Castanea sativa is one of the main multipurpose tree species valued for its timber and nuts. This species is susceptible to two major diseases, ink disease and chestnut blight, caused by Phytophthora spp. and Cryphonectria parasitica, respectively. The loss-of-function mutations of genes required for the onset of pathogenesis, referred to as plant susceptibility (S) genes, are one mechanism of plant resistance against pathogens. On the basis of sequence homology, functional domain identification, and phylogenetic analyses, we report for the first time on the identification of S-genes (mlo1, dmr6, dnd1, and pmr4) in the Castanea genus. The expression dynamics of S-genes were assessed in C. sativa and C. crenata plants inoculated with P. cinnamomi and C. parasitica. Our results highlighted the upregulation of pmr4 and dmr6 in response to pathogen infection. Pmr4 was strongly expressed at early infection phases of both pathogens in C. sativa, whereas in C. crenata, no significant upregulation was observed. The infection of P. cinnamomi led to a higher increase in the transcript level of dmr6 in C. sativa compared to C. crenata-infected samples. For a better understanding of plant responses, the transcript levels of defense genes gluB and chi3 were also analyzed.
- chestnut
- susceptibility genes
- Phytophthora cinnamomi
- Cryphonectria parasitica
1. Introduction
The
genus belongs to the Fagaceae family and includes four major species of commercial and ecosystemic interest:
Mill. (European chestnut),
Sieb. et Zucc. (Japanese chestnut),
Bl. (Chinese chestnut), and
Borkh (American chestnut).
is a woody species common in all Mediterranean countries and Asia Minor. It has been widely used since ancient times, not only for the consumption of its edible nuts, but also for wood and the products of its ecosystem, such as mushrooms and honey. It is a forest tree, relevant for landscape ecology and biodiversity of mountain and rural areas [1].
2. History
Over the last century, the number of chestnut trees decreased in growing areas in Europe due to the depopulation of mountains, climate change, and the spread of two severe diseases: ink disease and chestnut blight [2][3]. Ink disease is caused by the Oomycete
and
. Both species are pathogenic to
, although
generally displays greater virulence than
species, only
exhibits high tolerance to
[6]. The disease, which affects both young and old trees, leads to subcortical necrosis of the root system and the basal part of the stem; this is followed by the appearance of wasting symptoms in the foliage until the total desiccation and death of the plant occur [7][8][9][10]. These pathogens spread mainly through the movement of soil harboring inoculum and the dissemination of asexual flagellated spores (i.e., zoospores) that can actively travel short distances or passively travel long distances in flowing water [10][11]. The use of resistant rootstocks represents one possible solution to protect against these pathogens, although, at present, only tolerant selections obtained from hybridization between
and
are available [12].
Chestnut blight stands among the most destructive fungal tree diseases ever [10][13]. The causal agent,
, infects trees through dead plant tissue and wounds, including those caused by pruning, graft, and hail [13][14]. The symptoms involve bark cankers that can develop on suckers, young branches, and adult branches and trunks [15]. Chestnut blight was one of the causes of the abandonment of chestnut orchards in Europe until the end of the 1970s, when the natural spread of the hypovirulent form of the fungus favored a slow but progressive recovery of chestnut orchards and coppices. However, the fungus still represents a relevant problem in many areas of Europe. It is very harmful to young grafted trees in particular, hampering the establishment of new orchards in many areas [10][13].
forests in Eastern North America were wiped out by
in the early 20th century [16]. Extensive studies and breeding activities have been carried out to restore the American chestnut species introgressing resistance genes of
[17][18]. More recently, researchers discovered that the onset of the disease is associated with the release of oxalic acid by the pathogen during infection. Blight-resistant
trees were obtained by transferring a wheat gene that encodes oxalate oxidase [19].
Recently, a new interest and sensitivity towards the preservation of the local landscape generated a growing interest in silviculture and chestnut trees [20]. Moreover, the market demand for chestnuts in European countries has been strong in the last two decades and has often been supplied by importations. This has been due in part to the gall wasp (
Yasumatsu) infestation, which only recently has been controlled effectively [1][21], and to the general difficulty of developing a modern chestnut industry based on quality cultivars of
that are more tolerant to pathogens. The elucidation of the genetic mechanism behind host–pathogen interaction could thus be useful for the development of novel breeding strategies aimed at achieving resistance or higher tolerance to these pathogens.
Plants take advantage of different defense mechanisms during pathogen attack, and pathogens trigger counter-defense mechanisms. Plants carry pattern recognition receptors (PRRs) able to perceive pathogen-associated molecular patterns (PAMPs); this perception leads to intracellular signal transduction culminating in PAMP-triggered immunity (PTI). PTI is characterized by the production of reactive oxygen species (ROS), the secretion of antimicrobial compounds, and hydrolytic enzymes targeting the pathogen cell wall (chitinase and glucanase) and local cell wall fortifications (through callose deposition) [22].
To suppress PTI, pathogens developed effector molecules able to facilitate pathogen infection by manipulating the host response to support compatibility. Plant resistance (R) genes can detect effectors and trigger effector-triggered immunity (ETI) [23]. The recognition between R genes and effectors causes a cascade of responses involving jasmonic acid (JA) and salicylic acid (SA), culminating in a hypersensitive response (HR) [24].
Most pathogens require the cooperation of the host to establish a compatible interaction. Plant genes supporting compatibility and facilitating infection are called susceptibility (S) genes. S-genes can be divided into three main classes: (a) genes required for the early pathogen infection step (basic compatibility); (b) genes encoding negative regulators of plant immunity; (c) genes necessary for pathogen proliferation (sustained compatibility) [22].
The mutation or loss of an S-gene can thus limit the ability of the pathogen to infect the host and the spread of the disease. The resistance mediated by the S-gene mutation can be pathogen-specific or broad-spectrum. In the former case, the pathway can be implicated in the penetration phase; in the latter, one of the target genes can be involved in constitutive defense responses [22]. Resistance due to the loss of S-genes is generally recessive, differing from the generally dominant resistance mediated by R genes.
Among the S-genes,
(
),
(
),
(
), and
(
) have been characterized in many plant species. The
gene family, encoding seven transmembrane domain proteins, has been characterized in many plant species [25]. Some
homologs act as PM susceptibility factors, as their loss of function results in a distinguished type of resistance known as
resistance. Originally discovered in barley (
L.),
resistance was later shown to occur in several monocot and eudicot species, namely Arabidopsis, tomato (
L.), pea (
L.), pepper (
L.), tobacco (
L.), and wheat (
L.) [26][27] plants. The callose synthase encoded by
is responsible for the production of callose in response to biotic and abiotic stresses. In tomato and potato plants, the knockout and silencing of
led to
and
is involved in the conversion of salicylic acid (SA) to 2,3-dihydroxybenzoic acid (2,3-DHBA) and negatively regulates defense gene expression [30]. Its silencing caused resistance to hemi-biotrophic
,
, and
[31]. Mutants of
, encoding for a cyclic nucleotide-gated cation channel, showed
resistance [29].
At the moment, studies on S-genes in woody plant species have been carried out only for
genes in rubber trees [32], poplar trees [33], apple trees, and grapevines [34]. In our work, we report on the S-genes identification and characterization in
on the basis of sequence homology, functional domain detection and phylogenetic relationships. In addition, the expression dynamics of S-genes were assessed in
and
plants inoculated with the two pathogens,
and
, belonging to different kingdoms. Using the same plant material, the transcription levels of key genes involved in pathogen resistance,
(
) and
(
, were also determined. Our analysis revealed the strong activation of
and
in response to infection by both
and
.
3. Development and Findings
C. sativais a European woody tree species commonly used across the globe in the food and timber industries. This chestnut species is susceptible to the two major pathogens,
P. cinnamomiand
C. parasitica [10,36]. In contrast, the Asian chestnut species[10][35]. In contrast, the Asian chestnut species
C. crenataand
C. mollissimaexhibit higher tolerance to
P. cinnamomiand
C. parasitica [6]. Achieving tolerance or resistance to pathogens is the major aim of rootstock breeding. Blight-resistant trees were obtained through backcross breeding of introgression genes from Asian species into American chestnut trees. [37]. However, this approach, although successful in developing blight-resistant American chestnut selections has been slowed by a lack of genetic tools. In Europe, ink disease tolerant hybrids were obtained through interspecific crosses between[6]. Achieving tolerance or resistance to pathogens is the major aim of rootstock breeding. Blight-resistant trees were obtained through backcross breeding of introgression genes from Asian species into American chestnut trees. [36]. However, this approach, although successful in developing blight-resistant American chestnut selections has been slowed by a lack of genetic tools. In Europe, ink disease tolerant hybrids were obtained through interspecific crosses between
C. sativaand
C. crenata, although the nut quality produced by these hybrids is below current market standards [38,39]., although the nut quality produced by these hybrids is below current market standards [37][38].
It has long been recognized that a deep understanding of a pathogen’s biology, host–pathogen interactions, and the resistance mechanisms are fundamental to improving breeding programs. Genomic and transcriptomic analyses have provided the first genetic insights into mechanisms underlying susceptible and resistant chestnut species responses to
P. cinnamomiand
C. parasitica [37,38,40,41,42]. Santos et al. [40] reported the upregulation of a set of candidate genes (e.g.,[36][37][39][40][41]. Santos et al. [39] reported the upregulation of a set of candidate genes (e.g.,
Cast_Gnk2-likeand
Calcium-dependent protein kinase) after
P. cinnamomiinfection, which may trigger HR-like cell death in
C. crenata cells. A significant number of genes involved in the defense against chestnut blight were identified [37].cells. A significant number of genes involved in the defense against chestnut blight were identified [36].
Traditionally, the introduction of resistance gene analogues into plants was the most promising approach to facilitate the acquisition of resistance. However, it did not prove to be durable enough because the widespread use of R genes caused the selection of pathogens capable of overcoming them [24]. Susceptibility (S) genes can be interesting candidates to be used in target breeding programs [22,23,24]. Furthermore, on the basis of previous studies, it was highlighted that the disabling of susceptibility genes may facilitate durable resistance since the pathogen needs to gain a new function to replace the lost host factor it was exploiting [43].Traditionally, the introduction of resistance gene analogues into plants was the most promising approach to facilitate the acquisition of resistance. However, it did not prove to be durable enough because the widespread use of R genes caused the selection of pathogens capable of overcoming them [24]. Susceptibility (S) genes can be interesting candidates to be used in target breeding programs [22][23][24]. Furthermore, on the basis of previous studies, it was highlighted that the disabling of susceptibility genes may facilitate durable resistance since the pathogen needs to gain a new function to replace the lost host factor it was exploiting [42].
In woody species, the investigation of S-genes has been performed only for MLO genes in rubber trees [32], poplar trees [33], apple trees, and grapevines [34,44]. Due to the absence of aIn woody species, the investigation of S-genes has been performed only for MLO genes in rubber trees [32], poplar trees [33], apple trees, and grapevines [34][43]. Due to the absence of a
C. sativagenome, highly similar S-genes were selected using the
C. mollissimav 1.1 genome. Based on the blastn survey, four loci with high similarity were identified in the
C. mollissima genome and attributed to different subclasses of S-genes [31,45,46,47] due to the presence of specific domains:genome and attributed to different subclasses of S-genes [31][44][45][46] due to the presence of specific domains:
mlo1,
dmr6,
dnd1, and
pmr4(
Figure 1,
Table 1). As previously observed [31], in the phylogenetic trees, monocot proteins formed a separate clade with respect to those of dicotyledonous species, supporting the hypothesis that an independent evolution occurred for these genes (
Figure 2). Quantitative PCR analysis has been carried out to identify the differential expression of candidate S-genes in response to
P. cinnamomiand
C. parasiticain the stems of a susceptible species,
C. sativa, and of a tolerant one,
C. crenata. Lesion analysis and DNA quantification of the pathogen (
Figure 4 and3 and
Figure 6) confirmed the higher tolerance level of4) confirmed the higher tolerance level of
C. crenatain response to both
P. cinnamomiand
C. parasiticainfection. Our qPCR results highlighted the main upregulation of
pmr4and
dmr6in response to infection by both
P. cinnamomiand
C. parasitica. As expected, a greater increase in the transcription of these susceptibility genes was observed in the susceptible species
C. sativa. Remarkably,
pmr4was strongly expressed at early infection phases of both pathogens in
C. sativa; in the tolerant
C. crenata, significant upregulation was observed (
Figure 5and
Figure 7).6).
Pmr4codifies for a callose synthase, which is necessary to create a physical barrier to avoid pathogen penetration and is also implicated in plant-triggered immunity suppression.
Pmr4 is thus not only involved in the synthesis of callose, but it also acts as a negative regulator of the salicylic acid pathway [28].is thus not only involved in the synthesis of callose, but it also acts as a negative regulator of the salicylic acid pathway [28].
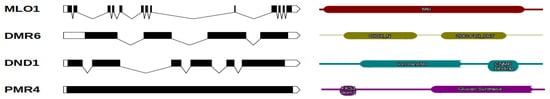
Figure 1. Chestnut S-genes and their protein structures. The graphical representations of gene exon/intron structures were generated using the http://wormweb.org/exonintron tool (accessed on 31 March 2021) and are shown in the left panel. The exons are indicated with black boxes, whereas introns are shown with lines. In the right panel, the protein structural domains are displayed.
Table 1. S-genes detected in the C. mollissima v1.1 genome and protein domain annotations.
| Gene Name | Scaffold | ORF Length (bp) | N° Exons | Size (aa) | Domains | PFAM DOMAINS |
|---|---|---|---|---|---|---|
| MLO1 | scaffold00101 | 1425 | 13 | 474 | Mlo | PF03094 |
| DMR6 | scaffold02358 | 1128 | 4 | 375 | 2OG-FeII_Oxy; DIOX_N | PF03171;PF14226 |
| DND1 | scaffold00410 | 1407 | 6 | 468 | cNMP_binding; Ion_trans | PF00027;PF00520 |
| PMR4 | scaffold00300 | 5346 | 1 | 1781 | FKS1_dom1; Glucan_synthase | PF14288;PF02364 |
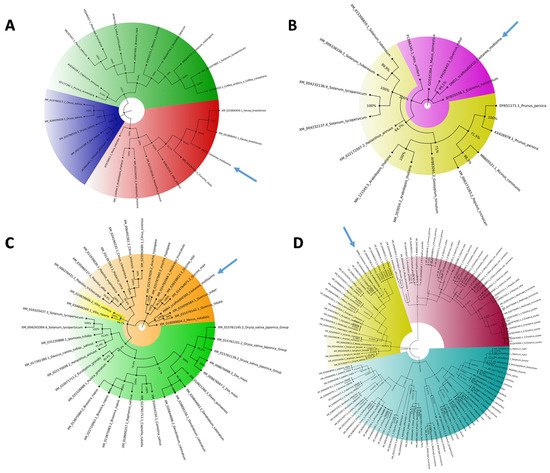
Figure 2. Phylogenetic analysis of the S-genes. The 4 phylogenetic trees of mlo1 (A), dnd1 (B), pmr4 (C), and dmr6 (D) were constructed using MEGAX software by aligning chestnut S-gene coding sequences with NCBI S-gene ortholog coding sequences. The colors indicate the main clades detected, and the arrows underline the location of C. mollissima.
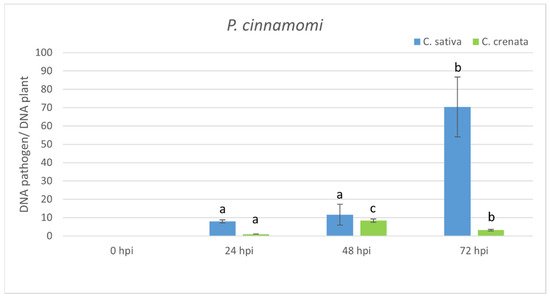
Figure 3. qRT-PCR pathogen DNA quantification after P. cinnamomi inoculation. Data were quantified using the 2−ΔΔCt method based on the Ct values of pathogen genes (ypt and mf1) and actin-7 as a housekeeping gene. Data are the means of three biological replicates ± SE. C. sativa data are normalized with C. sativa 0 hpi control; C. crenata data are normalized with C. crenata 0 hpi control. Different letters associated with the set of means indicate a significant difference based on Tukey’s HSD test (p ≤ 0.05).
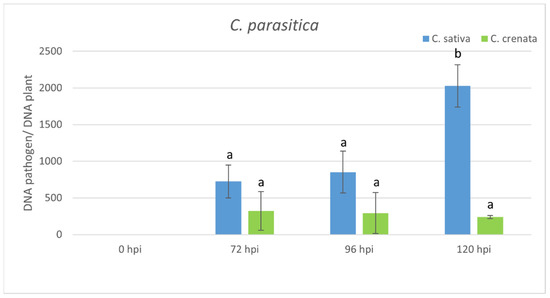
Figure 4. qRT-PCR pathogen DNA quantification after C. parasitica inoculation. Data were quantified using the 2−ΔΔCt method based on the Ct values of fungal genes (ypt and mf1) with actin-7 as a housekeeping gene. Data are the means of three biological replicates ± SE. C. sativa data are normalized with the C. sativa 0 hpi control; C. crenata data are normalized with C. crenata 0 hpi control. Different letters associated with the set of means indicate a significant difference based on Tukey’s HSD test (p ≤ 0.05).
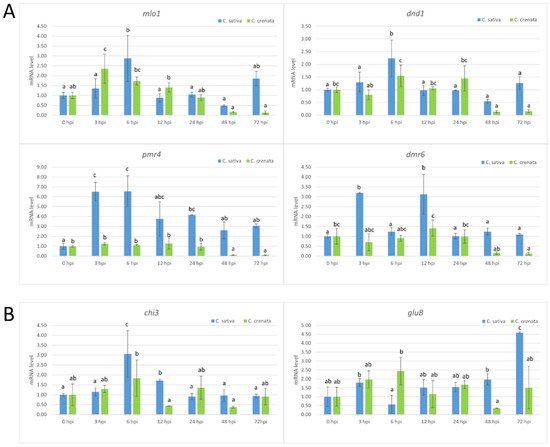
Figure 5. qRT-PCR-based transcription profiling after P. cinnamomi inoculation. (A) The S-gene transcription profiles in C. sativa (blue) and C. crenata (green) chestnut species. (B) The expression analysis of genes coding for several pathogenesis-related (PR) proteins in C. sativa (blue) and C. crenata (green) species. In all analyses, Cm7-actin was used as a housekeeping gene. Data are the means of three biological replicates ± SE. C. sativa data are normalized with C. sativa 0 hpi control; C. crenata data are normalized with C. crenata 0 hpi control. Different letters associated with the set of means indicate a significant difference based on Tukey’s HSD test (p ≤ 0.05).
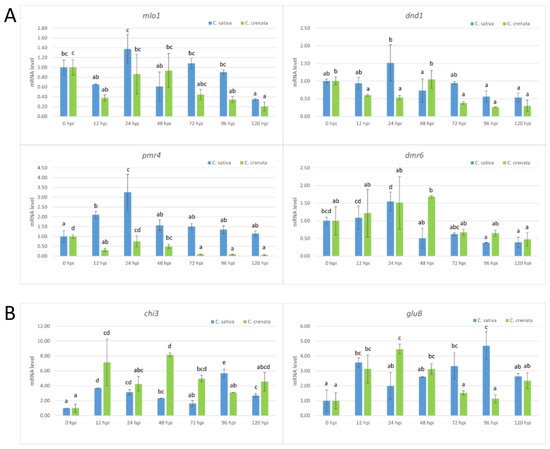
Figure 6. qRT-PCR-based transcription profiling after C. parasitica inoculation. (A) The S-gene transcription profile in C. sativa (blue) and C. crenata (green) chestnut species. (B) The expression analysis of genes coding for several pathogenesis-related (PR) proteins of C. sativa (blue) and C. crenata (green) species. In all the analyses, Cm7-actin was used as the housekeeping gene. The data are the means of three biological replicates ± SE. C. sativa data are normalized with C. sativa 0 hpi control; C. crenata data are normalized with C. crenata 0 hpi control. Different letters associated with the set of means indicate a significant difference based on Tukey’s HSD test (p ≤ 0.05).
Huibers et al. [47] demonstrated that resistance due to the silencing of
Pmr4 is associated with salicylic acid (SA) accumulation rather than with the callose deposition absence. Salicylic acid signaling plays a key role protecting against biotrophic pathogens through the establishment of a hypersensitive response (HR). Saiz-Fernandez et al. [49] revealed the increment of SA levels inis associated with salicylic acid (SA) accumulation rather than with the callose deposition absence. Salicylic acid signaling plays a key role protecting against biotrophic pathogens through the establishment of a hypersensitive response (HR). Saiz-Fernandez et al. [48] revealed the increment of SA levels in
P. cinnamomiinoculated stems, indicating that
P. cinnamomiactivates a defense response similar to that triggered by biotrophic pathogens. Inoculation with both virulent and hypovirulent strains of
C. parasitica led to SA accumulation in European chestnut plantlets that were grown in vitro [50]. Transcriptome analyses carried out in bothled to SA accumulation in European chestnut plantlets that were grown in vitro [49]. Transcriptome analyses carried out in both
C. dentataand
C. mollissima highlighted activation of salicylic-acid-related genes in canker tissues [37].highlighted activation of salicylic-acid-related genes in canker tissues [36].
In chestnut trees, callose deposition around
P. cinnamomi hyphae was detected early in the infection process; however, it does not seem to play a key role in the associated interactions since the pathogen can reach the vascular cylinder in both susceptible and resistant plant genotypes [51]. This result was validated by transcriptomes analyses ofhyphae was detected early in the infection process; however, it does not seem to play a key role in the associated interactions since the pathogen can reach the vascular cylinder in both susceptible and resistant plant genotypes [50]. This result was validated by transcriptomes analyses of
C. sativaand
C. crenata, in which no overexpression of
Callose synthasesafter
P. cinnamomi infection was observed [38].infection was observed [37].
Based on our results and the literature, we can hypothesize that callose accumulation due to the
pmr4upregulation in inoculated
C. sativalines may not be responsible for controlling
P. cinnamomicolonization. We suggest that the upregulation of
pmr4could lead to a negative regulation of the SA pathway that in turn provokes the susceptibility of
C. sativato both
P. cinnamomiand
C. parasitica. A clear link with SA pathway has emerged even with the other chestnut gene candidate
dmr6(downy mildew resistance 6). The mutation of
Arabidopsis dmr6gene, associated with salicylic acid (SA) homeostasis [31], results in the generation of plants that are resistant to bacteria and oomycetes.
Dmr6is involved in the conversion of salicylic acid (SA) to 2,3-dihydroxybenzoic acid (2,3-DHBA) and negatively regulates the expression of defense genes (PR-1, PR-2, and PR-5) [30].
The expression trend of the
Dmr6gene in response to
P. cinnamomiinfection turned out to be similar to the profile of
pmr4. Indeed,
dmr6was strongly expressed at early infection phases of
P. cinnamomiin
C. sativa; in
C. crenatano significant upregulation was detected (
Figure 5). No upregulation of
dmr6in response to
C. parasiticawas highlighted in both plant species (
Figure 7). We can thus hypothesize that6). We can thus hypothesize that
dmr6upregulation observed in
C. sativacould negatively regulate defense gene expression, leading to susceptibility to
P. cinnamomi.
Plants produce a variety of hydrolytic defense enzymes against pathogens, including chitinases, proteases, and also glucanases [52]. The genes coding for several pathogenesis-related (PR) proteins,Plants produce a variety of hydrolytic defense enzymes against pathogens, including chitinases, proteases, and also glucanases [51]. The genes coding for several pathogenesis-related (PR) proteins,
Acidic 26 kDa endochitinasegene (
chi3) and
Glucan endo-1,3-beta-glucosidase Bgene (
gluB), were selected in our analysis because they are considered as responsive to SA-dependent signaling [53,54].), were selected in our analysis because they are considered as responsive to SA-dependent signaling [52][53].
Chi3and
gluB are enzymes that cause the lysis of hyphae of various pathogens, resulting in growth inhibition [55,56,57].are enzymes that cause the lysis of hyphae of various pathogens, resulting in growth inhibition [54][55][56].
In both
C. sativaand
C. crenataplants inoculated with
C. parasitica,
chi3and
gluBwere significantly upregulated. The transcription of
chi3was higher in
C. crenatathan in
C. sativa, presumably as a consequence of the improved defense mechanism against
C. parasitica. Our results are in agreement with Shain et al. [58], who demonstrated the involvement of. Our results are in agreement with Shain et al. [57], who demonstrated the involvement of
b-1,3-glucanaseand
chitinasein chestnut species affected by
C. parasitica. Studies on the role of
chitinasein blight infection mostly involved
C. sativa as a model system [50,59]. In bothas a model system [49][58]. In both
C. dentataand
C. mollissima, transcripts of several compounds expressing
chitinase accumulated more in canker tissues than healthy stem tissues [37]. In order to obtain chestnut plants with potentially increased resistance/tolerance to chestnut blight, the endogenous Ch3gene encoding a chitinase-like protein was over-expressed in the European chestnut through Agrobacterium-mediated transformation [60].accumulated more in canker tissues than healthy stem tissues [36]. In order to obtain chestnut plants with potentially increased resistance/tolerance to chestnut blight, the endogenous Ch3gene encoding a chitinase-like protein was over-expressed in the European chestnut through Agrobacterium-mediated transformation [59].
The emergent CRISPR/Cas9 technology is expected to play a key role in future crop breeding as it allows highly efficient gene editing and generates genetic changes indistinguishable from those arising spontaneously in nature or through conventional breeding [61]. Several examples of edited plants resistant to fungal pathogens have been described [62,63]. For example, genome editing was successfully applied to knock outThe emergent CRISPR/Cas9 technology is expected to play a key role in future crop breeding as it allows highly efficient gene editing and generates genetic changes indistinguishable from those arising spontaneously in nature or through conventional breeding [60]. Several examples of edited plants resistant to fungal pathogens have been described [61][62]. For example, genome editing was successfully applied to knock out
mlo S-genes, leading to Powdery mildew (PM) resistance [44,64,65,66].S-genes, leading to Powdery mildew (PM) resistance [43][63][64][65].
Pmr4and
dmr6 loss-of-function through CRISPR/Cas reduced the susceptibility to PM in tomato plants [28,67]. In our laboratory, we are setting up a CRISPR/Cas9 transformation protocol inloss-of-function through CRISPR/Cas reduced the susceptibility to PM in tomato plants [28][66]. In our laboratory, we are setting up a CRISPR/Cas9 transformation protocol in
Castanea sativa. Our future goal will be to perform the functional characterization using the CRISPR/Cas9 approach of the two candidate genes (
dmr6and
pmr4), while checking if the two genes may also play a role in the interaction between
C. sativaand the emergent nut rot and canker agent
Gnomoniopsis castaneae [68].[67].
References
- Torello Marinoni, D.; Nishio, S.; Valentini, N.; Shirasawa, K.; Acquadro, A.; Portis, E.; Alma, A.; Akkak, A.; Pavese, V.; Cavalet-Giorsa, E.; et al. Development of High-Density Genetic Linkage Maps and Identification of Loci for Chestnut Gall Wasp Resistance in Castanea spp. Plants 2020, 9, 1048.
- Conedera, M.; Krebs, P.; Tinner, W.; Pradella, M.; Torriani, D. The cultivation of Castanea sativa (Mill.) in Europe, from its origin to its diffusion on a continental scale. Veg. Hist. Archaeobot. 2004, 13, 161–179.
- Müllerová, J.; Hédl, R.; Szabó, P. Coppice abandonment and its implications for species diversity in forest vegetation. For. Ecol. Manag. 2015, 343, 88–100.
- Zentmyer, G.A. Phytophthora cinnamomi and The Diseases it Causes; The American Phytopathological Society: St. Paul, MN, USA, 1980; Volume 10, p. 96.
- Cristinzio, G.; Grassi, G. Valutazione di resistenza a Phytophthora cambivora e Phytophthora cinnamomi in cultivar di Castanea sativa. Monti Boschi 1993, 1, 54–58.
- Serrazina, S.; Machado, H.; Costa, R.; Duque, P.; Malhó, R. Expression of Castanea crenata Allene Oxide Synthase in Arabidopsis Improves the Defense to Phytophthora cinnamomi. bioRxiv 2020.
- Maurel, M.; Robin, C.; Simonneau, T.; Loustau, D.; Dreyer, E.; Desprez-Loustau, M.-L. Stomatal conductance and root-to-shoot signalling in chestnut saplings exposed to Phytophthora cinnamomi or partial soil drying. Funct. Plant Biol. 2004, 31, 41–51.
- Vannini, A.; Vettraino, A.M. Ink disease in chestnuts: Impact on the European chestnut. For. Snow Landsc. Res. 2002, 76, 345–350.
- Robin, C.; Smith, I.; Hansen, E.M. Phythophthora cinnamomi. For. Phytophthoras 2012, 2, 1.
- Gonthier, P.; Robin, C. Diseases. In The Chestnut Handbook: Crop and Forest Management; Beccaro, G., Alma, A., Bounous, G., Gomes-Laranjo, J., Eds.; CRC Press, Taylor & Francis Group: Boca Raton, FL, USA, 2019; pp. 297–316, Chapter 11; ISBN 9781138334021.
- Hayden, K.J.; Garbelotto, M.; Dodd, R.; Wright, J.W. Scaling up from greenhouse resistance to fitness in the field for a host of an emerging forest disease. Evol. App. 2013, 6, 970–982.
- Bounous, G. Il Castagno: Risorsa Multifunzionale in Italia e nel Mondo; Edagricole: Bologna, Italy, 2014.
- Rigling, D.; Prospero, S. Cryphonectria parasitica, the causal agent of chestnut blight: Invasion history, population biology and disease control. Mol. Plant Pathol. 2018, 1, 7–20.
- Lione, G.; Giordano, L.; Turina, M.; Gonthier, P. Hail-Induced Infections of the Chestnut Blight Pathogen Cryphonectria parasitica Depend on Wound Size and May Lead to Severe Diebacks. Phytopathology 2020, 110, 1280–1293.
- Trapiello, E.; González-Varela, G.; González, A.J. Chestnut Blight Control by Agrochemicals in Castanea sativa under Managed Conditions. J. Plant Dis. Prot. 2015, 122, 120–124.
- Westbrook, J.W.; Zhang, Q.; Mandel, M.K.; Jenkins, E.V.; Barth, L.E.; Jenkins, J.W.; Grimwood, J.; Schmutz, J.; Holliday, J.A. Genomic selection analyses reveal tradeoff between blight tolerance and genome inheritance from American chestnut (Castenea dentata) in [C. dentata x C. mollissima] x C. dentata backcross populations. BioRxiv 2019.
- Kubisiak, T.L.; Hebard, F.V.; Nelson, C.D.; Zhang, J.; Bernatzky, R.; Huang, H.; Anagnostakis, S.L.; Doudrick, R.L. Molecular Mapping of Resistance to Blight in an Interspecific Cross in the genus Castanea. Phytopathology 1997, 87, 751–759.
- Kubisiak, T.L.; Nelson, C.D.; Staton, M.E.; Zhebentyayeva, T.; Smith, C.; Olukolu, B.A.; Fang, G.-C.; Hebard, F.V.; Anagnostakis, S.; Wheeler, N.; et al. A transcriptome-based genetic map of Chinese chestnut (Castanea mollissima) and identification of regions of segmental homology with peach (Prunus persica). Tree Genet. Genomes 2012, 9, 557–571.
- Zhang, B.; Oakes, A.D.; Newhouse, A.E.; Baier, K.M.; Maynard, C.A.; Powell, W.A. A threshold level of oxalate oxidase transgene expression reduces Cryphonectria parasitica-induced necrosis in a transgenic American chestnut (Castanea dentata) leaf bioassay. Transgenic Res. 2013, 22, 973–982.
- Marcolin, E.; Manetti, M.C.; Pelleri, F.; Conedera, M.; Pezzatti, G.B.; Lingua, E.; Pividori, M. Seed regeneration of sweet chestnut (Castanea sativa Miller) under different coppicing approaches. For. Ecol. Manag. 2020, 472, 118273.
- Aebi, A.; Schönrogge, K.; Melika, G.; Alma, A.; Bosio, G.; Quacchia, A.; Picciau, L.; Abe, Y.; Moriya, S.; Yara, K.; et al. Parasitoid recruitment to the globally invasive chestnut gall wasp Dryocosmus kuriphilus. In Ecology and Evolution of Galling Arthropods and Their Associates; Ozaki, K., Yukwa, J., Ohgushi, T., Price, P.W., Eds.; Springer: Tokyo, Japan, 2006; pp. 103–121.
- Van Schie, C.C.; Takken, F.L. Susceptibility genes 101: How to be a good host. Annu. Rev. Phytopathol. 2014, 52, 551–581.
- Zaidi, S.S.; Mukhtar, M.S.; Mansoor, S. Genome editing: Targeting susceptibility genes for plant disease resistance. Trends Biotechnol. 2018, 36, 898–906.
- Pavan, S.; Jacobsen, E.; Visser, R.G.; Bai, Y. Loss of susceptibility as a novel breeding strategy for durable and broad-spectrum resistance. Mol. Breed. 2010, 25, 1–12.
- Devoto, A.; Hartmann, H.A.; Piffanelli, P.; Elliott, C.; Simmons, C.; Taramino, G.; Goh, C.S.; Cohen, F.E.; Emerson, B.C.; Schulze-Lefert, P.; et al. Molecular phylogeny and evolution of the plant-specific seven-transmembrane MLO family. J. Mol. Evol. 2003, 56, 77–88.
- Büschges, R.; Hollricher, K.; Panstruga, R.; Simons, G.; Wolter, M.; Frijters, A.; van Daelen, R.; van der Lee, T.; Diergaarde, P.; Groenendijk, J.; et al. The barley Mlo gene: A novel control element of plant pathogen resistance. Cell 1997, 88, 695–705.
- Appiano, M.; Pavan, S.; Catalano, D.; Zheng, Z.; Bracuto, V.; Lotti, C.; Visser, R.G.; Ricciardi, L.; Bai, Y. Identification of candidate MLO powdery mildew susceptibility genes in cultivated Solanaceae and functional characterization of tobacco NtMLO1. Transgenic Res. 2015, 24, 847–858.
- Santillán Martínez, M.I.; Bracuto, V.; Koseoglou, E.; Appiano, M.; Jacobsen, E.; Visser, R.G.F.; Wolters, A.A.; Bai, Y. CRISPR/Cas9-targeted mutagenesis of the tomato susceptibility gene PMR4 for resistance against powdery mildew. BMC Plant Biol. 2020, 20, 284.
- Sun, K.; Wolters, A.M.A.; Loonen, A.E.; Huibers, R.P.; van der Vlugt, R.; Goverse, A.; Jacobsen, E.; Visser, R.G.; Bai, Y. Down-regulation of Arabidopsis DND1 orthologs in potato and tomato leads to broad-spectrum resistance to late blight and powdery mildew. Transgenic Res. 2016, 25, 123–138.
- Van Damme, M.; Huibers, R.P.; Elberse, J.; Van Den Ackerveken, G. Arabidopsis DMR6 encodes a putative 2OG-Fe(II) oxygenase that is defense-associated but required for susceptibility to downy mildew. Plant J. 2008, 54, 785–793.
- Zeilmaker, T.; Ludwig, N.R.; Elberse, J.; Seidl, M.F.; Berke, L.; Van Doorn, A.; Schuurink, R.C.; Snel, B.; Van den Ackerveken, G. Downy mildew resistant 6 and DMR6-like oxygenase 1 are partially redundant but distinct suppressors of immunity in Arabidopsis. Plant J. 2015, 81, 210–222.
- Liyanage, K.K.; Khan, S.; Herath, V.; Brooks, S.; Mortimer, P.E.; Nadir, S.; Hyde, K.D.; Xu, J. Genome Wide Identification of the MLO Gene Family Associated with Powdery Mildew Resistance in Rubber Trees (Hevea brasiliensis). Trop. Plant Biol. 2020, 13, 331–342.
- Filiz, E.; Vatansever, R. Genome-wide identification of mildew resistance locus O (MLO) genes in tree model poplar (Populus trichocarpa): Powdery mildew management in woody plants. Eur. J. Plant Pathol. 2018, 152, 95–109.
- Pessina, S.; Lenzi, L.; Perazzolli, M.; Campa, M.; Dalla Costa, L.; Urso, S.; Valè, G.; Salamini, F.; Velasco, R.; Malnoy, M. Knockdown of MLO genes reduces susceptibility to powdery mildew in grapevine. Hortic. Res. 2016, 3, 16016.
- Pereira-Lorenzo, S.; Ballester, A.; Corredoira, E.; Bounous, G.; Botta, R.; Beccaro, G.L.; Kubisiak, T.L.; Conedera, M.; Krebs, P.; Yamamoto, T.; et al. Chestnut. In Fruit Breeding; Badenes, M.L., Byrne, D.H., Eds.; Springer Science + Business Media: New York, NY, USA, 2012; pp. 729–769.
- Barakat, A.; Staton, M.; Cheng, C.H.; Park, J.; Yassin, N.B.M.; Ficklin, S.; Yeh, C.C.; Hebard, F.; Baier, K.; Powell, W.; et al. Chestnut resistance to the blight disease: Insights from transcriptome analysis. BMC Plant Biol. 2012, 12, 38.
- Serrazina, S.; Santos, C.; Machado, H.; Pesquita, C.; Vicentini, R.; Pais, M.S.; Sebastiana, M.; Costa, R. Castanea root transcriptome in response to Phytophthora cinnamomi challenge. Tree Genet. Genomes 2015, 11, 19.
- Santos, C.; Machado, H.; Correia, I.; Gomes, F.; Gomes-Laranjo, J.; Costa, R. Phenotyping Castanea hybrids for Phytophthora cinnamomi resistance. Plant Pathol. 2015, 64, 901–910.
- Santos, C.; Duarte, S.; Tedesco, S.; Fevereiro, P.; Costa, R.L. Expression profiling of Castanea genes during resistant and susceptible interactions with the oomycete pathogen Phytophthora cinnamomi reveal possible mechanisms of immunity. Front. Plant Sci. 2017, 8, 515.
- Santos, C.; Nelson, C.D.; Zhebentyayeva, T.; Machado, H.; Gomes-Laranjo, J.; Costa, R.L. First interspecific genetic linkage map for Castanea sativa x Castanea crenata revealed QTLs for resistance to Phytophthora cinnamomi. PLoS ONE 2017, 12, e0184381.
- Barakat, A.; DiLoreto, D.S.; Zhang, Y.; Smith, C.; Baier, K.; Powell, W.A.; Wheeler, N.; Sederoff, R.; Carlson, J.E. Comparison of the transcriptomes of American chestnut (Castanea dentata) and Chinese chestnut (Castanea mollissima) in response to the chestnut blight infection. BMC Plant Biol. 2009, 9, 51.
- van Esse, H.P.; Reuber, T.L.; van der Does, D. Genetic modification to improve disease resistance in crops. New Phytol. 2020, 225, 70–86.
- Wan, D.Y.; Guo, Y.; Cheng, Y.; Hu, Y.; Xiao, S.; Wang, Y.; Wen, Y.Q. CRISPR/Cas9-mediated mutagenesis of VvMLO3 results in enhanced resistance to powdery mildew in grapevine (Vitis vinifera). Hortic. Res. 2020, 7, 116.
- Clough, S.J.; Fengler, K.A.; Yu, I.C.; Lippok, B.; Smith, R.K.; Bent, A.F. The Arabidopsis dnd1 “defense, no death” gene encodes a mutated cyclic nucleotide-gated ion channel. Proc. Natl. Acad. Sci. USA 2000, 97, 9323–9328.
- Nishimura, M.T.; Stein, M.; Hou, B.H.; Vogel, J.P.; Edwards, H.; Somerville, S.C. Loss of a callose synthase results in salicylic acid-dependent disease resistance. Science 2003, 301, 969–972.
- Zhang, Y.; Zhao, L.; Zhao, J.; Li, Y.; Wang, J.; Guo, R.; Gan, S.; Liu, C.J.; Zhang, K. S5H/DMR6 encodes a salicylic acid 5-hydroxylase that fine-tunes salicylic acid homeostasis. Plant Physiol. 2017, 175, 1082–1093.
- Huibers, R.P.; Loonen, A.E.; Gao, D.; Van den Ackerveken, G.; Visser, R.G.; Bai, Y. Powdery mildew resistance in tomato by impairment of SlPMR4 and SlDMR1. PLoS ONE 2013, 8, e67467.
- Saiz-Fernández, I.; Milenković, I.; Berka, M.; Černý, M.; Tomšovský, M.; Brzobohatý, B.; Kerchev, P. Integrated Proteomic and Metabolomic Profiling of Phytophthora cinnamomi Attack on Sweet Chestnut (Castanea sativa) Reveals Distinct Molecular Reprogramming Proximal to the Infection Site and Away from It. Int. J. Mol. Sci. 2020, 21, 8525.
- Lovat, C.-A.; Donnelly, D.J. Mechanisms and metabolomics of the host-pathogen interactions between Chestnut (Castanea species) and Chestnut blight (Cryphonectria parasitica). For. Pathol. 2019, 49, e12562.
- Fernandes, P.; Machado, H.; Silva, M.D.C.; Costa, R.L. A Histopathological Study Reveals New Insights Into Responses of Chestnut (Castanea spp.) to Root Infection by Phytophthora cinnamomi. Phytopathology 2021.
- Veluthakkal, R.; Dasgupta, M.G. Pathogenesis-related genes and proteins in forest tree species. Trees 2010, 24, 993–1006.
- Puthoff, D.P.; Holzer, F.M.; Perring, T.M.; Walling, L.L. Tomato Pathogenesis-related Protein Genes are Expressed in Response to Trialeurodes vaporariorum and Bemisia tabaci Biotype B Feeding. J. Chem. Ecol. 2010, 36, 1271–1285.
- Molinari, S.; Fanelli, E.; Leonetti, P. Expression of tomato salicylic acid (SA)-responsive pathogenesis-related genes in Mi-1-mediated and SA-induced resistance to root-knot nematodes. Mol. Plant Pathol. 2014, 15, 255–264.
- Broglie, K.; Chet, I.; Holliday, M.; Cressman, R.; Biddle, P.; Knowlton, S.; Mauvais, C.J.; Broglie, R. Transgenic plants with enhanced resistance to the fungal pathogen, Rhizoctonia solani. Science 1991, 254, 1194–1197.
- Mauch, F.; Staehelin, L.A. Functional Implications of the Subcellular Localization of Ethylene-Induced Chitinase and [beta]-1,3-Glucanase in Bean Leaves. Plant Cell 1989, 1, 447–457.
- Schlumbaum, A.; Mauch, F.; Vögeli, U.; Boller, T. Plant chitinases are potent inhibitors of fungal growth. Nature 1986, 324, 365–367.
- Shain, L.; Miller, J.B.; Spalding, R.J. Responses of American and Chinese Chestnut to Cryphonectria parasitica and Ethylene. In Proceedings of the International Chestnut Conference; Double, M.L., MaCDonald, D.H., Eds.; West Virginia University Press: Morgantown, WV, USA, 1994; pp. 97–101.
- Vannini, A.; Caruso, C.; Leonardi, L.; Rugini, E.; Chiarot, E.; Caporale, C.; Buonocore, V. Antifungal properties of chitinases from Castanea sativa against hypovirulent and virulent strains of the chestnut blight fungus Cryphonectria parasitica. Physiol. Mol. Plant Pathol. 1999, 55, 29–35.
- Corredoira, E.; San José, M.C.; Vieitez, A.M.; Allona, I.; Aragoncillo, C.; Ballester, A. Agrobacterium-mediated transformation of European chestnut somatic embryos with a Castanea sativa (Mill.) endochitinase gene. New For. 2016, 47, 669–684.
- Zhu, H.; Li, C.; Gao, C. Applications of CRISPR–Cas in agriculture and plant biotechnology. Nat. Rev. Mol. Cell Biol. 2020, 21, 661–677.
- Schenke, D.; Cai, D. Applications of CRISPR/Cas to Improve Crop Disease Resistance: Beyond Inactivation of Susceptibility Factors. iScience 2020, 23, 101478.
- Tyagi, S.; Kesiraju, K.; Saakre, M.; Rathinam, M.; Raman, V.; Pattanayak, D.; Sreevathsa, R. Genome editing for resistance to insect pests: An emerging tool for crop improvement. ACS Omega 2020, 5, 20674–20683.
- Wang, Y.; Cheng, X.; Shan, Q.; Zhang, Y.; Liu, J.; Gao, C.; Qiu, J.L. Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat. Biotechnol. 2014, 32, 947–951.
- Malnoy, M.; Viola, R.; Jung, M.H.; Koo, O.-J.; Kim, S.; Kim, J.-S.; Velasco, R.; Kanchiswamy, C.N. DNA-Free Genetically Edited Grapevine and Apple Protoplast Using CRISPR/Cas9 Ribonucleoproteins. Front. Plant Sci. 2016, 7, 1904.
- Nekrasov, V.; Wang, C.; Win, J.; Lanz, C.; Weigel, D.; Kamoun, S. Rapid generation of a transgene-free powdery mildew resistant tomato by genome deletion. Sci. Rep. 2017, 7, 482.
- de Toledo Thomazella, D.P.; Brail, Q.; Dahlbeck, D.; Staskawicz, B.J. CRISPR-Cas9 mediated mutagenesis of a DMR6 ortholog in tomato confers broad-spectrum disease resistance. bioRxiv 2016, 1–23.
- Lione, G.; Danti, R.; Fernandez-Conradi, P.; Ferreira-Cardoso, J.V.; Lefort, F.; Marques, G.; Meyer, J.B.; Prospero, S.; Radòcz, L.; Robin, C.; et al. The emerging pathogen of chestnut Gnomoniopsis castaneae: The challenge posed by a versatile fungus. Eur. J. Plant Pathol. 2019, 153, 671–685.
