The bacterial flagellum is a complex and dynamic nanomachine that propels bacteria through liquids. It consists of a basal body, a hook, and a long filament. The flagellar filament is composed of thousands of copies of the protein flagellin (FliC) arranged helically and ending with a filament cap composed of an oligomer of the protein FliD. The overall structure of the filament core is preserved across bacterial species, while the outer domains exhibit high variability, and in some cases are even completely absent. Flagellar assembly is a complex and energetically costly process triggered by environmental stimuli and, accordingly, highly regulated on transcriptional, translational and post-translational levels. Apart from its role in locomotion, the filament is critically important in several other aspects of bacterial survival, reproduction and pathogenicity, such as adhesion to surfaces, secretion of virulence factors and formation of biofilms. Additionally, due to its ability to provoke potent immune responses, flagellins have a role as adjuvants in vaccine development.
- bacterial flagella
- flagellin
- filament
- FliD
- FliC
1. Flagellins
Bacillus cereus
Desulfotalea psychrophila [1][2]. In
E. coli
S.
E. coli
Salmonella mutants functional flagellins of 41 and 42 kDa have been reported [3][4]. Terminal regions of flagellins are highly homologous among species and they build the core of the filament [2]. Conversely, a hypervariable region in the middle of the polypeptide sequence makes outer domains significantly different, and in some bacteria such as
B.
subtilis it is completely absent [2][5].
fliC
E. coli
P. aeruginosa
Salmonella
hag
B. subtilis
flaA
C. jejuni
Helicobacter pylori
Magnetococus with 15 [6]. While the
E. coli
fliC
Salmonella
fliC
fljB [7]. However, only one of them is expressed at any point; the switch between them is called phase variation [8]. These two proteins diverge in the middle region providing distinct antigenicities. The studies on mutants locked in one of the two phases showed the different swimming properties and advantages of the FliC-expressing strains in colonization and infection of the gastrointestinal tract [9][10].
H. pylori
Campylobacter
Shewanella putrefaciens, both are present in the filament, the minor flagellin predominantly in the proximal region, while the major flagellin forms the remainder of the filament [11][12][13][14]. Although knocking out the minor flagellin does not usually affect the length of the filament, mutants exhibit changes in motility, especially in more viscous media, suggesting that having two flagellins in the filament provides optimal swimming characteristics in different environments [14][15]. The minor flagellin of
C. jejuni also plays a role in defense against bacteriophage infection [16].
P. aeruginosa has two types of flagellin genes, A and B, which differ in molecular weight and recognition by antibodies [17][18]. They are not, however, simultaneously present in the genome, but rather they are strain-specific with strains such as PAK containing type A FliC, while strains like PAO1 have type B FliC. While their N- and C-termini are highly conserved, the central region of the type A FliC is significantly shorter compared to the type B [19].
Caulobacter crescentus
Vibrio vulnificus
Rhizobium leguminosarum) [6][20][21]. In
V. vulnificus, mass spectrometry confirmed the presence of five distinct flagellins in the filament, with three having higher impact on motility, adhesion and cytotoxicity [20]. Some of these are redundant, such as for
C. crescentus in which five of six flagellins can make filaments alone [6].
1.1. Flagellin Structure
Early biophysical characterization showed that the N- and C-termini of monomeric flagellin are mainly disordered and critical for polymerization, although they become highly structured in the filament [22][23]. Due to the tendency of full-length flagellin to polymerize, crystallization of
S. Typhimurium flagellin FliC was achieved only after the removal of its terminal regions by limited proteolysis [24]. The overall shape of FliC obtained by combining X-ray crystallography and cryo-EM approaches revealed a structure in the form of a Greek letter Γ with four distinct domains (
Figure 1a) [25][26]. Starting from the N-terminus, the polypeptide chain makes a single helix that forms one half of an α-helical domain called D0, continues with the partial formation of domains D1 and D2, and forms domain D3 prior to completing the folds of the previous three domains. Thus, domains D0-D2 have N- and C-terminal moieties.
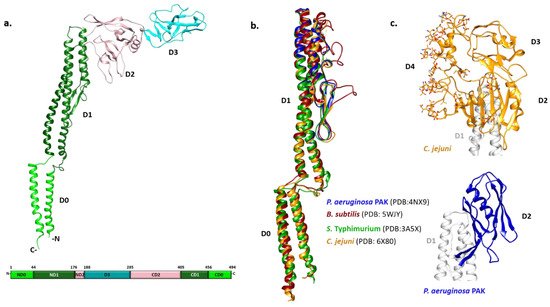
Figure 1.
a
S.
b
c
C. jejuni
P. aeruginosa
The N- and C-terminal helices of D0 form a coiled coil in the filament. The organization of D1 with three α-helices and a β-hairpin resembles a four-helix bundle with an extensive hydrophobic core. D2 and D3 consist primarily of β-strands organized in a specific fold called β-foliums in which the tips of the β-hairpins are bent or twisted [25][26].
Figure 1
P. aeruginosa
S. Typhimurium [27]. Conversely, the D2 domain adopts a different fold with two β-sheets and one α-helix between them forming a less flexible cup-like structure positioned parallel to the D1 domain, instead of pointing away from it as in
Salmonella
Figure 1
Campylobacter
Figure 1c) [28]. The D2 and D3 domains of the
Campylobacter
Pseudomonas
1.2. Posttranslational Modifications of Flagellins
Pseudomonas type A flagellin was glycosylated [19]. A genomic island between
flgL
fliC genes containing 14 open reading frames (ORFs) is responsible for glycosylation and it specifically modifies type A flagellin, while having no effect on type B [29]. These glycan chains in the PAK strain are
O-linked to residues T189 and S260 through rhamnose and vary in length (up to 11 monosaccharides) and composition [30].
O-linked glycans at serine residues 191 and 195, although in this case it is a much simpler modification with a single monosaccharide [31]. The genomic island responsible for type B glycosylation is smaller than its type A counterpart and contains four ORFs.
Pseudomonas type A and B mutants in which glycosylation is absent or incomplete do not affect motility in general, although the lack of glycosylation significantly decreased virulence of both PAK and PAO1 strains in a burned-mouse model of infection [32]. Apart from
O
N
P. aeruginosa strain PA14, with three sites on the D0 and D1 domains proposed as the most likely candidate positions [33].
Campylobacter
O-linked pseudaminic acid or related derivatives [34][35][36]. Glycosylation in
C. jejuni
flaA
flaB
Campylobacter glycosylation machinery [37]. Glycosylation of flagellin in
Campylobacter is necessary for filament assembly as mutants that produce unglycosylated flagellin do not have filaments and flagellin accumulates intracellularly [38].
H. pylori flagellins are also glycosylated with pseudaminic acid with most glycosylation sites in the central core region of the protein [39]. As in
Campylobacter, glycosylation plays an important role during assembly of the filament; abolishing glycosylation leads to loss of the filament and motility, even though the levels of flagellin mRNA are unaffected [39].
Listeria monocytogenes
O
N-acetylglucosamines at up to six sites located in the central surface exposed region of the protein [30]. The flagellin of
P. syringae is also glycosylated and these modifications are important for its virulence and swarming [40][41].
S. Typhimurium, the presence of methylated lysines was reported as early as 1959 [42]. Lysine methylation is performed by the enzyme FliB. Abolishing methylation does not affect filament assembly or motility [43]. However, it was shown that it promotes adhesion and host cell invasion [44].
N
Shewanella oneidensis [45].
2. Filament
The filament is a tubular structure made of more than 20,000 copies of the flagellin protein arranged in a helical fashion [46]. Flagellin is rotated and translated 11 times along the screw axis, making two turns, before it reaches its original position. Such an arrangement results in 11 stacks of flagellin along the axis called protofilaments, which are the basic functional units of the filament (
Figure 2
Figure 2b) [47]. During swimming, native filaments adopt a superhelical or corkscrew shape and their rotation creates thrust. Depending on the number of L- and R-protofilaments, the pitch of the superhelix changes, modulating the swimming properties [48]. If all protofilaments are in the L- or R-state, the filament is straight and cannot create a thrust and, thus, cells are immotile. Apart from handedness, L- and R-filament also differ in length, with the R-filament being 1.5% shorter. During the normal smooth swimming of
S.
E. coli, two R- and nine L-protofilaments form a filament that is a left-handed superhelix with a pitch of around 2–3 µm. The increased number of R-protofilaments eventually results in right-handed curly flagella with a shorter pitch typical for tumbling movements [48].
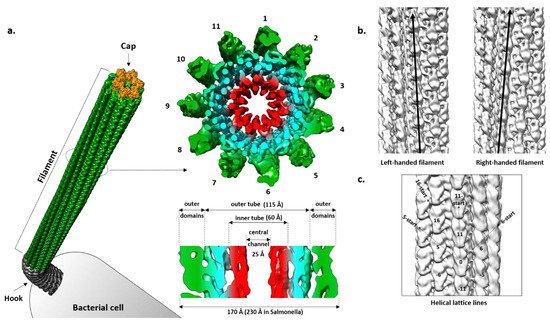
Figure 2.
P. aeruginosa
a
b
c
Straight filaments in which all protofilaments are locked in either L- or R-state are much more amenable for structural studies than the native wavy forms in which, due to the presence of a mixture of L- and R-protofilaments, the strict helical symmetry is disturbed [49]. Several mutations of the
fliC
Salmonella result in formation of straight filaments and were used in X-ray and EM studies [47]. Additionally, these mutations served as a basis to design the equivalent L- and R-mutants in other bacteria like
P. aeruginosa
C. jejuni and to study their structures [28][50].
Salmonella filament reveals a densely packed helical core 115 Å in diameter around a central channel that is ~25 Å wide, and an outer region exposed to the surface where domains of the individual molecules are clearly separated [51][52]. The diameter of the whole filament is ~230 Å. The core is organized into two tubes, inner and outer, connected through eleven short spokes, while the outer region contains two domains that are clearly defined.
Using cryo-EM, Yonekura et al. [26] obtained an electron density map of ~4 Å resolution that allowed them, in combination with the already known structure of FliC, to build the first complete atomic model of the R-filament from
Salmonella
Salmonella R-filament is a helical structure in which the FliC molecule is rotated 65.8° to the right followed by the translation of 4.7 Å. Protofilaments are tilted to the right by 3.5° relative to the longitudinal axis. The inner and outer tube are made of the D0 and D1 domains, respectively. In the inner tube, interactions between subunits are found along 11-start and 5-start helices and they are mostly hydrophobic, stabilizing the filament. The outer tube has a similar pattern with interactions found in the 11-start, 5-start and 16-start helices but they are predominantly polar–polar and charge–polar. Along the 11-start helix, the D1 domain of the upper subunit forms a concave surface that is complementary to the convex surface of the lower subunit and the two neighboring molecules make numerous Van der Waals contacts [25]. In the 5-start direction, the N-terminal helices of one molecule interact with the β-hairpin and C-terminal helix of the other molecule. N-terminal helices also make contact with the spoke in the 16-start direction. According to this rigid model, except for the small region of D2 proximal to the core, D2 and D3 do not engage in obvious inter-molecular contacts and they are projected away from the core. However, they do contribute to the overall stability of the filament, since deletion of the large part of this region makes the filament much more fragile in comparison to the wild type and mutant filaments, showing a contact between the adjacent outer domains along the 5-start helix that is absent in the mutant [4][53]. Therefore, it is likely that D2 and D3 exhibit flexibility in vivo, transiently interacting with the neighboring subunits and contributing to the filament integrity.
B. subtilis
P. aeruginosa appear as a single domain in which the D3 portion extends along the axis and seems to form a dimer with the D2 of the subunit above, although the structure could not be resolved due to low resolution in that region [50]. Another distinctive feature of the
Pseudomonas
Rhizobium lupini
Pseudomonas rhodos
P. aeruginosa it is present only at the level of the outer domains [50]. As mentioned previously, the outer region in
Campylobacter is significantly larger containing 3 domains and, apart from interactions along the protofilament, it also makes contacts in 5-start and 6-start directions, contributing to filament integrity [28].
2.1. Comparison of L- and R-Filaments
One of the questions that draws much attention is the structural basis for the polymorphic switching of the filament. Comparison of the L- and R-structures obtained by cryo-EM revealed that the D0 domains align well with some local changes in conformation, while the rest of the molecule is twisted by 5° in the counterclockwise direction [54]. Previous X-ray diffraction study on L- and R-types from
Salmonella showed that there were no changes in helical parameters in the inner region of the core made of D0 domains [55]. However, the two types differ in the outer regions and the distance between two subunits in the protofilament is 0.8 Å shorter in the R-type compared to the L-type. Maki-Yonekura et al. [54] found that the packing of the hydrophobic side chains at the lower portion of D1 is different between L- and R-type and that the intersubunit interactions along the 5-start helix are important in stabilizing the conformation. Based on molecular dynamics simulations of the polymorphic supercoiling mechanism, Kitao et al. [56] identified three types of interaction between subunits: permanent (the same pair of residues in different states), sliding (the same types of interaction with variable partners) and switch interactions that were responsible for locking the protofilament interface in R- or L-handed state. The switch interactions were found only along the 5-start helices. However, this was not supported by the analysis of the corresponding interactions in L- and R-filaments of
B. subtilis [50].
2.2. Filament Cap
Salmonella
While they vary in size, FliDs from different bacteria have conserved N- and C-terminal regions predicted to be coiled-coils and are largely unstructured in the monomeric state [57]. The first high-resolution structure of the
P. aeruginosa FliD showed that this 50 kDa protein has three distinct domains: a highly flexible D1 domain that corresponds to the leg domains seen in the cap structure, and compact D2 and D3 domains that comprise the head [58]. Much as in flagellin, a continuous sequence in the middle of the polypeptide folds into D3 while domains D1 and D2 consist of N- and C-terminal portions of the chain relative to D3. Domains D2 and D3 are each made of two antiparallel β-sheets, while D1 could not be resolved with confidence apart from one helix. The structure of the
E. coli
Figure 3a,c) [59]. These structures, together with the ones from
Salmonella
Serratia marcescens [59][60], indicated that the structural organization of the capping proteins among different bacteria is preserved. However, not all FliDs are of the same size. For example, the FliD of
H. pylori
Figure 3b) [61]. These domains may be related to the specific role FliD plays as a virulence factor during
H. pylori infection; stronger antigenicity of D4 and D5 further supports this [61].
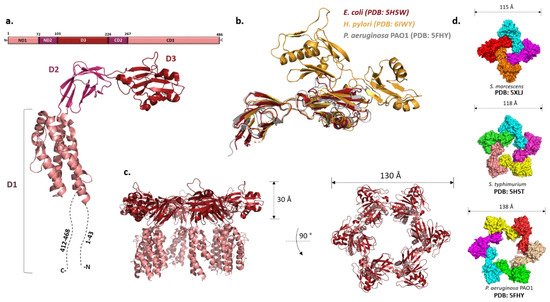
Figure 3.
a
E. coli
b
H. pylori
c
E. coli
d
Salmonella
C. jejuni cap supports it [62]. However, both
Pseudomonas
E. coli
Serratia
Figure 3d) [58][59][60]. Moreover, tetramers and pentamers of
Pseudomonas
Salmonella FliD were observed in vitro [58][62]. While these stoichiometries could arise due to non-physiological conditions, in the case of
P. aeruginosa hexamers were also detected in vivo [58].
3. Protein Synthesis, Export and Filament Elongation
3.1. Genetic Regulation of FliC and FliD
The genetic organization of the flagellar components is highly complex and relatively well conserved across bacteria with the same type of flagellar arrangement (i.e., peritrhichous, monotrichous/polar, etc.) but quite different between them [63][64][65]. The bacterial flagellar regulon commonly consists of dozens of genes grouped in several operons, encoding all structural proteins of the flagellum, the chemosensory apparatus and the regulators that control gene expression. For example, there are about 50 genes involved in the synthesis of the polar
P. aeruginosa
fla regulon [66][67]. On the other hand, in peritrichous bacteria such as
Salmonella
Escherichia coli, the flagellar regulon contains 70 genes distributed in at least 25 operons [43][64]. The transcription of these genes is hierarchical and controlled by different promoter classes that are temporally regulated (
Figure 4
Table 1
28
Actinoplanes missouriensis zoospore in which there is no hierarchical coordination and 33 flagellar genes are transcribed simultaneously during the sporangium formation [68]. The filament cap gene
fliD
fliD is expressed as a class II gene, it is not assembled into the growing flagellar structure until the hook and the junction proteins are expressed from class III (including flagellin genes) and incorporated into the flagellar organelle [69]. Hence, this cascade serves to control the timing of gene expression to coincide with the assembly of the flagellar apparatus and filament.
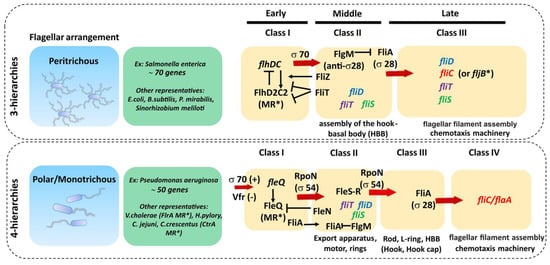
Figure 4.
Table 1.
| Protein | Function | Sigma Factor | Class in Flagellar Regulatory Hierarchy | Transcriptional Expression Regulators | Post-Transcriptional Regulators | Specific Secretion Chaperone | |
|---|---|---|---|---|---|---|---|
| 3-Tiered | 4-Tiered | ||||||
| Flagellin (FliC, FlaA) | Structural component of the filament | Sigma28 Sigma54 Sigma43 |
Class III gene | Class IV gene | CodY Environmental factors (nutrients, c-di-GMP, ppGpp, BCAA, temperature…) |
Self-regulating CsrA-FliW |
FliS |
| FliD | Filament cap | Sigma70 Sigma28 Sigma54 |
Class II and III | Class II | Cognate flagellin | ? | FliT |
S.
fliC
fljB
fljB
fljB
fljA, a transcriptional repressor that inhibits the expression of FliC [70].
Salmonella is bimodal, meaning that in a population of genetically identical cells under the same conditions only a fraction of them is motile [71][72]. This bimodality is present at both the class II and class III levels and is governed by separate mechanisms. While a double negative feedback loop involving two flagellar regulatory proteins, RflP and FliZ, controls the expression of the class II genes in response to nutrient availability, class III gene expression is tuned by the secretion of FlgM and there is a minimum number of HBBs necessary for the cell to pass this checkpoint.
Figure 4) [66]. Gene regulation in
Pseudomonas
54
Salmonella
28
Pseudomonas independently of other flagellar genes, which makes it one of the class I genes [66].
Helicobacter
Campylobacter
28
54 [73][74]. Although the major flagellin is usually under the control of σ
28
V. cholera
S. oneidensis
54
Eubacterium
Roseburia
28
43 [75][76]. In the phylogenetically related
Butyrivibrio fibrisolvens
fliC gene is driven from two different promoters, yielding two transcripts with alternative transcription start-sites [76].
In some Gram-positive bacteria, changes in the environmental conditions such as nutrient limitation induce variations in the levels of the intracellular messengers guanosine tetra/pentaphosphate, (p)ppGpp, guanosine nucleoside triphosphate (GTP) and branched chain amino-acid pools. These variations are detected by a conserved GTP-sensing protein CodY and a global regulator CsrA that modulate flagellin expression. Cell motility can be repressed under elevated intracellular cyclic-di-GMP levels, by impeding transcription of some of the flagellum genes [77][78][79][80][81]. Indeed, the expression of
fliD
fliC is repressed in phosphodiesterase 3 (PDE3) knockout mutants triggered by elevated c-di-GMP accumulation [82].
Because the presence of flagellin can be deleterious to the bacterium on account of inducing host immunity, flagellated bacteria downregulate (or turn off) flagellin expression during the host invasion to avoid host immune responses. Therefore, normal microbiota within the healthy adult mammalian gut has been shown to have overall relatively low levels of flagellin expression, while TLR5−/− mice exhibited a diversity of gut microbiome members with overexpressed flagellar genes [83][84].
E. coli
S. Typhimurium strains strongly down-regulate their genes coding for flagellar machinery and lose their motility once inside the host [85][86]. Under environmental temperature conditions (22–30 °C), the expression of flagellar genes in the human pathogens
Listeria monocytogenes
Legionela pneumophila is normal, but significantly reduced when raised to 37 °C, revealing temperature-dependent transcription mainly controlled by the protein GmaR, which acts as a protein expression thermostat [87][88]. These types of system provide a pathogen with the ability to turn off immune-stimulating antigens before they trigger adverse host defense mechanisms once inside their target host.
Regulation of flagellin expression also occurs on the post-transcriptional level by proteins that bind to the untranslated leader region of the flagellin mRNA, affecting transcript stability and/or ribosome access [89][90][91]. For instance, the RNA-binding protein CsrA of
B. burgdorferi specifically mediates synthesis of the major flagellin by inhibiting translation initiation of its transcript. In some cases, post-transcriptional regulators repress the accumulation of flagellin when cells are defective for flagellar hook assembly [92][93].
B. subtilis. This is governed by a homeostatic mechanism in which the flagellin protein itself is a critical regulator. In a partner switching mechanism, the flagellar assembly factor FliW binds flagellin, while a global regulator CsrA binds flagellin mRNA, repressing its translation. After completion of the hook, flagellin is secreted, and the released FliW binds CsrA, thus derepressing flagellin translation [94]. An interesting case is
A. missouriensis, in which all flagellar genes are transcribed simultaneously during sporangium formation and there is no checkpoint mechanism in the process of flagellar gene transcription to optimize the efficiency of the flagellar assembly [68]. The post-transcriptional level of regulation between protein production and assembly could play an important role, although this remains unknown.
3.2. Chaperones
Accumulation and premature oligomerization of flagellar axial proteins in the cytosol could be wasteful and detrimental to the cell. For this reason, bacteria encode flagellar-specific chaperones dedicated to facilitating the assembly and export of the flagellum components mainly by protecting and/or preventing their cognate flagellar protein substrates from aggregation or avoiding premature undesired interactions of flagellar proteins in the cytoplasm prior to interaction with the export gate [95]. Due to the small diameter of the flagellar export central channel (20–30 Å), the proteins destined for incorporation into the growing flagellum must be exported in a partially or completely unfolded state, implying that premature folding and oligomerization in the cytosol must be prevented to keep them in a secretion-competent conformation for flagellum assembly [96]. Hence, the external flagellar components require T3SS-specific chaperones to facilitate their efficient export [64].
3.2.1. FliS—Flagellin Chaperone
FliS acts as a flagellin-specific T3SS chaperone of FliC, preventing premature folding and inappropriate interaction of newly synthesized flagellin subunits in the cytosol, thus facilitating its export and polymerization upon completion of the HBB assembly. Yeast two-hybrid assays indicated that the C-terminal disordered region of flagellin is essential for FliS binding and, accordingly, spontaneous mutations causing flagellin accumulation in the cytoplasm map to the C-terminal region of FliC [97]. Thermodynamic experiments indicated that FliS does not function as an anti-folding factor keeping flagellin in a secretion-competent conformation. Instead, FliS binding stabilizes the flagellin conformation through formation of the α-helical secondary structure in the last 40 C-terminal residues of FliC (residues 454–494) [70][96][98].
Aquifex aeolicus FliS (AaFliS), revealed a novel, mainly α-helical fold, different from those of the T3SS chaperones [99]. The structure of
Aa
S. typhimurium cell extracts subjected to gel-filtration chromatography, suggesting that this interaction may be transient in vivo. Such rapid chaperone dissociation would favor the subsequent export of FliC [100].
The interaction between FliS chaperone and FlhA is a key step preceding the efficient transfer of FliC to the platform of the flagellar type III export apparatus for a rapid export during flagellar filament assembly [101][102][103]. Filament proteins in complex with their cognate chaperones bind to a highly conserved hydrophobic pocket of FlhA
C
C for the chaperone substrate complexes may be the key to defining the correct order of protein export among the filament-type proteins [104]. Recently, a structure of the ternary complex formed by FliC, FliS and the export gate protein FlhA revealed that FliC does not interact directly with FlhA (
Figure 5). Instead, the presence of FliC induces a binding-competent conformation of FliS that exposes the motif which is specifically recognized by FlhA [105]. Moreover, SAXS and HDX-MS experiments showed the formation of a heterotrimeric FliC-FliS-FliW complex that interacts with FlhA suggesting that FliS and FliW are released during flagellin export. FliW and FliS bind to opposing interfaces located at the N- and C-termini of flagellin, respectively, and these proteins seems to synchronize the production of flagellin with the capacity of the T3SS to secrete flagellin [106].
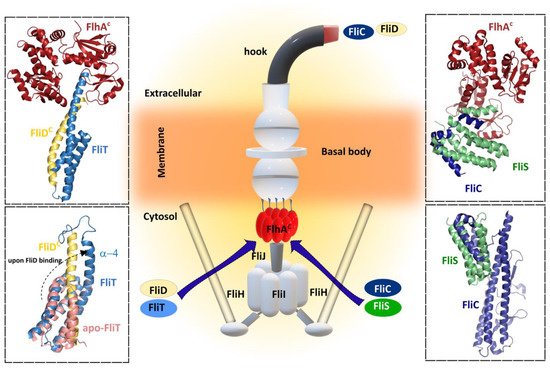
Figure 5.
C
Upper left
Salmonella enterica
enterica
Upper right
Salmonella enterica
enterica
Lower left
Salmonella enterica
enterica
Lower right
B. subtilis
Center figure
Salmonella. The loss of FliS results in a short filament phenotype despite high expression levels of FliC, which is explained by the increase in the secretion level of FlgM [101]. Bypass mutants have been isolated from a
Salmonella ΔfliS mutant, and all those mutations were identified in FliC [107][108].
S. typhimurium
Yersinia pseudotuberculosis, respectively [100][109]. Using a number of different approaches, they showed that these proteins specifically interact to form a 1:1 complex, and that this interaction protects FlgM from proteolysis. FliS acts as a negative inhibitor of FlgM secretion, keeping this intrinsically disordered protein stable before FliA is expressed in cells. The binding site of FliA on FlgM is close to or even overlaps with the binding site of FliS, suggesting that FliA binding removes FliS from the complex. In addition, FliS from
S. typhimurium
3.2.2. FliT—Chaperone of FliD
For the correct formation of the filament, a filament-capping protein FliD should be exported first during the filament assembly to form a penta- or hexameric cap that promotes self-assembly of FliC. In order to do this, FliD requires the assistance of its chaperon FliT [110]. FliT act as key flagellar chaperone in the assembly and operation of the flagellum because it binds to several flagellar proteins in the cytoplasm, such as the export apparatus components FliI, FliJ, and FlhA, beyond interaction with its cognate FliD. As an example of its versatility, FliT also functions as a negative transcriptional regulator of flagellar genes by inhibiting the formation of a DNA complex with the master regulator FlhDC. Lately, several crystal structures of FliT alone and in complex with FliD or FliI have become available [95][110]. Although in the crystal structure of
Salmonella
Figure 5) [105].
