Recent reports suggest a link between positive regulation of the Hippo pathway with bipolar disorder (BD), and the Hippo pathway is known to interact with multiple other signaling pathways previously associated with BD and other psychiatric disorders. In this study, neuronal-like NT2 cells were treated with amisulpride (10 µM), aripiprazole (0.1 µM), clozapine (10 µM), lamotrigine (50 µM), lithium (2.5 mM), quetiapine (50 µM), risperidone (0.1 µM), valproate (0.5 mM), or vehicle control for 24 h. Genome-wide mRNA expression was quantified and analyzed using gene set enrichment analysis (GSEA), with genes belonging to Hippo, Wnt, Notch, TGF- β, and Hedgehog retrieved from the KEGG database. Five of the eight drugs downregulated the genes of the Hippo pathway and modulated several genes involved in the interacting pathways. We speculate that the regulation of these genes, especially by aripiprazole, clozapine, and quetiapine, results in a reduction of MAPK and NFκB pro-inflammatory signaling through modulation of Hippo, Wnt, and TGF-β pathways. We also employed connectivity map analysis to identify compounds that act on these pathways in a similar manner to the known psychiatric drugs. Thirty-six compounds were identified. The presence of antidepressants and antipsychotics validates our approach and reveals possible new targets for drug repurposing
1. Introduction
The Hippo pathway is a signaling cascade that integrates a broad range of different biological, chemical, and mechanical cues to control several cellular processes through its downstream effectors YAP (yes-associated protein) and transcriptional co-activator with PDZ-binding motif (TAZ)
[1][2][1,2]. It was first discovered in Drosophila melanogaster, where a genetic mutation in one of its core components (Hippo/Hpo) was associated with tissue overgrowth of the eyes, wings, and limbs, a phenotype that gave name to the pathway
[1]. The Hippo pathway regulates cell proliferation, differentiation, and spatial patterning governing organ size, tissue homeostasis, and regeneration, and is highly conserved from Drosophila to mammals
[1][2][1,2].
The canonical Hippo signaling pathway is a key regulator of organ size and tissue remodeling
[2]. The non-canonical Hippo signaling pathway, due to its diverse interplay with a variety of signaling cascades, i.e., TGF-β, Wnt, Hedgehog (HH), and Notch signaling pathways
[3], has been associated with diversified mechanisms according to different microenvironments
[4]: neurogenesis
[5], neuronal development
[6], neuronal dendritic field formation
[7][8][7,8], and, more recently, neuroinflammation
[9] and immunology
[10]. In the central nervous system, the Hippo pathway has been associated with the response to neuroinflammation
[11], dendrite development and organization
[7], the balance between apoptosis and proliferation of the neural progenitor cell pool
[12], glioma proliferation
[13], Huntington’s disease
[14], and Alzheimer’s disease
[15].
Due to its role in both healthy and pathologic processes in the central nervous system and its crosstalk with other signaling pathways, it is unsurprising that the genes involved in the Hippo pathway have recently been associated with various psychiatric conditions
[16][17][18][19][16,17,18,19]. Indeed, Liu and colleagues
[18], identified positive regulation in the transcription of Hippo pathway genes in post-mortem prefrontal cortex of bipolar disorder (BD) patients as significantly enriched compared to healthy controls. Together with 30 hub genes, including YAP, the authors suggested that this pathway might have important implications in understanding the pathophysiology of BD, and could be a source of new targets for treatment. Further, the Hippo pathway genes were also reported as hypermethylated in a twin affected with BD compared with the non-affected twin
[16]. Although the Hippo pathway was not differentially methylated in a second pair of twins, the authors proposed that the patient-specific differences might reflect the effects of antipsychotic medications, resulting in hyper/hypomethylation differences. In addition, different components of the Hippo pathway appear to be targeted by chlorpromazine, an antipsychotic used to treat schizophrenia (SZ), leading to apoptosis in cancer cells
[20]. Valproic acid used to treat BD is reported to interact with the Hippo pathway through RASSF1A in myeloid leukemia, allowing YAP to associate with p73 and induce the expression of pro-apoptotic genes
[21].
Therefore, this study aimed to evaluate the effects of commonly prescribed psychoactive drugs used in treating affective disorders (BD and SZ) on the expression of genes in the Hippo pathway. We expect these medications to downregulate the genes involved in the Hippo pathway, as upregulation of the Hippo pathway was previously observed in people with BD compared to healthy controls
[18].
2. GSEA
The list of genes involved in the Hippo pathway was extracted from the KEGG database, and the acute effects of the eight drugs on these genes was analyzed by GSEA. Five of the eight drugs, amisulpride, aripiprazole, clozapine, quetiapine, and risperidone, significantly downregulated genes in the Hippo pathway, as shown in
Table 1.
Table 1. Effects of psychoactive drugs on KEGG Hippo pathway gene expression.
| Drug |
ES |
NES |
p | -Value |
Wnt | q | -Value |
| Notch |
Hedgehog |
TGF-β |
| |
LogFC |
p-Value |
LogFC |
p-Value |
LogFC |
p-Value |
LogFC |
p-Value |
| |
Mean |
SEM |
Mean |
SEM |
Mean |
SEM |
Mean |
SEM |
| Clozapine |
−0.61 |
−2.09 |
0.00013 |
0.0012 |
| Aripiprazole |
−0.54 |
| Amisulpride |
−0.13 |
0.04 |
0.0018 |
−0.08 |
0.03 |
0.0032 |
−0.03 |
0.02 |
0.18 |
−0.08 |
0.05 |
0.19 |
| −2.04 |
0.00015 |
| Aripiprazole |
−0.21 |
0.04 | 0.0044 |
| 4.80 × 10 | −6 |
−0.07 |
0.03 |
0.043 |
−0.04 |
0.02 |
0.07 |
−0.05 |
0.1 |
0.66 |
Risperidone |
−0.49 |
−1.92 |
0.00024 |
0.0049 |
| Clozapine |
−0.11 |
0.03 |
0.0013 |
−0.08 |
0.03 |
0.0033 |
−0.05 |
0.02 |
0.025 |
−0.21 |
0.08 |
0.07 |
Quetiapine |
−0.60 |
−1.79 |
0.00027 |
0.0052 |
| Quetiapine |
−0.02 |
0.05 |
0.64 |
0.02 |
0.05 |
0.63 |
−0.04 |
0.03 |
0.26 |
−0.27 |
0.15 |
Amisulpride |
−0.45 |
−1.61 |
0.00085 |
0.016 |
| Lithium |
−0.40 |
−1.32 |
0.048 |
0.21 |
| Lamotrigine |
−0.29 |
−1.15 |
0.14 |
0.35 |
| Valproate |
0.31 |
0.89 |
0.67 |
0.66 |
Medications listed based on the p-value, lowest to highest. Abbreviations: ES = enrichment score; NES = normalized enrichment score.
3. Hippo Core Genes and Transcription Factors
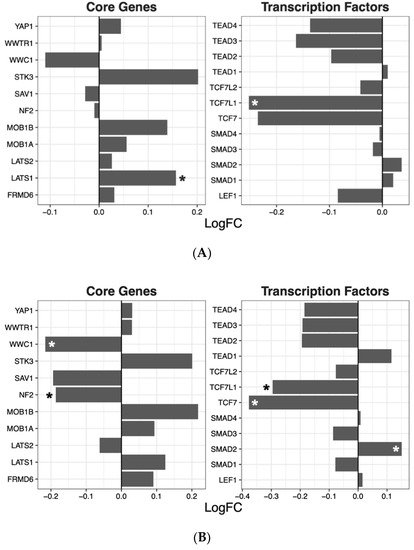
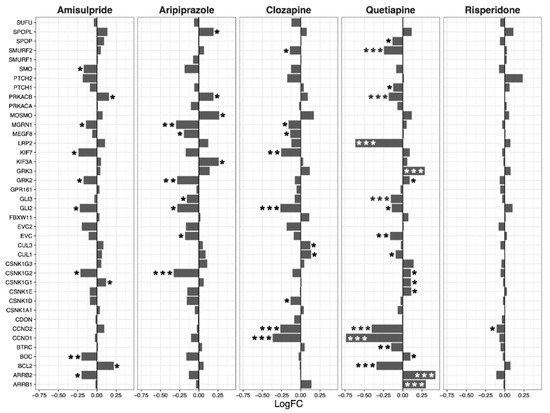
Having ascertained that five of the psychoactive drugs significantly altered the expression of genes in the Hippo pathway, we further investigated the effects of these drugs on a smaller set of genes. This smaller set of genes included the core genes of the Hippo pathway and its transcription factors, which are the critical effectors of the pathway (
Figure 1 A–E).
Figure 1. Effects of five drugs in the core genes and transcriptional factors of the Hippo pathway. (A) Amisulpride. (B) Aripiprazole. (C) Clozapine. (D) Quetiapine. (E) Risperidone * p < 0.05, ** p < 0.005, and *** p < 0.001.
Amisulpride, aripiprazole, clozapine, and quetiapine significantly reduced the expression of the transcription factors involved in the Hippo pathway (p = 0.007, p = 0.025, p = 0.001, and p = 0.008, respectively). Risperidone and quetiapine downregulated the core genes of the Hippo pathway (p= 0.03 and p = 0.02, respectively), and amisulpride (p = 0.05) showed a tendency to downregulate the core genes of the Hippo pathway.
4. Interacting Pathways
The Hippo signaling pathway interacts extensively with other closely related signaling pathways, including the TGF-β, WNT, Notch, and Hedgehog signaling pathways. Therefore, the effects of these drugs were further investigated on these pathways (
Table 2).
Table 2. The effects of the drugs on genes involved in interacting pathways.
| 0.15 |
| Risperidone |
−0.06 |
0.02 |
0.02 |
−0.01 |
0.02 |
0.72 |
0 |
0.01 |
0.67 |
−0.05 |
0.02 |
0.06 |
Abbreviations: LogFC= logarithmic fold change; SEM = standard error of the mean.
The Wnt signaling pathway was transcriptionally downregulated by amisulpride (p = 0.0018), aripiprazole (p = 4.80 × 10
−6), clozapine (p = 0.0013), and risperidone (p = 0.02;
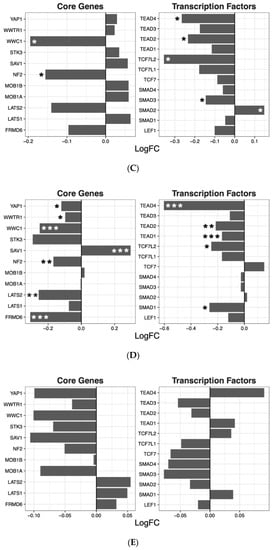
Table 2). These four drugs significantly regulated several individual genes in the Wnt pathway. While quetiapine also positively or negatively affected the transcription of a number of genes in the pathway, the overall effect of quetiapine on the Wnt signaling pathway was not significant (
Figure 2).
Figure 2. Wnt signaling pathway genes regulated by psychotropic drug treatment in NT2-N cells expressed as log fold change relative to vehicle-treated cells. * p < 0.05, ** p < 0.005, and *** p < 0.001.
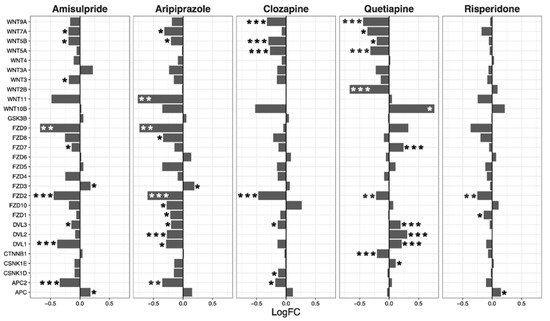
Overall, amisulpride (p = 0.0032), aripiprazole (p = 0.043), and clozapine (p = 0.0033) reduced the expression of genes in the notch signaling pathway. Again, quetiapine increased or decreased the expression of several genes in the pathway, but did not have a significant overall effect (
Figure 3).
Figure 3. Notch signaling pathway genes regulated by psychotropic drug treatment in NT2-N cells expressed as log fold change relative to vehicle-treated cells. * p < 0.05, ** p < 0.005, and *** p < 0.001.
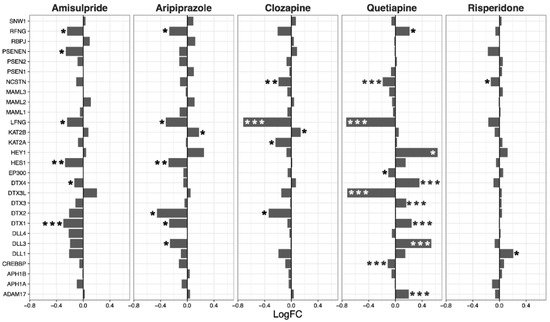
Expression of genes in the hedgehog signaling pathway was decreased overall by clozapine (p = 0.025), and tended to be reduced by aripiprazole (p = 0.07,
Figure 4). Clozapine significantly reduced the expression of CCND1, CCND2, and GLI2 (
Figure 4).
Figure 4. Hedgehog signaling pathway genes regulated by psychotropic drug treatment in NT2-N cells expressed as log fold change relative to vehicle-treated cells. * p < 0.05, ** p < 0.005, and *** p < 0.001.
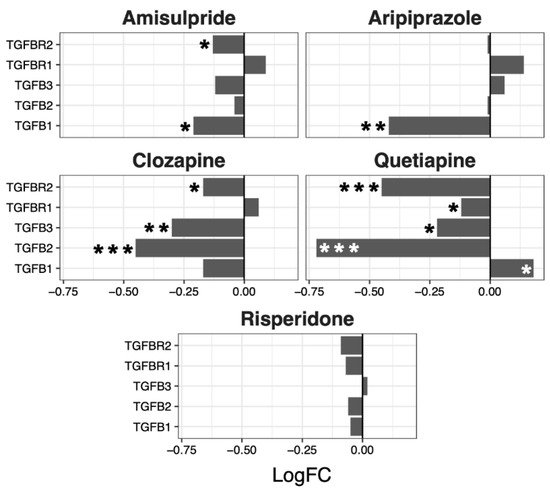
Although no overall significant effects were observed in the regulation of the TGF-β pathway, several genes were up- or downregulated by the different drugs, with a tendency to be downregulated by clozapine (p = 0.07) and risperidone (p = 0.06) (
Figure 5).
Figure 5. TGFβ signaling pathway genes regulated by psychotropic drug treatment in NT2-N cells expressed as log fold change relative to vehicle-treated cells. * p < 0.05, ** p < 0.005, and *** p < 0.001.
5. CMap
The genes in these signaling pathways that were regulated by the five drugs in the same direction (n = 22 genes with evidence of mean log fold change compared with the vehicle of p < 0.1;
Table S1) were submitted for CMap analysis, which identified 36 drugs with a positive CMap enrichment score of >0 and p < 0.05 (
Table 3).
Table 3. Compounds that affect the expression of the gene expression signature genes similar to the five drugs that affected the Hippo pathway.
| Rank |
CMap Name |
Enrichment |
p-Value |
Class |
| 1 |
Ursolic acid |
0.91 |
0.00008 |
Triterpenoid |
| 2 |
Levothyroxine sodium |
0.89 |
0.00014 |
Thyroid hormones |
| 3 |
Ajmaline |
0.94 |
0.00022 |
Antiarrhythmic agent |
| 4 |
5707885 |
0.87 |
0.00044 |
Unknown |
| 6 |
Carbimazole |
0.90 |
0.00182 |
Imidazole–thyroid function |
| 7 |
0297417–0002b |
0.88 |
0.00324 |
Unknown |
| 9 |
Ns-398 |
0.87 |
0.00463 |
COX-2 inhibitor (anti-inflammatory) |
| 10 |
Cefapirin |
0.78 |
0.00473 |
Antibiotic |
| 11 |
Estrone |
0.77 |
0.00539 |
Estrogen steroid |
| 12 |
Strophanthidin |
0.77 |
0.00561 |
Cardiac glycoside |
| 17 |
Picotamide |
0.68 |
0.00937 |
Platelet aggregation inhibitor |
| 20 |
Maprotiline |
0.72 |
0.0117 |
Tetracyclic antidepressant |
| 22 |
Monastrol |
0.53 |
0.01256 |
Antimitotic agent |
| 24 |
Sr-95639a (Aminopyridazine) |
0.72 |
0.01331 |
Muscarinic agonist |
| 28 |
Benzethonium chloride |
0.78 |
0.02097 |
Antiseptics and Disinfectants |
| 30 |
Ciprofibrate |
0.68 |
0.0228 |
Fibrate (lipid-lowering agent) |
| 31 |
5255229 |
0.90 |
0.02286 |
Unknown |
| 32 |
Methazolamide |
0.68 |
0.02316 |
Carbonic anhydrase inhibitors |
| 33 |
Bumetanide |
0.68 |
0.02379 |
Diuretic |
| 37 |
Sodium phenylbutyrate |
0.52 |
0.02663 |
Urea cycle disorder treatment agents |
| 38 |
Tenoxicam |
0.67 |
0.02795 |
NSAID (Nonsteroidal anti-inflammatory drug) |
| 40 |
Ah-23848 |
0.76 |
0.0291 |
Prostanoid EP4 antagonist |
| 41 |
Furazolidone |
0.66 |
0.03058 |
Oxazolidine–antibiotic agent |
| 42 |
Iloprost |
0.75 |
0.03201 |
Vasodilator |
| 43 |
Minoxidil |
0.60 |
0.03344 |
Vasodilator |
| 46 |
Hyoscyamine |
0.59 |
0.03475 |
Anticholinergic/antispasmodic |
| 45 |
Prestwick-642 (Epicatechin) |
0.65 |
0.03475 |
Catechin–antioxidant flavonoid |
| 48 |
Dihydrostreptomycin |
0.59 |
0.03529 |
Antibiotic |
| 49 |
Nisoxetine |
0.65 |
0.03539 |
Antidepressant -SNRI |
| 50 |
Ergocalciferol |
0.65 |
0.03589 |
Vitamin D analogue |
| 52 |
Ceforanide |
0.65 |
0.03804 |
Antibiotic |
| 55 |
Pheniramine |
0.58 |
0.04175 |
Antihistamine |
| 56 |
Ifenprodil |
0.64 |
0.04245 |
NMDA receptor antagonist |
| 57 |
Pha-00745360 |
0.46 |
0.04312 |
Unknown |
| 58 |
Clozapine |
0.32 |
0.04422 |
Atypical antipsychotic |
| 60 |
Diphemanil methylsulfate |
0.57 |
0.04552 |
Muscarinic antagonist |
| 63 |
Piperacetazine |
0.63 |
0.04729 |
Antipsychotic prodrug |
| 66 |
Trifluoperazine |
0.63 |
0.04882 |
Phenothiazine antipsychotic |
| 68 |
Lidoflazine |
0.71 |
0.04974 |
Vasodilator |
6. Summary
Downregulation of
NF2 and
WWC1 was observed following administration of aripiprazole, clozapine, and quetiapine. Likely consequences include reduced inflammatory signaling mediated by MST1/2 and NF2, and reduced MAPK and NFκB activity levels following upregulation of YAP/TAZ. Aripiprazole also downregulates T-cell factors, potentially compromising Wnt signaling
[22][51].
Several authors have demonstrated that MST1/2 directly regulates many aspects of immune activation and the activity of inflammatory pathways
[23][24][52,53]. Examples of pathways upregulated include MAPK (mitogen-activated protein kinase) and p53
[25][36]. MST1/2 also plays a major role in Toll-like receptor (TLR) signaling and subsequent activation of NFκB and pro-inflammatory cytokine production
[26][27][54,55]. Similarly,
YAP deletion in astrocytes induced JAK-STAT pathways, inducing reactive astrogliosis, microglial activation, and BBB dysfunction in mice
[28][56].
When considered as a whole, the net effects of these changes would reduce pro-inflammatory signaling mediated by MAPK and NFκB. This is of interest as aripiprazole
[29][57], quetiapine
[30][58], and clozapine
[31][32][59,60] are known to inhibit NFκB. The data produced in this study suggests that such inhibition may be achieved, at least in part, by effects on Hippo, Wnt, and TGFβ signaling (
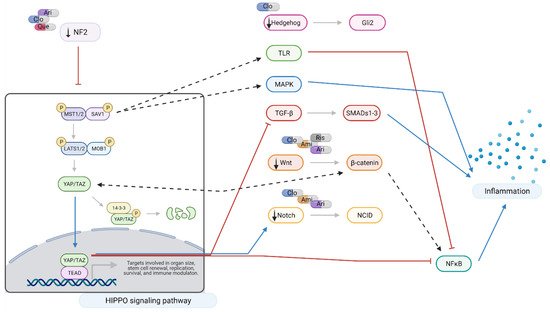
Figure 6).
Figure 6. Hippo signaling at a glance: effects of amisulpride, aripiprazole, clozapine, quetiapine, and risperidone. Legend: Ami = amisulpride; Ari = aripiprazole; Clo = clozapine; Que = quetiapine; Ris = risperidone; red arrows = inhibition; blue arrows = activation; dotted arrows = modulation; and black arrows = drugs action. In neuronal-like NT2 cells, inhibition of Hippo pathway reduces the production of pro-inflammatory cytokines through the crosstalk with TRL, MAPK, TGF-β, Wnt, and Notch signaling pathways. Aripiprazole, clozapine, and quetiapine downregulate NF2, leading to activation and nuclear translocation of YAP/TAZ which results in a reduction in NFκB and TGF-β signaling. NF2 also closely interacts with Hedgehog, TGF-β, Wnt, and Notch pathways. Modulation of MST1/2 expression also results in reduced pro-inflammatory signaling through Toll-like receptor (TRL) and MAPK signaling. Amisulpride, aripiprazole, clozapine, and risperidone downregulate Wnt signaling interfering with NFκB and pro-inflammatory cytokine production. Created with BioRender.com.