Yeasts affiliated with the Metschnikowia pulcherrima clade (subclade) of the large ascomycetous genus Metschnikowia frequently turn out to produce the characteristic maroon-red pulcherrimin when tested for pigment production and prove to exert antagonistic effects on many types of microorganisms. The determination of the exact taxonomic position of the strains is hampered by the shortage of distinctive morphological and physiological properties of the species of the clade and the lack of rDNA barcode gaps. The rDNA repeats of the type strains of the species are not homogenized and are assumed to evolve by a birth-and-death mechanism combined with reticulation. The taxonomic division is further hampered by the incomplete biological (reproductive) isolation of the species: certain type strains can be hybridized and genome sequencing revealed chimeric genome structures in certain strains that might have evolved from interspecies hybrids (alloploid genome duplication). Various mechanisms have been proposed for the antimicrobial antagonism. One is related to pulcherrimin production. The diffusible precursor of pulcherrimin, the pulcherriminic acid is secreted by the cells into the environment where it forms the insoluble pulcherrimin with the ferric ions. The lack of free iron caused by the immobilization of ferric ions inhibits the growth of many microorganisms.
Alert! Revision description should not be blank.
Where is the place for revision description?
- yeast
- antimicrobial antagonism
- bioprotection
- wine
- pulcherrimin
- rDNA reticulation
- iron immobilisation
1. Definition
Metschnikowia Kamienski (1899) is a large ascomycetous genus currently comprising 79 species (Mycobank, 04. 2020) but the number or species is continuously growing. The M. pulcherrima clade of the genus contains seven validly described species that share the ability to produce pulcherrimin, a maroon-red pigment (reviewed in [1][2]). These species and the strains closely related to them have broad biotechnological potential for application in various industrial processes. In wine fermentation, these yeasts can modulate the population dynamics of the fermenting yeast communities and produce enzymes and a broad range of compounds that improve the aromatic complexity of the wine (for a review, see [3]).
2. Introduction
Since the Kamechanisms underlying their antagonistic effect are not associated with the production of toxic compounds, these strains can safely be used as bioprotective agents to curb the invasion of pathogenic and rotting (saprophytic) microorganisms (Figure 1D) and/or additives in food technologies to modulate the dynamics of microbial populations. Numerous technological innovations involving antagonistic Metschnikowia straienski (1899) is a large ascomycetous genus currently comprising 79 specines have been patented (e.g., JPH01117778A, 1989; US6991930B1, 2006; NZ528225A, 2008; P0800775, 2008; ITTO20070655A1, 2009; WO2010149370, 2010; WO2010149369, 2010; CN101946805A 2011; CN103642705A, 2014; EP3266305A1, 2018; CN107904180A, 2018; CN110684678A, 2020;) and several Metschnikowia-(Mycobank, 04. 2020) based products have been commercialiZed [Excellence Bio-Nature (Lamothe-Abiet), Flavia and Gaïa (Lallemand), Shemer (Bayer, Koppert Biological Systems) Zymaflore Egide (Laffort)] as ADYs (Active Dry Yeast) for inoculated fermentation or as biocontrol agents for application against plant pathogens and post-harvest plant diseases.
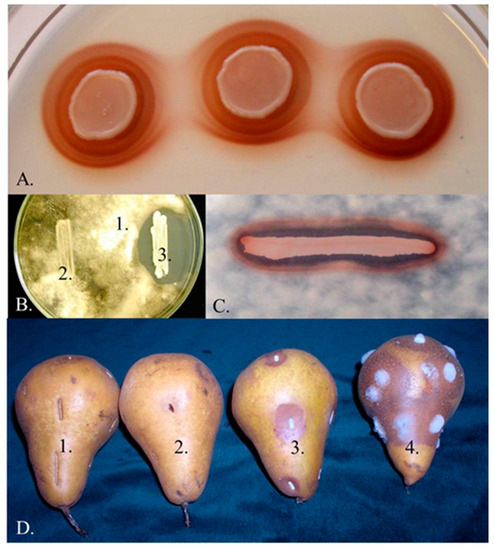
Figure 1. Pit the number or species is continuously gment production and antagonism. (A) Pigmented wing. Thalos around the colonies of the Mets. pulchnikowia strain 02.11.1.21. (B) Inhibition zone around the colony of the same strain (1: Botrytis cinerea 3318, 2: Saccharoimyces cerevisiae S288c, 3: Metschnikowia 02.11.1.21). (C) Coinciladence of pigmented halo and inhibition zone around the colony of Metschnikowia 02.11.1.21 on a medithe genum supplemented with 0.005 mg/mL FeCl3 and flooded with conidita of B. cinerea 3318. (D) Inhibition of rotting caused by Botrytis (1: untreats seved, 2: dipped in a suspension Metschnikowia 02.11.1.21 cevalls, 3: dipped in a mixed suspension of B. cinerea 3318 dly desconridia and Metschnikowia 02.11.1.21 cbells, 4: dipped in a suspension of B. cinerea 3318 speconidia). See reference [4] for es theat description of the strains.
Ovsharer the past two decades, large numbers of strains isolated from various substrates have been assigned to one or the other of these species (preferentially to M. pulcherrima) on the basis of barcode sequence ability to produce pulcherrimin, a maroon-red pigment (preferentially the D1/D2 domains of the LSU rRNA genes and the ITS1-5.8S-ITS2 segments of the rDNA repeatsviewed in [1,2]) identities/similarities. The general practice of sequence-based strain identification is a search with the sequence of the strain in nucleotide databases for identical/similar sequences and assigning the strain to the species whose database sequence is found most similar. The sequence of the strain is then routinely deposited in the database (most journals request accession numbers) under this taxonomic name usually without an expert taxonomic verification. Since small sequence differences are usually tolerated during identification, the new entries will gradually fill up the barcode gaps separating the closely related species; the species boundaries gradually become fuzzy. Thus, the taxonomic identification of new isolates by comparing their rDNA sequences with those deposited in databases can easily lead to false results. In addition to this general problem, other difficulties can also arise when pulcherrimin-producing Metschnikowia strse species and the strains closely related to them have broad biotechnological potential for application in various industrial processes. In wine fermentation, these yeasts can modulate the population dynains are to be identified taxonomically. The resultss of the in-depth analyses of the rDNA repeats of certain type strainsfermenting [5][6], the hybridization of type strains with each-other [6] and the analysis of genoast comme sequences (e.g., [7][8][9]) ranitised doubts as to whether the taxonomic division of the M. pulcherrima clas and prode is ucorrect at all. The incongruences around the mechanism of the antimicrobial antagonism pose another problem. While many researchers associate it with iron depletion, others prefer the view that non-iron-related mechanisms similar to those known in other antagonistic microorganisms are also involved or even play the major role.
3. The M. pulcherrima Clade
Pulcherrie enzymes and a broad range of compounds that improve the aromin-producing Metschnikowia satrains are common componentsc complexity of the yeast communities that colonise ripening fruits, flowers (nectar), tree sap fluxes and also frequently occur in fruit juices and fermenting wine (ewine (for a review, see [3]).g
2., [2][3][4][10][11][12][13][14][15][16][17][18][19][20][21][22][23][24][25][26]). New yeast isolates produciIng the characteristic maroon-red pulcherrimin halos around their colroductionies (Figure 1A) are frequently declared to belong to

3. The M. pulcherrima Clade
Pulcherrimin-producing Metschnikowia strains are common components of the yeast communities that colonise ripening fruits, flowers (nectar), tree sap fluxes and also frequently occur in fruit juices and fermenting wine (e.g., [2,3,6,9,12,18,19,20,21,22,23,24,25,26,27,28,29,30,31,32]). New yeast isolates producing the characteristic maroon-red pulcherrimin halos around their colonies (Figure 1A) are frequently declared to belong to M. pulcherrima
without taking into account that M. pulcherrima
is not the only pigment-producing Metschnikowia
species. Over the past two decades, five additional species (M. andauensis, M. rubicola, M. shanxiensis, M. sinensis, M. zizyphicola
) were validly described and M. fructicola, originally described as a pigment-less species has also turned out to produce pulcherrimin (for a review, see [1]). The intensity of pigment production is variable and highly dependent on the culturing conditions [1][33][34] and probably also on ploidy. The phylogenetic analysis of the barcode sequences of the type strains of the genus clustered these species in a group designated
, originally described as a pigment-less species has also turned out to produce pulcherrimin (for a review, see [1]). The intensity of pigment production is variable and highly dependent on the culturing conditions [6,33,34] and probably also on ploidy [35]. The phylogenetic analysis of the barcode sequences of the type strains of the genus clustered these species in a group designated
Recently, two additional species,
clade [1,2].
Recently, two additional species, M
. persimmonesis
and M. citriensis
were proposed to accommodate pulcherrimin-producing strains. The taxonomic name M. persimmonesis
was proposed for a single Korean isolate but without providing a complete taxonomic description [27]. The phylogenetic position of the strain is uncertain because its different rDNA barcode sequences (D1/D2, ITS and 18S) show the highest similarities to sequences of the type strains of different species. M. citriensis
is based on two strains isolated from citrus leaves [30]. The taxonomic position of these strains is also somewhat obscure, because the authors found them closely related to M. koreensis
based on the neighbour-joining analysis of the D1/D2 domains of the 26S rRNA genes, but the M. koreenesis
sequence used in the analysis was a direct GenBank submission amplified from a strain for which no taxonomic description is available. The other closest relatives were strains of three pigment-producing members of the M. pulcherrima
clade and the non-pigmented M. chrysoperlae
, but only one sequence used in the phylogenetic analysis represented a type strain. As previous analyses found M. koreensis
separated by a large phylogenetic distance from the clade [1,33], the proposed simultaneous close relationship to M. koreensis
and the M. pulcherrima
clade needs to be revised or reinforced by the analysis or more sequences. Interestingly, when the ITS sequences are examined, the M. citriensis
type strain differ more from the other M. citriensis
strain than from the M. persimmonesis
type strain. Besides, both the D1/D2 and the ITS sequences were cloned, and a phylogenetic analysis based on cloned sequences can easily be misleading in this group of yeast species because of the very high intragenomic diversity of the rDNA repeats (see below). The formation of spheroidal ascospores is another problematic feature of these isolates. M. pulcherrima
and its relatives have needle-shape spores [33]. Because of these uncertainties, further examination is required to validate the status of M. persimmonesis
and M. citriensis
as distinct species. Nevertheless, most properties of their strains and the results of the sequence analyses indicate taxonomic affinity with the M. pulcherrima
clade. Many pulcherrimin-producing isolates were not identified at the species level or could not be assigned to any species and were therefore only classified as Metschnikowia
sp., M.
aff. pulcherrima
or M.
aff. fructicola. On the other hand, many strains have been classified into these species without presenting sufficient taxonomic evidence. Since pigmentation is an irrelevant property in most biotechnological processes, the strains isolated for industrial purposes are normally not tested for pulcherrimin production. Therefore and because of the sensitivity of pulcherrimin synthesis to the culturing conditions, it is unknown whether pigmentation is a general ability of all strains of the clade.
. On the other hand, many strains have been classified into these species without presenting sufficient taxonomic evidence. Since pigmentation is an irrelevant property in most biotechnological processes, the strains isolated for industrial purposes are normally not tested for pulcherrimin production. Therefore and because of the sensitivity of pulcherrimin synthesis to the culturing conditions, it is unknown whether pigmentation is a general ability of all strains of the clade.
References
- Lachance, M.A. Metschnikowia: Half tetrads, a regicide and the fountain of youth. Yeast 2016, 33, 563–574. [Google Scholar] [CrossRef]
- Kurtzman, C.P.; Robnett, C.J.; Basehoar, E.; Ward, T.J. Four new species of Metschnikowia and the transfer of seven Candida species to Metschnikowia and Clavispora as new combinations. Antonie Van Leeuwenhoek 2018, 111, 2017–2035. [Google Scholar] [CrossRef]
- Morata, A.; Loira, I.; Escott, C.; del Fresno, J.M.; Bañuelos, M.A.; Suárez-Lepe, J.A. Applications of Metschnikowia pulcherrima in wine biotechnology. Fermentation 2019, 5, 63. [Google Scholar] [CrossRef]
- Abeln, F.; Chuck, C.J. Achieving a high-density oleaginous yeast culture: Comparison of four processing strategies using Metschnikowia pulcherrima. Biotechnol. Bioeng. 2019, 116, 3200–3214. [Google Scholar] [CrossRef]
- Kurtzman, C.P.; Droby, S. Metschnikowia fructicola, a new ascosporic yeast with potential for biocontrol of postharvest fruit rots. Syst. Appl. Microbiol. 2001, 24, 395–399. [Google Scholar] [CrossRef]
- Sipiczki, M. Metschnikowia strains isolated from botrytized grapes antagonize fungal and bacterial growth by iron depletion. Appl. Environ. Microbiol. 2006, 72, 6716–6724. [Google Scholar] [CrossRef]
- Saravanakumar, D.; Ciavorella, A.; Spadaro, D.; Garibaldi, A.; Gullino, M.L. Metschnikowia pulcherrima strain MACH1 outcompetes Botrytis cinerea, Alternaria alternata and Penicillium expansum in apples through iron depletion. Postharvest Biol. Technol. 2008, 49, 121–128. [Google Scholar] [CrossRef]
- Türkel, S.; Korukluoglu, M.; Yavuz, M. Biocontrol activity of the local strain of Metschnikowia pulcherrima on different postharvest pathogens. Biotechnol. Res. Int. 2014, 2014, 397167. [Google Scholar] [CrossRef] [PubMed]
- Kántor, A.; Hutková, J.; Petrová, J.; Hleba, L.; Kačániová, M. Antimicrobial activity of pulcherrimin pigment produced by Metschnikowia pulcherrima against various yeast species. J. Microbiol. Biotechnol. Food Sci. 2015, 5, 282–285. [Google Scholar] [CrossRef]
- Parafati, L.; Vitale, A.; Restuccia, C.; Cirvilleri, G. Biocontrol ability and action mechanism of food-isolated yeast strains against Botrytis cinerea causing post-harvest bunch rot of table grape. Food Microbiol. 2015, 47, 85–92. [Google Scholar] [CrossRef]
- Liu, Y.; Yi, L.; Ruan, C.; Yao, S.; Deng, L.; Zeng, K. Proline increases pigment production to improve oxidative stress tolerance and biocontrol ability of Metschnikowia citriensis. Front. Microbiol. 2019, 10, 1273. [Google Scholar] [CrossRef] [PubMed]
- Pawlikowska, E.; James, S.A.; Breierova, E.; Antolak, H.; Kregiel, D. Biocontrol capability of local Metschnikowia sp. isolates. Antonie Van Leeuwenhoek 2019, 112, 1425–1445. [Google Scholar] [CrossRef] [PubMed]
- Sipiczki, M.; Pfliegler, W.P.; Holb, I.J. Metschnikowia species share a pool of diverse rRNA genes differing in regions that determine hairpin-loop structures and evolve by reticulation. PLoS ONE 2013, 8, e67384. [Google Scholar] [CrossRef]
- Sipiczki, M.; Horvath, E.; Pfliegler, W.P. Birth-and-death evolution and reticulation of ITS segments of Metschnikowia andauensis and Metschnikowia fructicola rDNA repeats. Front. Microbiol. 2018, 9, 1193. [Google Scholar] [CrossRef]
- Piombo, E.; Sela, N.; Wisniewski, M.; Hoffmann, M.; Gullino, M.L.; Allard, M.W.; Levin, E.; Spadaro, D.; Droby, S. Genome sequence, assembly and characterization of two Metschnikowia fructicola strains used as biocontrol agents of postharvest diseases. Front. Microbiol. 2018, 9, 593. [Google Scholar] [CrossRef]
- Venkatesh, A.; Murray, A.L.; Boyle, A.B.; Quinn Farrington, L.; Maher, T.J.; Ó’Gaora, P.; Wolfe, K.H.; O’Brien, C.E.; Butler, G. Draft genome sequence of a highly heterozygous yeast strain from the Metschnikowia pulcherrima subclade, UCD127. Genome Announc. 2018, 6, e00550-18. [Google Scholar] [CrossRef]
- Gore-Lloyd, D.; Sumann, I.; Brachmann, A.O.; Schneeberger, K.; Ortiz-Merino, R.A.; Moreno-Beltrán, M.; Schläfli, M.; Kirner, P.; Santos Kron, A.; Rueda-Mejia, M.P.; et al. Snf2 controls pulcherriminic acid biosynthesis and antifungal activity of the biocontrol yeast Metschnikowia pulcherrima. Mol. Microbiol. 2019, 112, 317–332. [Google Scholar] [CrossRef] [PubMed]
- Molnar, O.; Prillinger, H. Analysis of yeast isolates related to Metschnikowia pulcherrima using the partial sequences of the large subunit rDNA and the actin gene; description of Metschnikowia andauensis sp. nov. Syst. Appl. Microbiol. 2005, 28, 717–726. [Google Scholar] [CrossRef]
- Xue, M.L.; Zhang, L.Q.; Wang, Q.M.; Zhang, J.S.; Bai, F.Y. Metschnikowia sinensis sp. nov., Metschnikowia zizyphicola sp. nov. and Metschnikowia shanxiensis sp. nov., novel yeast species from jujube fruit. Int. J. Syst. Evol. Microbiol. 2006, 56, 2245–2250. [Google Scholar] [CrossRef]
- Sipiczki, M. Overwintering of vineyard yeasts: Survival of interacting yeast communities in grapes mummified on vines. Front. Microbiol. 2016, 7, 212. [Google Scholar] [CrossRef]
- Türkel, S.; Ener, B. Isolation and characterization of new Metschnikowia pulcherrima strains as producers of the antimicrobial pigment pulcherrimin. Z. Naturforsch. C J. Biosci. 2009, 64, 405–410. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Spadaro, D.; Garibaldi, A.; Gullino, M.L. Selection and evaluation of new antagonists for their efficacy against postharvest brown rot of peaches. Postharvest Biol. Technol. 2009, 55, 174–181. [Google Scholar] [CrossRef]
- Vadkertiova, R.; Molnarova, J.; Vranova, D.; Slavikova, E. Yeasts and yeast-like organisms associated with fruits and blossoms of different fruit trees. Can. J. Microbiol. 2012, 58, 1344–1352. [Google Scholar] [CrossRef] [PubMed]
- Csutak, O.; Vassu, T.; Cornea, P.; Grebenisan, I. Genetic characterization of two new Metschnikowia strains with antifungal activity. Rom. Biotech. Lett. 2007, 12, 3175–3182. [Google Scholar]
- Csutak, O.; Vassu, T.; Sarbu, I.; Stoica, I.; Cornea, P. Antagonistic activity of three newly isolated yeast strains from the surface of fruits. Food Technol. Biotechnol. 2013, 51, 70–77. [Google Scholar]
- Hadwiger, L.A.; McDonel, H.; Glawe, D. Wild yeast strains as prospective candidates to induce resistance against potato late blight (Phytophthora infestans). Am. J. Potato Res. 2015, 92, 379–386. [Google Scholar] [CrossRef]
- Kang, Y.M.; Choi, J.E.; Komakech, R.; Park, J.H.; Kim, D.W.; Cho, K.M.; Kang, S.M.; Choi, S.H.; Song, K.C.; Ryu, C.M.; et al. Characterization of a novel yeast species Metschnikowia persimmonesis KCTC 12991BP (KIOM G15050 type strain) isolated from a medicinal plant, Korean persimmon calyx (Diospyros kaki Thumb). AMB Express 2017, 7, 199. [Google Scholar] [CrossRef]
- Pawlikowska, E.; Kręgiel, D. Enzymatic profiles and antimicrobial activity of the yeast Metschnikowia pulcherrima. Acta Innov. 2017, 23, 17–24. [Google Scholar]
- Liu, Y.; Wang, W.; Zhou, Y.; Yaoa, S.; Denga, L.; Zeng, K. Isolation, identification and in vitro screening of Chongqing orangery yeasts for the biocontrol of Penicillium digitatum on citrus fruit. Biol. Control 2017, 110, 18–24. [Google Scholar] [CrossRef]
- Liu, Y.; Yao, S.; Deng, L.; Ming, J.; Zeng, K. Metschnikowia citriensis sp. nov., a novel yeast species isolated from leaves with potential for biocontrol of postharvest fruit rot. Biol. Control 2018, 125, 15–19. [Google Scholar] [CrossRef]
- Tian, Y.Q.; Li, W.; Jiang, Z.T.; Jing, M.M.; Shao, Y.Z. The preservation effect of Metschnikowia pulcherrima yeast on anthracnose of postharvest mango fruits and the possible mechanism. Food Sci. Biotechnol. 2017, 27, 95–105. [Google Scholar] [CrossRef] [PubMed]
- Barbosa, C.; Lage, P.; Esteves, M.; Chambel, L.; Mendes-Faia, A.; Mendes-Ferreira, A. Molecular and phenotypic characterization of Metschnikowia pulcherrima strains from Douro wine region. Fermentation 2018, 4, 8. [Google Scholar] [CrossRef]
- Lachance, M.A. Metschnikowia Kamienski (1899). In The Yeasts. A Taxonomic Study; Kurtzman, C.P., Fell, J.W., Boekhout, T., Eds.; Elsevier: Amsterdam, The Netherlands, 2011; pp. 575–620. [Google Scholar]
M. pulcherrima
M. pulcherrima
Metschnikowia
M. andauensis, M. rubicola, M. shanxiensis, M. sinensis, M. zizyphicola
M. fructicola, originally described as a pigment-less species has also turned out to produce pulcherrimin (for a review, see [1]). The intensity of pigment production is variable and highly dependent on the culturing conditions [1][33][34] and probably also on ploidy. The phylogenetic analysis of the barcode sequences of the type strains of the genus clustered these species in a group designated
Recently, two additional species,
Recently, two additional species, M
. persimmonesis
and M. citriensis
were proposed to accommodate pulcherrimin-producing strains. The taxonomic name M. persimmonesis
was proposed for a single Korean isolate but without providing a complete taxonomic description [27]. The phylogenetic position of the strain is uncertain because its different rDNA barcode sequences (D1/D2, ITS and 18S) show the highest similarities to sequences of the type strains of different species. M. citriensis
is based on two strains isolated from citrus leaves [30]. The taxonomic position of these strains is also somewhat obscure, because the authors found them closely related to M. koreensis
based on the neighbour-joining analysis of the D1/D2 domains of the 26S rRNA genes, but the M. koreenesis
sequence used in the analysis was a direct GenBank submission amplified from a strain for which no taxonomic description is available. The other closest relatives were strains of three pigment-producing members of the M. pulcherrima
clade and the non-pigmented M. chrysoperlae
, but only one sequence used in the phylogenetic analysis represented a type strain. As previous analyses found M. koreensis
separated by a large phylogenetic distance from the clade [1,33], the proposed simultaneous close relationship to M. koreensis
and the M. pulcherrima
clade needs to be revised or reinforced by the analysis or more sequences. Interestingly, when the ITS sequences are examined, the M. citriensis
type strain differ more from the other M. citriensis
strain than from the M. persimmonesis
type strain. Besides, both the D1/D2 and the ITS sequences were cloned, and a phylogenetic analysis based on cloned sequences can easily be misleading in this group of yeast species because of the very high intragenomic diversity of the rDNA repeats (see below). The formation of spheroidal ascospores is another problematic feature of these isolates. M. pulcherrima
and its relatives have needle-shape spores [33]. Because of these uncertainties, further examination is required to validate the status of M. persimmonesis
and M. citriensis
as distinct species. Nevertheless, most properties of their strains and the results of the sequence analyses indicate taxonomic affinity with the M. pulcherrima
clade. Many pulcherrimin-producing isolates were not identified at the species level or could not be assigned to any species and were therefore only classified as Metschnikowia
sp., M.
aff. pulcherrima
or M.
aff. fructicola. On the other hand, many strains have been classified into these species without presenting sufficient taxonomic evidence. Since pigmentation is an irrelevant property in most biotechnological processes, the strains isolated for industrial purposes are normally not tested for pulcherrimin production. Therefore and because of the sensitivity of pulcherrimin synthesis to the culturing conditions, it is unknown whether pigmentation is a general ability of all strains of the clade.
. On the other hand, many strains have been classified into these species without presenting sufficient taxonomic evidence. Since pigmentation is an irrelevant property in most biotechnological processes, the strains isolated for industrial purposes are normally not tested for pulcherrimin production. Therefore and because of the sensitivity of pulcherrimin synthesis to the culturing conditions, it is unknown whether pigmentation is a general ability of all strains of the clade.
References
- Lachance, M.A. Metschnikowia: Half tetrads, a regicide and the fountain of youth. Yeast 2016, 33, 563–574. [Google Scholar] [CrossRef]
- Kurtzman, C.P.; Robnett, C.J.; Basehoar, E.; Ward, T.J. Four new species of Metschnikowia and the transfer of seven Candida species to Metschnikowia and Clavispora as new combinations. Antonie Van Leeuwenhoek 2018, 111, 2017–2035. [Google Scholar] [CrossRef]
- Morata, A.; Loira, I.; Escott, C.; del Fresno, J.M.; Bañuelos, M.A.; Suárez-Lepe, J.A. Applications of Metschnikowia pulcherrima in wine biotechnology. Fermentation 2019, 5, 63. [Google Scholar] [CrossRef]
- Abeln, F.; Chuck, C.J. Achieving a high-density oleaginous yeast culture: Comparison of four processing strategies using Metschnikowia pulcherrima. Biotechnol. Bioeng. 2019, 116, 3200–3214. [Google Scholar] [CrossRef]
- Kurtzman, C.P.; Droby, S. Metschnikowia fructicola, a new ascosporic yeast with potential for biocontrol of postharvest fruit rots. Syst. Appl. Microbiol. 2001, 24, 395–399. [Google Scholar] [CrossRef]
- Sipiczki, M. Metschnikowia strains isolated from botrytized grapes antagonize fungal and bacterial growth by iron depletion. Appl. Environ. Microbiol. 2006, 72, 6716–6724. [Google Scholar] [CrossRef]
- Saravanakumar, D.; Ciavorella, A.; Spadaro, D.; Garibaldi, A.; Gullino, M.L. Metschnikowia pulcherrima strain MACH1 outcompetes Botrytis cinerea, Alternaria alternata and Penicillium expansum in apples through iron depletion. Postharvest Biol. Technol. 2008, 49, 121–128. [Google Scholar] [CrossRef]
- Türkel, S.; Korukluoglu, M.; Yavuz, M. Biocontrol activity of the local strain of Metschnikowia pulcherrima on different postharvest pathogens. Biotechnol. Res. Int. 2014, 2014, 397167. [Google Scholar] [CrossRef] [PubMed]
- Kántor, A.; Hutková, J.; Petrová, J.; Hleba, L.; Kačániová, M. Antimicrobial activity of pulcherrimin pigment produced by Metschnikowia pulcherrima against various yeast species. J. Microbiol. Biotechnol. Food Sci. 2015, 5, 282–285. [Google Scholar] [CrossRef]
- Parafati, L.; Vitale, A.; Restuccia, C.; Cirvilleri, G. Biocontrol ability and action mechanism of food-isolated yeast strains against Botrytis cinerea causing post-harvest bunch rot of table grape. Food Microbiol. 2015, 47, 85–92. [Google Scholar] [CrossRef]
- Liu, Y.; Yi, L.; Ruan, C.; Yao, S.; Deng, L.; Zeng, K. Proline increases pigment production to improve oxidative stress tolerance and biocontrol ability of Metschnikowia citriensis. Front. Microbiol. 2019, 10, 1273. [Google Scholar] [CrossRef] [PubMed]
- Pawlikowska, E.; James, S.A.; Breierova, E.; Antolak, H.; Kregiel, D. Biocontrol capability of local Metschnikowia sp. isolates. Antonie Van Leeuwenhoek 2019, 112, 1425–1445. [Google Scholar] [CrossRef] [PubMed]
- Sipiczki, M.; Pfliegler, W.P.; Holb, I.J. Metschnikowia species share a pool of diverse rRNA genes differing in regions that determine hairpin-loop structures and evolve by reticulation. PLoS ONE 2013, 8, e67384. [Google Scholar] [CrossRef]
- Sipiczki, M.; Horvath, E.; Pfliegler, W.P. Birth-and-death evolution and reticulation of ITS segments of Metschnikowia andauensis and Metschnikowia fructicola rDNA repeats. Front. Microbiol. 2018, 9, 1193. [Google Scholar] [CrossRef]
- Piombo, E.; Sela, N.; Wisniewski, M.; Hoffmann, M.; Gullino, M.L.; Allard, M.W.; Levin, E.; Spadaro, D.; Droby, S. Genome sequence, assembly and characterization of two Metschnikowia fructicola strains used as biocontrol agents of postharvest diseases. Front. Microbiol. 2018, 9, 593. [Google Scholar] [CrossRef]
- Venkatesh, A.; Murray, A.L.; Boyle, A.B.; Quinn Farrington, L.; Maher, T.J.; Ó’Gaora, P.; Wolfe, K.H.; O’Brien, C.E.; Butler, G. Draft genome sequence of a highly heterozygous yeast strain from the Metschnikowia pulcherrima subclade, UCD127. Genome Announc. 2018, 6, e00550-18. [Google Scholar] [CrossRef]
- Gore-Lloyd, D.; Sumann, I.; Brachmann, A.O.; Schneeberger, K.; Ortiz-Merino, R.A.; Moreno-Beltrán, M.; Schläfli, M.; Kirner, P.; Santos Kron, A.; Rueda-Mejia, M.P.; et al. Snf2 controls pulcherriminic acid biosynthesis and antifungal activity of the biocontrol yeast Metschnikowia pulcherrima. Mol. Microbiol. 2019, 112, 317–332. [Google Scholar] [CrossRef] [PubMed]
- Molnar, O.; Prillinger, H. Analysis of yeast isolates related to Metschnikowia pulcherrima using the partial sequences of the large subunit rDNA and the actin gene; description of Metschnikowia andauensis sp. nov. Syst. Appl. Microbiol. 2005, 28, 717–726. [Google Scholar] [CrossRef]
- Xue, M.L.; Zhang, L.Q.; Wang, Q.M.; Zhang, J.S.; Bai, F.Y. Metschnikowia sinensis sp. nov., Metschnikowia zizyphicola sp. nov. and Metschnikowia shanxiensis sp. nov., novel yeast species from jujube fruit. Int. J. Syst. Evol. Microbiol. 2006, 56, 2245–2250. [Google Scholar] [CrossRef]
- Sipiczki, M. Overwintering of vineyard yeasts: Survival of interacting yeast communities in grapes mummified on vines. Front. Microbiol. 2016, 7, 212. [Google Scholar] [CrossRef]
- Türkel, S.; Ener, B. Isolation and characterization of new Metschnikowia pulcherrima strains as producers of the antimicrobial pigment pulcherrimin. Z. Naturforsch. C J. Biosci. 2009, 64, 405–410. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Spadaro, D.; Garibaldi, A.; Gullino, M.L. Selection and evaluation of new antagonists for their efficacy against postharvest brown rot of peaches. Postharvest Biol. Technol. 2009, 55, 174–181. [Google Scholar] [CrossRef]
- Vadkertiova, R.; Molnarova, J.; Vranova, D.; Slavikova, E. Yeasts and yeast-like organisms associated with fruits and blossoms of different fruit trees. Can. J. Microbiol. 2012, 58, 1344–1352. [Google Scholar] [CrossRef] [PubMed]
- Csutak, O.; Vassu, T.; Cornea, P.; Grebenisan, I. Genetic characterization of two new Metschnikowia strains with antifungal activity. Rom. Biotech. Lett. 2007, 12, 3175–3182. [Google Scholar]
- Csutak, O.; Vassu, T.; Sarbu, I.; Stoica, I.; Cornea, P. Antagonistic activity of three newly isolated yeast strains from the surface of fruits. Food Technol. Biotechnol. 2013, 51, 70–77. [Google Scholar]
- Hadwiger, L.A.; McDonel, H.; Glawe, D. Wild yeast strains as prospective candidates to induce resistance against potato late blight (Phytophthora infestans). Am. J. Potato Res. 2015, 92, 379–386. [Google Scholar] [CrossRef]
- Kang, Y.M.; Choi, J.E.; Komakech, R.; Park, J.H.; Kim, D.W.; Cho, K.M.; Kang, S.M.; Choi, S.H.; Song, K.C.; Ryu, C.M.; et al. Characterization of a novel yeast species Metschnikowia persimmonesis KCTC 12991BP (KIOM G15050 type strain) isolated from a medicinal plant, Korean persimmon calyx (Diospyros kaki Thumb). AMB Express 2017, 7, 199. [Google Scholar] [CrossRef]
- Pawlikowska, E.; Kręgiel, D. Enzymatic profiles and antimicrobial activity of the yeast Metschnikowia pulcherrima. Acta Innov. 2017, 23, 17–24. [Google Scholar]
- Liu, Y.; Wang, W.; Zhou, Y.; Yaoa, S.; Denga, L.; Zeng, K. Isolation, identification and in vitro screening of Chongqing orangery yeasts for the biocontrol of Penicillium digitatum on citrus fruit. Biol. Control 2017, 110, 18–24. [Google Scholar] [CrossRef]
- Liu, Y.; Yao, S.; Deng, L.; Ming, J.; Zeng, K. Metschnikowia citriensis sp. nov., a novel yeast species isolated from leaves with potential for biocontrol of postharvest fruit rot. Biol. Control 2018, 125, 15–19. [Google Scholar] [CrossRef]
- Tian, Y.Q.; Li, W.; Jiang, Z.T.; Jing, M.M.; Shao, Y.Z. The preservation effect of Metschnikowia pulcherrima yeast on anthracnose of postharvest mango fruits and the possible mechanism. Food Sci. Biotechnol. 2017, 27, 95–105. [Google Scholar] [CrossRef] [PubMed]
- Barbosa, C.; Lage, P.; Esteves, M.; Chambel, L.; Mendes-Faia, A.; Mendes-Ferreira, A. Molecular and phenotypic characterization of Metschnikowia pulcherrima strains from Douro wine region. Fermentation 2018, 4, 8. [Google Scholar] [CrossRef]
- Lachance, M.A. Metschnikowia Kamienski (1899). In The Yeasts. A Taxonomic Study; Kurtzman, C.P., Fell, J.W., Boekhout, T., Eds.; Elsevier: Amsterdam, The Netherlands, 2011; pp. 575–620. [Google Scholar]
M
persimmonesis
M. citriensis
M. persimmonesis
M. citriensis
M. koreensis
M. koreenesis
M. pulcherrima
M. chrysoperlae
M. koreensis
M. koreensis
M. pulcherrima
M. citriensis
M. citriensis
M. persimmonesis
M. pulcherrima
M. persimmonesis
M. citriensis
M. pulcherrima
Metschnikowia
M.
pulcherrima
M.
fructicola. On the other hand, many strains have been classified into these species without presenting sufficient taxonomic evidence. Since pigmentation is an irrelevant property in most biotechnological processes, the strains isolated for industrial purposes are normally not tested for pulcherrimin production. Therefore and because of the sensitivity of pulcherrimin synthesis to the culturing conditions, it is unknown whether pigmentation is a general ability of all strains of the clade.
References
- Lachance, M.A. Metschnikowia: Half tetrads, a regicide and the fountain of youth. Yeast 2016, 33, 563–574. [Google Scholar] [CrossRef]
- Kurtzman, C.P.; Robnett, C.J.; Basehoar, E.; Ward, T.J. Four new species of Metschnikowia and the transfer of seven Candida species to Metschnikowia and Clavispora as new combinations. Antonie Van Leeuwenhoek 2018, 111, 2017–2035. [Google Scholar] [CrossRef]
- Morata, A.; Loira, I.; Escott, C.; del Fresno, J.M.; Bañuelos, M.A.; Suárez-Lepe, J.A. Applications of Metschnikowia pulcherrima in wine biotechnology. Fermentation 2019, 5, 63. [Google Scholar] [CrossRef]
- Abeln, F.; Chuck, C.J. Achieving a high-density oleaginous yeast culture: Comparison of four processing strategies using Metschnikowia pulcherrima. Biotechnol. Bioeng. 2019, 116, 3200–3214. [Google Scholar] [CrossRef]
- Kurtzman, C.P.; Droby, S. Metschnikowia fructicola, a new ascosporic yeast with potential for biocontrol of postharvest fruit rots. Syst. Appl. Microbiol. 2001, 24, 395–399. [Google Scholar] [CrossRef]
- Sipiczki, M. Metschnikowia strains isolated from botrytized grapes antagonize fungal and bacterial growth by iron depletion. Appl. Environ. Microbiol. 2006, 72, 6716–6724. [Google Scholar] [CrossRef]
- Saravanakumar, D.; Ciavorella, A.; Spadaro, D.; Garibaldi, A.; Gullino, M.L. Metschnikowia pulcherrima strain MACH1 outcompetes Botrytis cinerea, Alternaria alternata and Penicillium expansum in apples through iron depletion. Postharvest Biol. Technol. 2008, 49, 121–128. [Google Scholar] [CrossRef]
- Türkel, S.; Korukluoglu, M.; Yavuz, M. Biocontrol activity of the local strain of Metschnikowia pulcherrima on different postharvest pathogens. Biotechnol. Res. Int. 2014, 2014, 397167. [Google Scholar] [CrossRef] [PubMed]
- Kántor, A.; Hutková, J.; Petrová, J.; Hleba, L.; Kačániová, M. Antimicrobial activity of pulcherrimin pigment produced by Metschnikowia pulcherrima against various yeast species. J. Microbiol. Biotechnol. Food Sci. 2015, 5, 282–285. [Google Scholar] [CrossRef]
- Parafati, L.; Vitale, A.; Restuccia, C.; Cirvilleri, G. Biocontrol ability and action mechanism of food-isolated yeast strains against Botrytis cinerea causing post-harvest bunch rot of table grape. Food Microbiol. 2015, 47, 85–92. [Google Scholar] [CrossRef]
- Liu, Y.; Yi, L.; Ruan, C.; Yao, S.; Deng, L.; Zeng, K. Proline increases pigment production to improve oxidative stress tolerance and biocontrol ability of Metschnikowia citriensis. Front. Microbiol. 2019, 10, 1273. [Google Scholar] [CrossRef] [PubMed]
- Pawlikowska, E.; James, S.A.; Breierova, E.; Antolak, H.; Kregiel, D. Biocontrol capability of local Metschnikowia sp. isolates. Antonie Van Leeuwenhoek 2019, 112, 1425–1445. [Google Scholar] [CrossRef] [PubMed]
- Sipiczki, M.; Pfliegler, W.P.; Holb, I.J. Metschnikowia species share a pool of diverse rRNA genes differing in regions that determine hairpin-loop structures and evolve by reticulation. PLoS ONE 2013, 8, e67384. [Google Scholar] [CrossRef]
- Sipiczki, M.; Horvath, E.; Pfliegler, W.P. Birth-and-death evolution and reticulation of ITS segments of Metschnikowia andauensis and Metschnikowia fructicola rDNA repeats. Front. Microbiol. 2018, 9, 1193. [Google Scholar] [CrossRef]
- Piombo, E.; Sela, N.; Wisniewski, M.; Hoffmann, M.; Gullino, M.L.; Allard, M.W.; Levin, E.; Spadaro, D.; Droby, S. Genome sequence, assembly and characterization of two Metschnikowia fructicola strains used as biocontrol agents of postharvest diseases. Front. Microbiol. 2018, 9, 593. [Google Scholar] [CrossRef]
- Venkatesh, A.; Murray, A.L.; Boyle, A.B.; Quinn Farrington, L.; Maher, T.J.; Ó’Gaora, P.; Wolfe, K.H.; O’Brien, C.E.; Butler, G. Draft genome sequence of a highly heterozygous yeast strain from the Metschnikowia pulcherrima subclade, UCD127. Genome Announc. 2018, 6, e00550-18. [Google Scholar] [CrossRef]
- Gore-Lloyd, D.; Sumann, I.; Brachmann, A.O.; Schneeberger, K.; Ortiz-Merino, R.A.; Moreno-Beltrán, M.; Schläfli, M.; Kirner, P.; Santos Kron, A.; Rueda-Mejia, M.P.; et al. Snf2 controls pulcherriminic acid biosynthesis and antifungal activity of the biocontrol yeast Metschnikowia pulcherrima. Mol. Microbiol. 2019, 112, 317–332. [Google Scholar] [CrossRef] [PubMed]
- Molnar, O.; Prillinger, H. Analysis of yeast isolates related to Metschnikowia pulcherrima using the partial sequences of the large subunit rDNA and the actin gene; description of Metschnikowia andauensis sp. nov. Syst. Appl. Microbiol. 2005, 28, 717–726. [Google Scholar] [CrossRef]
- Xue, M.L.; Zhang, L.Q.; Wang, Q.M.; Zhang, J.S.; Bai, F.Y. Metschnikowia sinensis sp. nov., Metschnikowia zizyphicola sp. nov. and Metschnikowia shanxiensis sp. nov., novel yeast species from jujube fruit. Int. J. Syst. Evol. Microbiol. 2006, 56, 2245–2250. [Google Scholar] [CrossRef]
- Sipiczki, M. Overwintering of vineyard yeasts: Survival of interacting yeast communities in grapes mummified on vines. Front. Microbiol. 2016, 7, 212. [Google Scholar] [CrossRef]
- Türkel, S.; Ener, B. Isolation and characterization of new Metschnikowia pulcherrima strains as producers of the antimicrobial pigment pulcherrimin. Z. Naturforsch. C J. Biosci. 2009, 64, 405–410. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Spadaro, D.; Garibaldi, A.; Gullino, M.L. Selection and evaluation of new antagonists for their efficacy against postharvest brown rot of peaches. Postharvest Biol. Technol. 2009, 55, 174–181. [Google Scholar] [CrossRef]
- Vadkertiova, R.; Molnarova, J.; Vranova, D.; Slavikova, E. Yeasts and yeast-like organisms associated with fruits and blossoms of different fruit trees. Can. J. Microbiol. 2012, 58, 1344–1352. [Google Scholar] [CrossRef] [PubMed]
- Csutak, O.; Vassu, T.; Cornea, P.; Grebenisan, I. Genetic characterization of two new Metschnikowia strains with antifungal activity. Rom. Biotech. Lett. 2007, 12, 3175–3182. [Google Scholar]
- Csutak, O.; Vassu, T.; Sarbu, I.; Stoica, I.; Cornea, P. Antagonistic activity of three newly isolated yeast strains from the surface of fruits. Food Technol. Biotechnol. 2013, 51, 70–77. [Google Scholar]
- Hadwiger, L.A.; McDonel, H.; Glawe, D. Wild yeast strains as prospective candidates to induce resistance against potato late blight (Phytophthora infestans). Am. J. Potato Res. 2015, 92, 379–386. [Google Scholar] [CrossRef]
- Kang, Y.M.; Choi, J.E.; Komakech, R.; Park, J.H.; Kim, D.W.; Cho, K.M.; Kang, S.M.; Choi, S.H.; Song, K.C.; Ryu, C.M.; et al. Characterization of a novel yeast species Metschnikowia persimmonesis KCTC 12991BP (KIOM G15050 type strain) isolated from a medicinal plant, Korean persimmon calyx (Diospyros kaki Thumb). AMB Express 2017, 7, 199. [Google Scholar] [CrossRef]
- Pawlikowska, E.; Kręgiel, D. Enzymatic profiles and antimicrobial activity of the yeast Metschnikowia pulcherrima. Acta Innov. 2017, 23, 17–24. [Google Scholar]
- Liu, Y.; Wang, W.; Zhou, Y.; Yaoa, S.; Denga, L.; Zeng, K. Isolation, identification and in vitro screening of Chongqing orangery yeasts for the biocontrol of Penicillium digitatum on citrus fruit. Biol. Control 2017, 110, 18–24. [Google Scholar] [CrossRef]
- Liu, Y.; Yao, S.; Deng, L.; Ming, J.; Zeng, K. Metschnikowia citriensis sp. nov., a novel yeast species isolated from leaves with potential for biocontrol of postharvest fruit rot. Biol. Control 2018, 125, 15–19. [Google Scholar] [CrossRef]
- Tian, Y.Q.; Li, W.; Jiang, Z.T.; Jing, M.M.; Shao, Y.Z. The preservation effect of Metschnikowia pulcherrima yeast on anthracnose of postharvest mango fruits and the possible mechanism. Food Sci. Biotechnol. 2017, 27, 95–105. [Google Scholar] [CrossRef] [PubMed]
- Barbosa, C.; Lage, P.; Esteves, M.; Chambel, L.; Mendes-Faia, A.; Mendes-Ferreira, A. Molecular and phenotypic characterization of Metschnikowia pulcherrima strains from Douro wine region. Fermentation 2018, 4, 8. [Google Scholar] [CrossRef]
- Lachance, M.A. Metschnikowia Kamienski (1899). In The Yeasts. A Taxonomic Study; Kurtzman, C.P., Fell, J.W., Boekhout, T., Eds.; Elsevier: Amsterdam, The Netherlands, 2011; pp. 575–620. [Google Scholar]
References
- Lachance, M.A. Metschnikowia: Half tetrads, a regicide and the fountain of youth. Yeast 2016, 33, 563–574. [Google Scholar] [CrossRef]
- Kurtzman, C.P.; Robnett, C.J.; Basehoar, E.; Ward, T.J. Four new species of Metschnikowia and the transfer of seven Candida species to Metschnikowia and Clavispora as new combinations. Antonie Van Leeuwenhoek 2018, 111, 2017–2035. [Google Scholar] [CrossRef]
- Morata, A.; Loira, I.; Escott, C.; del Fresno, J.M.; Bañuelos, M.A.; Suárez-Lepe, J.A. Applications of Metschnikowia pulcherrima in wine biotechnology. Fermentation 2019, 5, 63. [Google Scholar] [CrossRef]
- Abeln, F.; Chuck, C.J. Achieving a high-density oleaginous yeast culture: Comparison of four processing strategies using Metschnikowia pulcherrima. Biotechnol. Bioeng. 2019, 116, 3200–3214. [Google Scholar] [CrossRef]
- Kurtzman, C.P.; Droby, S. Metschnikowia fructicola, a new ascosporic yeast with potential for biocontrol of postharvest fruit rots. Syst. Appl. Microbiol. 2001, 24, 395–399. [Google Scholar] [CrossRef]
- Sipiczki, M. Metschnikowia strains isolated from botrytized grapes antagonize fungal and bacterial growth by iron depletion. Appl. Environ. Microbiol. 2006, 72, 6716–6724. [Google Scholar] [CrossRef]
- Saravanakumar, D.; Ciavorella, A.; Spadaro, D.; Garibaldi, A.; Gullino, M.L. Metschnikowia pulcherrima strain MACH1 outcompetes Botrytis cinerea, Alternaria alternata and Penicillium expansum in apples through iron depletion. Postharvest Biol. Technol. 2008, 49, 121–128. [Google Scholar] [CrossRef]
- Türkel, S.; Korukluoglu, M.; Yavuz, M. Biocontrol activity of the local strain of Metschnikowia pulcherrima on different postharvest pathogens. Biotechnol. Res. Int. 2014, 2014, 397167. [Google Scholar] [CrossRef] [PubMed]
- Kántor, A.; Hutková, J.; Petrová, J.; Hleba, L.; Kačániová, M. Antimicrobial activity of pulcherrimin pigment produced by Metschnikowia pulcherrima against various yeast species. J. Microbiol. Biotechnol. Food Sci. 2015, 5, 282–285. [Google Scholar] [CrossRef]
- Parafati, L.; Vitale, A.; Restuccia, C.; Cirvilleri, G. Biocontrol ability and action mechanism of food-isolated yeast strains against Botrytis cinerea causing post-harvest bunch rot of table grape. Food Microbiol. 2015, 47, 85–92. [Google Scholar] [CrossRef]
- Liu, Y.; Yi, L.; Ruan, C.; Yao, S.; Deng, L.; Zeng, K. Proline increases pigment production to improve oxidative stress tolerance and biocontrol ability of Metschnikowia citriensis. Front. Microbiol. 2019, 10, 1273. [Google Scholar] [CrossRef] [PubMed]
- Pawlikowska, E.; James, S.A.; Breierova, E.; Antolak, H.; Kregiel, D. Biocontrol capability of local Metschnikowia sp. isolates. Antonie Van Leeuwenhoek 2019, 112, 1425–1445. [Google Scholar] [CrossRef] [PubMed]
- Sipiczki, M.; Pfliegler, W.P.; Holb, I.J. Metschnikowia species share a pool of diverse rRNA genes differing in regions that determine hairpin-loop structures and evolve by reticulation. PLoS ONE 2013, 8, e67384. [Google Scholar] [CrossRef]
- Sipiczki, M.; Horvath, E.; Pfliegler, W.P. Birth-and-death evolution and reticulation of ITS segments of Metschnikowia andauensis and Metschnikowia fructicola rDNA repeats. Front. Microbiol. 2018, 9, 1193. [Google Scholar] [CrossRef]
- Piombo, E.; Sela, N.; Wisniewski, M.; Hoffmann, M.; Gullino, M.L.; Allard, M.W.; Levin, E.; Spadaro, D.; Droby, S. Genome sequence, assembly and characterization of two Metschnikowia fructicola strains used as biocontrol agents of postharvest diseases. Front. Microbiol. 2018, 9, 593. [Google Scholar] [CrossRef]
- Venkatesh, A.; Murray, A.L.; Boyle, A.B.; Quinn Farrington, L.; Maher, T.J.; Ó’Gaora, P.; Wolfe, K.H.; O’Brien, C.E.; Butler, G. Draft genome sequence of a highly heterozygous yeast strain from the Metschnikowia pulcherrima subclade, UCD127. Genome Announc. 2018, 6, e00550-18. [Google Scholar] [CrossRef]
- Gore-Lloyd, D.; Sumann, I.; Brachmann, A.O.; Schneeberger, K.; Ortiz-Merino, R.A.; Moreno-Beltrán, M.; Schläfli, M.; Kirner, P.; Santos Kron, A.; Rueda-Mejia, M.P.; et al. Snf2 controls pulcherriminic acid biosynthesis and antifungal activity of the biocontrol yeast Metschnikowia pulcherrima. Mol. Microbiol. 2019, 112, 317–332. [Google Scholar] [CrossRef] [PubMed]
- Molnar, O.; Prillinger, H. Analysis of yeast isolates related to Metschnikowia pulcherrima using the partial sequences of the large subunit rDNA and the actin gene; description of Metschnikowia andauensis sp. nov. Syst. Appl. Microbiol. 2005, 28, 717–726. [Google Scholar] [CrossRef]
- Xue, M.L.; Zhang, L.Q.; Wang, Q.M.; Zhang, J.S.; Bai, F.Y. Metschnikowia sinensis sp. nov., Metschnikowia zizyphicola sp. nov. and Metschnikowia shanxiensis sp. nov., novel yeast species from jujube fruit. Int. J. Syst. Evol. Microbiol. 2006, 56, 2245–2250. [Google Scholar] [CrossRef]
- Sipiczki, M. Overwintering of vineyard yeasts: Survival of interacting yeast communities in grapes mummified on vines. Front. Microbiol. 2016, 7, 212. [Google Scholar] [CrossRef]
- Türkel, S.; Ener, B. Isolation and characterization of new Metschnikowia pulcherrima strains as producers of the antimicrobial pigment pulcherrimin. Z. Naturforsch. C J. Biosci. 2009, 64, 405–410. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Spadaro, D.; Garibaldi, A.; Gullino, M.L. Selection and evaluation of new antagonists for their efficacy against postharvest brown rot of peaches. Postharvest Biol. Technol. 2009, 55, 174–181. [Google Scholar] [CrossRef]
- Vadkertiova, R.; Molnarova, J.; Vranova, D.; Slavikova, E. Yeasts and yeast-like organisms associated with fruits and blossoms of different fruit trees. Can. J. Microbiol. 2012, 58, 1344–1352. [Google Scholar] [CrossRef] [PubMed]
- Csutak, O.; Vassu, T.; Cornea, P.; Grebenisan, I. Genetic characterization of two new Metschnikowia strains with antifungal activity. Rom. Biotech. Lett. 2007, 12, 3175–3182. [Google Scholar]
- Csutak, O.; Vassu, T.; Sarbu, I.; Stoica, I.; Cornea, P. Antagonistic activity of three newly isolated yeast strains from the surface of fruits. Food Technol. Biotechnol. 2013, 51, 70–77. [Google Scholar]
- Hadwiger, L.A.; McDonel, H.; Glawe, D. Wild yeast strains as prospective candidates to induce resistance against potato late blight (Phytophthora infestans). Am. J. Potato Res. 2015, 92, 379–386. [Google Scholar] [CrossRef]
- Kang, Y.M.; Choi, J.E.; Komakech, R.; Park, J.H.; Kim, D.W.; Cho, K.M.; Kang, S.M.; Choi, S.H.; Song, K.C.; Ryu, C.M.; et al. Characterization of a novel yeast species Metschnikowia persimmonesis KCTC 12991BP (KIOM G15050 type strain) isolated from a medicinal plant, Korean persimmon calyx (Diospyros kaki Thumb). AMB Express 2017, 7, 199. [Google Scholar] [CrossRef]
- Pawlikowska, E.; Kręgiel, D. Enzymatic profiles and antimicrobial activity of the yeast Metschnikowia pulcherrima. Acta Innov. 2017, 23, 17–24. [Google Scholar]
- Liu, Y.; Wang, W.; Zhou, Y.; Yaoa, S.; Denga, L.; Zeng, K. Isolation, identification and in vitro screening of Chongqing orangery yeasts for the biocontrol of Penicillium digitatum on citrus fruit. Biol. Control 2017, 110, 18–24. [Google Scholar] [CrossRef]
- Liu, Y.; Yao, S.; Deng, L.; Ming, J.; Zeng, K. Metschnikowia citriensis sp. nov., a novel yeast species isolated from leaves with potential for biocontrol of postharvest fruit rot. Biol. Control 2018, 125, 15–19. [Google Scholar] [CrossRef]
- Tian, Y.Q.; Li, W.; Jiang, Z.T.; Jing, M.M.; Shao, Y.Z. The preservation effect of Metschnikowia pulcherrima yeast on anthracnose of postharvest mango fruits and the possible mechanism. Food Sci. Biotechnol. 2017, 27, 95–105. [Google Scholar] [CrossRef] [PubMed]
- Barbosa, C.; Lage, P.; Esteves, M.; Chambel, L.; Mendes-Faia, A.; Mendes-Ferreira, A. Molecular and phenotypic characterization of Metschnikowia pulcherrima strains from Douro wine region. Fermentation 2018, 4, 8. [Google Scholar] [CrossRef]
- Lachance, M.A. Metschnikowia Kamienski (1899). In The Yeasts. A Taxonomic Study; Kurtzman, C.P., Fell, J.W., Boekhout, T., Eds.; Elsevier: Amsterdam, The Netherlands, 2011; pp. 575–620. [Google Scholar]
