AS1 aggravates osteosarcoma carcinogenesis through epigenetically inhibiting p57 via EZH2. Biomed. Pharmacother. 2018, 106, 890–895. [CrossRef]
- Li, L.; Wang, Y.; Zhang, X.; Huang, Q.; Diao, Y.; Yin, H.; Liu, H. Long non-coding RNA HOXD-AS1 in cancer. Clin. Chim. Acta 2018, 487, 197–201.Li, L.;Wang, Y.; Zhang, X.; Huang, Q.; Diao, Y.; Yin, H.; Liu, H. Long non-coding RNA HOXD-AS1 in cancer. Clin. Chim. Acta 2018, 487, 197–201. [CrossRef] [PubMed]
- Xia, H.; Jing, H.; Li, Y.; Lv, X. Long noncoding RNA HOXD-AS1 promotes non-small cell lung cancer migration and invasion through regulating miR-133b/MMP9 axis. Biomed. Pharmacother. 2018, 106, 156–162.Xia, H.; Jing, H.; Li, Y.; Lv, X. Long noncoding RNA HOXD-AS1 promotes non-small cell lung cancer migration and invasion through regulating miR-133b/MMP9 axis. Biomed. Pharmacother. 2018, 106, 156–162. [CrossRef] [PubMed]
- Yang, M.H.; Zhao, L.; Wang, L.; Ou-Yang, W.; Hu, S.S.; Li, W.L.; Ai, M.L.; Wang, Y.Q.; Han, Y.; Li, T.T.; et al. Nuclear lncRNA HOXD-AS1 suppresses colorectal carcinoma growth and metastasis via inhibiting HOXD3-induced integrin β3 transcriptional activating and MAPK/AKT signalling. Mol. Cancer 2019, 18, 31.Yang, M.H.; Zhao, L.; Wang, L.; Ou-Yang, W.; Hu, S.S.; Li, W.L.; Ai, M.L.; Wang, Y.Q.; Han, Y.; Li, T.T.; et al. Nuclear lncRNA HOXD-AS1 suppresses colorectal carcinoma growth and metastasis via inhibiting HOXD3-induced integrin _3 transcriptional activating and MAPK/AKT signalling. Mol. Cancer 2019, 18, 31. [CrossRef] [PubMed]
- Sun, W.; Nie, W.; Wang, Z.; Zhang, H.; Li, Y.; Fang, X. Lnc HAGLR Promotes Colon Cancer Progression Through Sponging miR-185-5p and Activating CDK4 and CDK6 in vitro and in vivo. Onco Targets Ther. 2020, 13, 5913–5925.Sun, W.; Nie, W.; Wang, Z.; Zhang, H.; Li, Y.; Fang, X. Lnc HAGLR Promotes Colon Cancer Progression Through Sponging miR-185-5p and Activating CDK4 and CDK6 in vitro and in vivo. Onco Targets Ther. 2020, 13, 5913–5925. [CrossRef] [PubMed]
- Pan, L.X.; Ding, W. LncRNA HAGLR accelerates femoral neck fracture healing through negatively regulating miRNA-19a-3p. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 4080–4087.Pan, L.X.; Ding,W. LncRNA HAGLR accelerates femoral neck fracture healing through negatively regulating miRNA-19a-3p. Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 4080–4087. [PubMed]
- Bae, E.; Calhoun, V.C.; Levine, M.; Lewis, E.B.; Drewell, R.A. Characterization of the intergenic RNA profile at abdominal-A and Abdominal-B in the Drosophila bithorax complex. Proc. Natl. Acad. Sci. USA 2002, 99, 16847–16852.Bae, E.; Calhoun, V.C.; Levine, M.; Lewis, E.B.; Drewell, R.A. Characterization of the intergenic RNA profile at abdominal-A and Abdominal-B in the Drosophila bithorax complex. Proc. Natl. Acad. Sci. USA 2002, 99, 16847–16852. [CrossRef]
- Qi, Y.; Wang, Z.; Wu, F.; Yin, B.; Jiang, T.; Qiang, B.; Yuan, J.; Han, W.; Peng, X. Long noncoding RNA HOXD-AS2 regulates cell cycle to promote glioma progression. J. Cell Biochem. 2018.Qi, Y.; Wang, Z.; Wu, F.; Yin, B.; Jiang, T.; Qiang, B.; Yuan, J.; Han, W.; Peng, X. Long noncoding RNA HOXD-AS2 regulates cell cycle to promote glioma progression. J. Cell Biochem. 2018. [CrossRef]
- Yao, L.; Ye, P.C.; Tan, W.; Luo, Y.J.; Xiang, W.P.; Liu, Z.L.; Fu, Z.M.; Lu, F.; Tang, L.H.; Xiao, J.W. Decreased expression of the long non-coding RNA HOXD-AS2 promotes gastric cancer progression by targeting HOXD8 and activating PI3K/Akt signaling pathway. World J. Gastrointest. Oncol. 2020, 12, 1237–1254.Yao, L.; Ye, P.C.; Tan, W.; Luo, Y.J.; Xiang, W.P.; Liu, Z.L.; Fu, Z.M.; Lu, F.; Tang, L.H.; Xiao, J.W. Decreased expression of the long non-coding RNA HOXD-AS2 promotes gastric cancer progression by targeting HOXD8 and activating PI3K/Akt signaling pathway. World J. Gastrointest. Oncol. 2020, 12, 1237–1254. [CrossRef] [PubMed]
- Galis, F.; Van Dooren, T.J.; Feuth, J.D.; Metz, J.A.; Witkam, A.; Ruinard, S.; Steigenga, M.J.; Wijnaendts, L.C. Extreme selection in humans against homeotic transformations of cervical vertebrae. Evolution 2006, 60, 2643–2654.Galis, F.; Van Dooren, T.J.; Feuth, J.D.; Metz, J.A.; Witkam, A.; Ruinard, S.; Steigenga, M.J.; Wijnaendts, L.C. Extreme selection in humans against homeotic transformations of cervical vertebrae. Evolution 2006, 60, 2643–2654. [CrossRef] [PubMed]
- Yang, L.; Wang, Y.; Lu, Y.; Li, B.; Chen, K.; Li, C. Genome-wide Identification and Characterization of Long Non-coding RNAs in Tribolium castaneum. Insect Sci. 2020.Yang, L.; Wang, Y.; Lu, Y.; Li, B.; Chen, K.; Li, C. Genome-wide Identification and Characterization of Long Non-coding RNAs in Tribolium castaneum. Insect Sci. 2020. [CrossRef]
- Wu, Y.; Cheng, T.; Liu, C.; Liu, D.; Zhang, Q.; Long, R.; Zhao, P.; Xia, Q. Systematic Identification and Characterization of Long Non-Coding RNAs in the Silkworm, Bombyx mori. PLoS ONE 2016, 11, e0147147.Wu, Y.; Cheng, T.; Liu, C.; Liu, D.; Zhang, Q.; Long, R.; Zhao, P.; Xia, Q. Systematic Identification and Characterization of Long Non-Coding RNAs in the Silkworm, Bombyx mori. PLoS ONE 2016, 11, e0147147. [CrossRef] [PubMed]
- Zhang, X.; Yuan, J.; Sun, Y.; Li, S.; Gao, Y.; Yu, Y.; Liu, C.; Wang, Q.; Lv, X.; Zhang, X.; et al. Penaeid shrimp genome provides insights into benthic adaptation and frequent molting. Nat. Commun. 2019, 10, 356.Zhang, X.; Yuan, J.; Sun, Y.; Li, S.; Gao, Y.; Yu, Y.; Liu, C.; Wang, Q.; Lv, X.; Zhang, X.; et al. Penaeid shrimp genome provides insights into benthic adaptation and frequent molting. Nat. Commun. 2019, 10, 356. [CrossRef]
- Gutekunst, J.; Andriantsoa, R.; Falckenhayn, C.; Hanna, K.; Stein, W.; Rasamy, J.; Lyko, F. Clonal genome evolution and rapid invasive spread of the marbled crayfish. Nat. Ecol. Evol. 2018, 2, 567–573.Gutekunst, J.; Andriantsoa, R.; Falckenhayn, C.; Hanna, K.; Stein,W.; Rasamy, J.; Lyko, F. Clonal genome evolution and rapid invasive spread of the marbled crayfish. Nat. Ecol. Evol. 2018, 2, 567–573. [CrossRef]
- Tang, B.; Zhang, D.; Li, H.; Jiang, S.; Zhang, H.; Xuan, F.; Ge, B.; Wang, Z.; Liu, Y.; Sha, Z.; et al. Chromosome-level genome assembly reveals the unique genome evolution of the swimming crab (Portunus trituberculatus). Gigascience 2020, 9, giz161.Tang, B.; Zhang, D.; Li, H.; Jiang, S.; Zhang, H.; Xuan, F.; Ge, B.; Wang, Z.; Liu, Y.; Sha, Z.; et al. Chromosome-level genome assembly reveals the unique genome evolution of the swimming crab (Portunus trituberculatus). Gigascience 2020, 9, giz161. [CrossRef] [PubMed]
- Chebbi, M.A.; Becking, T.; Moumen, B.; Giraud, I.; Gilbert, C.; Peccoud, J.; Cordaux, R. The Genome of Armadillidium vulgare (Crustacea, Isopoda) Provides Insights into Sex Chromosome Evolution in the Context of Cytoplasmic Sex Determination. Mol. Biol. Evol. 2019, 36, 727–741.Chebbi, M.A.; Becking, T.; Moumen, B.; Giraud, I.; Gilbert, C.; Peccoud, J.; Cordaux, R. The Genome of Armadillidium vulgare (Crustacea, Isopoda) Provides Insights into Sex Chromosome Evolution in the Context of Cytoplasmic Sex Determination. Mol. Biol. Evol. 2019, 36, 727–741. [CrossRef] [PubMed]
- Thomas, G.W.C.; Dohmen, E.; Hughes, D.S.T.; Murali, S.C.; Poelchau, M.; Glastad, K.; Anstead, C.A.; Ayoub, N.A.; Batterham, P.; Bellair, M.; et al. Gene content evolution in the arthropods. Genome Biol. 2020, 21, 15.Thomas, G.W.C.; Dohmen, E.; Hughes, D.S.T.; Murali, S.C.; Poelchau, M.; Glastad, K.; Anstead, C.A.; Ayoub, N.A.; Batterham, P.; Bellair, M.; et al. Gene content evolution in the arthropods. Genome Biol. 2020, 21, 15. [CrossRef]
- Pang, Y.; Mao, C.; Liu, S. Encoding activities of non-coding RNAs. Theranostics 2018, 8, 2496–2507.Pang, Y.; Mao, C.; Liu, S. Encoding activities of non-coding RNAs. Theranostics 2018, 8, 2496–2507. [CrossRef]
- Mori, K.; Weng, S.M.; Arzberger, T.; May, S.; Rentzsch, K.; Kremmer, E.; Schmid, B.; Kretzschmar, H.A.; Cruts, M.; Van Broeckhoven, C.; et al. The C9orf72 GGGGCC repeat is translated into aggregating dipeptide-repeat proteins in FTLD/ALS. Science 2013, 339, 1335–1338.Mori, K.; Weng, S.M.; Arzberger, T.; May, S.; Rentzsch, K.; Kremmer, E.; Schmid, B.; Kretzschmar, H.A.; Cruts, M.; Van Broeckhoven, C.; et al. The C9orf72 GGGGCC repeat is translated into aggregating dipeptide-repeat proteins in FTLD/ALS. Science 2013, 339, 1335–1338. [CrossRef] [PubMed]
- Wang, X.Q.; Dostie, J. Reciprocal regulation of chromatin state and architecture by HOTAIRM1 contributes to temporal collinear HOXA gene activation. Nucleic Acids Res. 2017, 45, 1091–1104.Wang, X.Q.; Dostie, J. Reciprocal regulation of chromatin state and architecture by HOTAIRM1 contributes to temporal collinear HOXA gene activation. Nucleic Acids Res. 2017, 45, 1091–1104. [PubMed]
- Jenkins, A.M.; Waterhouse, R.M.; Muskavitch, M.A. Long non-coding RNA discovery across the genus anopheles reveals conserved secondary structures within and beyond the Gambiae complex. BMC Genom. 2015, 16, 337.Jenkins, A.M.; Waterhouse, R.M.; Muskavitch, M.A. Long non-coding RNA discovery across the genus anopheles reveals conserved secondary structures within and beyond the Gambiae complex. BMC Genom. 2015, 16, 337. [CrossRef] [PubMed]
- Legeai, F.; Derrien, T. Identification of long non-coding RNAs in insects genomes. Curr. Opin. Insect Sci. 2015, 7, 37–44.Legeai, F.; Derrien, T. Identification of long non-coding RNAs in insects genomes. Curr. Opin. Insect Sci. 2015, 7, 37–44. [CrossRef] [PubMed]
- Chang, Z.X.; Ajayi, O.E.; Guo, D.Y.; Wu, F. Genome-wide characterization and developmental expression profiling of long non-coding RNAs in Sogatella furcifera. Insect Sci. 2020, 27, 987–997.Chang, Z.X.; Ajayi, O.E.; Guo, D.Y.; Wu, F. Genome-wide characterization and developmental expression profiling of long non-coding RNAs in Sogatella furcifera. Insect Sci. 2020, 27, 987–997. [CrossRef]
- Guan, R.; Li, H.; Zhang, H.; An, S. Comparative analysis of dsRNA-induced lncRNAs in three kinds of insect species. Arch. Insect Biochem. Physiol. 2020, 103, e21640.Guan, R.; Li, H.; Zhang, H.; An, S. Comparative analysis of dsRNA-induced lncRNAs in three kinds of insect species. Arch. Insect Biochem. Physiol. 2020, 103, e21640. [CrossRef] [PubMed]
- Choudhary, C.; Sharma, S.; Meghwanshi, K.K.; Patel, S.; Mehta, P.; Shukla, N.; Do, D.N.; Rajpurohit, S.; Suravajhala, P.; Shukla, J.N. Long Non-Coding RNAs in Insects. Animals 2021, 11, 1118.Choudhary, C.; Sharma, S.; Meghwanshi, K.K.; Patel, S.; Mehta, P.; Shukla, N.; Do, D.N.; Rajpurohit, S.; Suravajhala, P.; Shukla, J.N. Long Non-Coding RNAs in Insects. Animals 2021, 11, 1118. [CrossRef]
- Ronshaugen, M.; Biemar, F.; Piel, J.; Levine, M.; Lai, E.C. The Drosophila microRNA iab-4 causes a dominant homeotic transformation of halteres to wings. Genes Dev. 2005, 19, 2947–2952.Ronshaugen, M.; Biemar, F.; Piel, J.; Levine, M.; Lai, E.C. The Drosophila microRNA iab-4 causes a dominant homeotic transformation of halteres to wings. Genes Dev. 2005, 19, 2947–2952. [CrossRef]
- Garaulet, D.L.; Lai, E.C. Hox miRNA regulation within the Drosophila Bithorax complex: Patterning behavior. Mech. Dev. 2015, 138, 151–159.Garaulet, D.L.; Lai, E.C. Hox miRNA regulation within the Drosophila Bithorax complex: Patterning behavior. Mech. Dev. 2015, 138, 151–159. [CrossRef]
- Hui, J.H.; Marco, A.; Hunt, S.; Melling, J.; Griffiths-Jones, S.; Ronshaugen, M. Structure, evolution and function of the bi-directionally transcribed iab-4/iab-8 microRNA locus in arthropods. Nucleic Acids Res. 2013, 41, 3352–3361.Hui, J.H.; Marco, A.; Hunt, S.; Melling, J.; Griffiths-Jones, S.; Ronshaugen, M. Structure, evolution and function of the bidirectionally transcribed iab-4/iab-8 microRNA locus in arthropods. Nucleic Acids Res. 2013, 41, 3352–3361. [CrossRef]
- Negre, B.; Ruiz, A. HOM-C evolution in Drosophila: Is there a need for Hox gene clustering? Trends Genet. 2007, 23, 55–59.Negre, B.; Ruiz, A. HOM-C evolution in Drosophila: Is there a need for Hox gene clustering? Trends Genet. 2007, 23, 55–59. [CrossRef] [PubMed]
- Shippy, T.D.; Ronshaugen, M.; Cande, J.; He, J.; Beeman, R.W.; Levine, M.; Brown, S.J.; Denell, R.E. Analysis of the Tribolium homeotic complex: Insights into mechanisms constraining insect Hox clusters. Dev. Genes Evol. 2008, 218, 127–139.Shippy, T.D.; Ronshaugen, M.; Cande, J.; He, J.; Beeman, R.W.; Levine, M.; Brown, S.J.; Denell, R.E. Analysis of the Tribolium homeotic complex: Insights into mechanisms constraining insect Hox clusters. Dev. Genes Evol. 2008, 218, 127–139. [CrossRef] [PubMed]
- Zeng, D.; Chen, X.; Peng, J.; Yang, C.; Peng, M.; Zhu, W.; Xie, D.; He, P.; Wei, P.; Lin, Y.; et al. Single-molecule long-read sequencing facilitates shrimp transcriptome research. Sci. Rep. 2018, 8, 16920.Zeng, D.; Chen, X.; Peng, J.; Yang, C.; Peng, M.; Zhu,W.; Xie, D.; He, P.;Wei, P.; Lin, Y.; et al. Single-molecule long-read sequencing facilitates shrimp transcriptome research. Sci. Rep. 2018, 8, 16920. [CrossRef] [PubMed]
- Wan, H.; Jia, X.; Zou, P.; Zhang, Z.; Wang, Y. The Single-molecule long-read sequencing of Scylla paramamosain. Sci. Rep. 2019, 9, 12401.Wan, H.; Jia, X.; Zou, P.; Zhang, Z.; Wang, Y. The Single-molecule long-read sequencing of Scylla paramamosain. Sci. Rep. 2019, 9, 12401. [CrossRef]
- Kim, H.S.; Kim, B.M.; Lee, B.Y.; Souissi, S.; Park, H.G.; Lee, J.S. Identification of Hox genes and rearrangements within the single homeobox (Hox) cluster (192.8 kb) of the cyclopoid copepod (Paracyclopina nana). J. Exp. Zool. B Mol. Dev. Evol. 2016, 326, 105–109.Kim, H.S.; Kim, B.M.; Lee, B.Y.; Souissi, S.; Park, H.G.; Lee, J.S. Identification of Hox genes and rearrangements within the single homeobox (Hox) cluster (192.8 kb) of the cyclopoid copepod (Paracyclopina nana). J. Exp. Zool. B Mol. Dev. Evol. 2016, 326, 105–109. [CrossRef]
- Kim, D.H.; Lee, B.Y.; Kim, H.S.; Jeong, C.B.; Hwang, D.S.; Kim, I.C.; Lee, J.S. Identification and characterization of homeobox (Hox) genes and conservation of the single Hox cluster (324.6 kb) in the water flea Daphnia magna. J. Exp. Zool. B Mol. Dev. Evol. 2018, 330, 76–82.Kim, D.H.; Lee, B.Y.; Kim, H.S.; Jeong, C.B.; Hwang, D.S.; Kim, I.C.; Lee, J.S. Identification and characterization of homeobox (Hox) genes and conservation of the single Hox cluster (324.6 kb) in the water flea Daphnia magna. J. Exp. Zool. B Mol. Dev. Evol. 2018, 330, 76–82. [CrossRef]
- Chipman, A.; Ferrier, D.; Brena, C.; Qu, J.; Hughes, D.S.T.; Schröder, R.; Torres-Oliva, M.; Znassi, N.; Jiang, H.; Almeida, F.; et al. The first myriapod genome sequence reveals conservative arthropod gene content and genome organisation in the centipede Strigamia maritima. PLoS Biol. 2014, 12, e1002005.Chipman, A.; Ferrier, D.; Brena, C.; Qu, J.; Hughes, D.S.T.; Schröder, R.; Torres-Oliva, M.; Znassi, N.; Jiang, H.; Almeida, F.; et al. The first myriapod genome sequence reveals conservative arthropod gene content and genome organisation in the centipede Strigamia maritima. PLoS Biol. 2014, 12, e1002005. [CrossRef] [PubMed]
- Qu, Z.; Nong, W.; So, W.L.; Barton-Owen, T.; Li, Y.; Leung, T.C.N.; Li, C.; Baril, T.; Wong, A.Y.P.; Swale, T.; et al. Millipede genomes reveal unique adaptations during myriapod evolution. PLoS Biol. 2020, 18, e3000636.Qu, Z.; Nong,W.; So,W.L.; Barton-Owen, T.; Li, Y.; Leung, T.C.N.; Li, C.; Baril, T.;Wong, A.Y.P.; Swale, T.; et al. Millipede genomes reveal unique adaptations during myriapod evolution. PLoS Biol. 2020, 18, e3000636. [CrossRef]
- Bakalenko, N.I.; Novikova, E.L.; Nesterenko, A.Y.; Kulakova, M.A. Hox gene expression during postlarval development of the polychaete Alitta virens. Evodevo 2013, 4, 13.Bakalenko, N.I.; Novikova, E.L.; Nesterenko, A.Y.; Kulakova, M.A. Hox gene expression during postlarval development of the polychaete Alitta virens. Evodevo 2013, 4, 13. [CrossRef] [PubMed]
- Novikova, E.L.; Bakalenko, N.I.; Nesterenko, A.Y.; Kulakova, M.A. Expression of Hox genes during regeneration of nereid polychaete Alitta (Nereis) virens (Annelida, Lophotrochozoa). Evodevo 2013, 4, 14.Novikova, E.L.; Bakalenko, N.I.; Nesterenko, A.Y.; Kulakova, M.A. Expression of Hox genes during regeneration of nereid polychaete Alitta (Nereis) virens (Annelida, Lophotrochozoa). Evodevo 2013, 4, 14. [CrossRef]
- Novikova, E.L.; Bakalenko, N.I.; Kulakova, M.A. Annelids win again: The first evidence of Hox antisense transcription in Spiralia. bioRxiv 2021.Novikova, E.L.; Bakalenko, N.I.; Kulakova, M.A. Annelids win again: The first evidence of Hox antisense transcription in Spiralia. bioRxiv 2021. [CrossRef]
- Andreeva, T.F.; Cook, C.; Korchagina, N.M.; Akam, M.; Dondya, A.K. Cloning and analysis of structural organization of Hox genes in the polychaete Nereis virens. Ontogenez 2001, 32, 225–233. (In Russian)Andreeva, T.F.; Cook, C.; Korchagina, N.M.; Akam, M.; Dondya, A.K. Cloning and analysis of structural organization of Hox genes in the polychaete Nereis virens. Ontogenez 2001, 32, 225–233. (In Russian) [PubMed]
- Deschamps, J.; Duboule, D. Embryonic timing, axial stem cells, chromatin dynamics, and the Hox clock. Genes Dev. 2017, 31, 1406–1416.Deschamps, J.; Duboule, D. Embryonic timing, axial stem cells, chromatin dynamics, and the Hox clock. Genes Dev. 2017, 31, 1406–1416. [CrossRef]
- Iimura, T.; Pourquié, O. Hox genes in time and space during vertebrate body formation. Dev. Growth Differ. 2007, 49, 265–275.Iimura, T.; Pourquié, O. Hox genes in time and space during vertebrate body formation. Dev. Growth Differ. 2007, 49, 265-275. [CrossRef]
- Hara, Y.; Yamaguchi, K.; Onimaru, K.; Kadota, M.; Koyanagi, M.; Keeley, S.D.; Tatsumi, K.; Tanaka, K.; Motone, F.; Kageyama, Y.; et al. Shark genomes provide insights into elasmobranch evolution and the origin of vertebrates. Nat. Ecol. Evol. 2018, 2, 1761–1771.Hara, Y.; Yamaguchi, K.; Onimaru, K.; Kadota, M.; Koyanagi, M.; Keeley, S.D.; Tatsumi, K.; Tanaka, K.; Motone, F.; Kageyama, Y.; et al. Shark genomes provide insights into elasmobranch evolution and the origin of vertebrates. Nat. Ecol. Evol. 2018, 2, 1761–1771. [CrossRef]
- Murray, S.C.; Mellor, J. Using both strands: The fundamental nature of antisense transcription. Bioarchitecture 2016, 6, 12–21.Murray, S.C.; Mellor, J. Using both strands: The fundamental nature of antisense transcription. Bioarchitecture 2016, 6, 12–21. [CrossRef] [PubMed]
- Xu, Z.; Wei, W.; Gagneur, J.; Clauder-Münster, S.; Smolik, M.; Huber, W.; Steinmetz, L.M. Antisense expression increases gene expression variability and locus interdependency. Mol. Syst. Biol. 2011, 7, 468.Xu, Z.;Wei,W.; Gagneur, J.; Clauder-Münster, S.; Smolik, M.; Huber,W.; Steinmetz, L.M. Antisense expression increases gene expression variability and locus interdependency. Mol. Syst. Biol. 2011, 7, 468. [CrossRef] [PubMed]
- Fusco, G. Trunk segment numbers and sequential segmentation in myriapods. Evol. Dev. 2005, 7, 608–617.Fusco, G. Trunk segment numbers and sequential segmentation in myriapods. Evol. Dev. 2005, 7, 608–617. [CrossRef]
